*My interview with George Daley has now been published in Development*
Last month I went to Boston for the Annual Meeting of the International Society for Stem Cell Research (ISSCR), which was celebrating its 15th birthday. Sally Temple handed over the Presidential reins to Hans Clevers with a reminder of what a remarkable 15 years it had been for the field – induced pluripotent cells (iPSCs), an explosion of differentiation protocols, the arrival of organoids, CRISPR and single cell sequencing, new windows onto human development, clinical trials and drug screening. She encouraged the audience to think “This is our century” – that is, the century of harnessing the power of stem cells in tackling disease. She also reiterated the ISSCR’s commitment to keep basic science at the core of what it does (you can hear more of her thoughts on the ISSCR in Caroline Hendry’s video interview for Development). In fact the whole meeting (or at least what I got out of it) was a healthy mix of basic and applied, from model organisms to drug trials, and developmental biology was repeatedly heralded as a vital foundation for any future clinical advance; my worries about wandering into a wall-to-wall differentiation protocol fest turned out to be unfounded.
So here’s my belated diary of highlights and impressions from the meeting. There were a thousand talks and posters so of course I missed a lot: if you went and want to write something about the meeting, just register and you’re free to post. Also check out the highlights from Agnes Soos, Cátia Bandeiras and RegenMedNet.

Tuesday, 13th June
Over the Atlantic the in-flight pasta meal tasted like it was cooked in popcorn butter. In Boston it was sweltering until out of nowhere a storm cooled the air down, and in the evening I went to a basement bar where Twitter had assembled a dozen strangers going to the meeting to meet in real life (a ‘Tweet Up’). For someone who knew hardly anyone at the meeting, it was a great way to meet new people and something I’ll try to do again (thanks to Samantha Yammine for instigating this!). After some local IPAs packed with enough hops to blow your head off, I stopped in at a Wallgreen’s on my way back to the hotel and had one of the more depressing late night dinners of my life: Salsitas followed by a flapjack.
Wednesday, 14th June
In the morning I went to a focus session on the ethics of organoids, which was chaired by Megan Munsie (more on Megan’s work below) and paired scientists with ethicists to discuss the impact and meaning of this new technology on wider society.
On the scientific side of things, Hans Clevers gave us the latest update on how organoids derived from adult (often patient) stem cells were helping to understand various pathologies and inform treatments. In an almost throwaway comment he told us that he could make lung organoids from patients’ sputum, and kidney organoids from their urine; you just need to spin the cells down and start the protocol. It struck me as one of those ‘living in the future’ moments. Melissa Little reminded us how critical an understanding normal development is for organoid derivation – you need a guide for which factors to add and in which order, something echoed for iPSC differentiation later in the meeting by George Daley. Indeed most iPSC-derived organoids are models for embryonic rather than mature tissues, which can complicate interpretations. Where the technology is particularly helping Little’s lab is to validate variants of unknown function in genetic kidney diseases – derive iPSCs from patients, ‘fix’ the sequence with gene editing, and see if organoids can form normally; this is such a cool clinical use of organoids that I hadn’t appreciated before.
On the ethics side, Annelien Bredenoord emphasised the inevitable collision of organoid technology and ethics – we have to decide what the moral and legal status of organoids are. Do we treat organoids like any other patient-derived tissue? Are they most akin to cell lines, or something else? What types of consent do we expect patients to give? How do patients see their own oragnoids? And what about if private companies seek to use organoids?
How we answer these questions might really depend on definitions: for Melissa Little, organoids are really just an extension of iPSC differentiation protocols, and if so we need not radically rethink the guidelines that exist for iPSCs (for instance, those written up by the ISSCR). There is also a lot of difference between different kinds of organoids, depending on how complex an organ they are modelling. Plus, according to Jeurgen Knoeblich, the ethical concerns around those most provocative of organoids, cortical ones – “when will they think?” – are based on a perceived rather than a real risk to societal ethics, given that we are light years away from making something capable of a thought or a feeling. Later in the meeting Knoeblich told us about his lab’s efforts to make more complex and representative brain organoids. You can dorsalise one organoid, and ventralise another, culture them together until they fuse, and then observe neural migration from ventral to dorsal, a feature that previous cortical organoids lacked.
Bredenoord’s work involves asking patients what they think about organoids derived from their cells, and revealed ambivalence in one particular cystic fibrosis patient community. Melissa Little had a different experience with kidney patients: they want the opportunity to donate tissue or cells to any initiative that might help them, and organoids are no different; I heard similar things from Jayaraj Rajagopal. Bredenoord also described a reluctance to donate tissue if it was to be commercialised by a private company, and this was echoed by others and presumably a feeling not unique to organoids. Hans Clevers described a fix for this, which was to establish not-for-profits to run the biobanks and profit from commercial customers. The question of consent in biobanking has recently been explored by Tim Caulfield and Blake Murdoch in PLoS Biology.
Insoo Hyun, who sits on the ISSCR’s ethics committee, made the distinction between two types of research ethics: philosophical and practical. Philosophical ethics cover the questions of “should we be doing this?” and “is it wise?”, while practical ethics emphasises the regulatory guidelines in place once we’ve said yes to the philosophical questions. I guess we’ve pretty much gone into the practical side of things – no one in the session questioned the use or potential of organoids, though some thought the hype might catch up with us soon – but he reminded us of the need to really educate the public and justify what we are doing to cover the philosophical part. To paraphrase Hyun, “don’t just forget the public and leave it to committee”.
It was quite a compelling session, though I didn’t exactly come out of it with a clear feeling for where organoids stand from an ethical perspective; I guess that was the point of the session, perhaps the field is too young. For a more coherent discussion of the various ethical issues in hand I’ll send you to a couple of reviews by the speakers, one in Development and one in Science.
The rest of the day was spent in the aircraft-hangar-sized hall where the plenaries were held. It was funny seeing the great and the good walk up to the podium to the snippets of EDM bangers or Coldplay, though I was disappointed that none of them danced their way up. A series of research talks was followed by Sanford Greenberg, who, as a roommate of Art Garfunkel in college (we can thank Greenberg for financing Art’s initial union with Paul Simon), went blind and has since dedicated much of his life to ending blindness, “this wretched curse”. You can check out the website detailing his 3 million dollar prize for blindness research at endblindnessby2020.com.

That evening I actually managed to have dinner, at the Barking Crab – a raucous place on the water, with snow crab and another great IPA (Boston’s a wonderful place for a beer drinker).

Thursday 15th June
In the afternoon I had the mindhurt of trying to choose from one of seven (!) concurrent sessions. The ‘Single Cell Heterogeneity’ session featured so much single cell sequencing of organs and organoids that one almost came out of it feeling blasé about the whole enterprise – just to think of telling a developmental biologist from the eighties that she could sequence the genome of every cell in a developing organ! It wasn’t all sequencing though: David Scadden explored the role of the stem cell niche in blood development, starting with some history (read more here). In the seventies, Raymond Schofield had proposed the niche hypothesis but apparently was at loggerheads with Ernest McCulloch about it – he ended up leaving science and becoming a sheep farmer in Wales, where Scadden met him for a pint in the village pub to discuss his ideas. Scadden’s approach was to test the role of the niche by taking stem cells from one animal and place them into another. Under these new conditions, they behaved pretty much the same. HSCs are very heterogeneous, but this is intrinsic rather than defined by the niche, and Scadden used a game-ey metaphor: rather than highly adaptive ‘transformers’, HSCs are like a collection of chess pieces with distinct functional attributes.

In the evening, dinner at an oyster restaurant with the Twitterati, and my first taste of these opinion-splitting molluscs (verdict: yup, with some hot sauce and lemon juice).
Friday 16th June
The highlight of the day was interviewing George Daley and Jayaraj Rajagopal for Development. Both are residents of Boston who come from clinical backgrounds but are in love with research, and are truly optimistic and excited about the wave of clinical translation of research into treatments. Both are also wonderful people who were generous in conversation with me; watch out for the final interviews in the coming months.
In a stimulating ‘Tissue Regeneration and Homeostasis’ session, Ben Simons began by apologising for being a physicist (though later he proudly showed us some of the first wet data from his lab). Simons uses mathematical approaches to understand developmental problems, and his talk focussed on the statistics of proliferation versus termination in mammary epithelial stem cells during pubertal morphogenesis. I’ve previously talked to Bill Harris about how Simons’ approach helped him understand the retina, and like the idea that sometimes we can be too burdened by knowledge, so a clear eye from a different field is needed to look at a problem. Given the number of collaborations Simons is involved in, his eye is in high demand.
In the evening I took the metro up to Cambridge though it was too rainy really to see much of it, and ate at a Tex Mex place with an old friend from old Cambridge (accompanied this time by Margaritas).

Saturday 17th June
I’d never been so moved at a scientific meeting – in the ‘Road to the Clinic’ session, Michele De Luca, well known for his promotion of responsible stem cell treatments and efforts to shut down charlatans, told us about epidermolysis bullosa, a terrible disease caused by a defective link between the dermis and the epidermis that can lead to skin blistering at even the slightest touch. De Luca described a patient with a severe genetic form of the disease, a child refugee from Syria who on his arrival to Germany had lost much of his epidermis to an infection and was in effect skinless, and placed in an induced coma for the pain it caused. De Luca’s solution was to grow skin from gene-corrected epidermal stem cells, and then graft this skin onto the child’s body (these were really huge grafts). And a couple of years on it seems to have worked – essentially the entire body is covered by transgenic skin that is now functioning like normal skin (faring better, indeed, than skin grafts used to treat burns, as the disease does not affect the underlying dermis like burns do). He ended his talk with a picture of the boy – standing slightly awkwardly in an ill-fitting suit in a hospital corridor, the most heart-wrenching contrast to the pre-treatment images De Luca had shown before. A kid in a coma to a kid standing and smiling.
An uplifting, miraculous story such as this shows us the potential of stem cells and gene therapy for treatment. As Megan Munsie then described, shadowing these remarkable advances is the growth of unregulated stem cell treatments. I remember a few years ago hearing that Texas Governor and GOP presidential candidate Rick Perry had had stem cells injected for back pain, and since we’ve seen an explosion in the number of clinics in the US and beyond offering stem cell treatments for just about anything (these are often covered by Paul Knoepfler’s excellent blog, the Niche). Not only might such treatments be expensive, and useless, they could of course also be dangerous. Munsie interviewed a number of patients who had sought such treatment – even if they understood the risk, for some it was really about the hope that it gave them, however small. Many of the patients (all based in Australia) felt let down by their own health care practitioners, which suggests we could do more to engage with patients who see no other way out than unregulated treatments.
A brief trip to see the tall ships that had gathered in the harbour, and that was that – back on the night flight to Heathrow, with the image of de Luca’s patient, smiling awkwardly, still in my head.
 (2 votes)
(2 votes)
 Loading...
Loading...
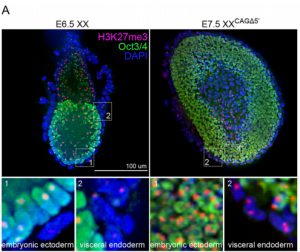 The 5′ region of Xist RNA contains a conserved element termed the A-repeat that is required for the silencing function of Xist in embryonic stem cells, but how this region functions during development is unclear. Now, Takashi Sado and co-workers explore this by introducing into mice a mutated Xist allele that produces Xist RNA lacking the A-repeat region (p. 2784). They first report that imprinted XCI is compromised upon paternal transmission of this allele. The authors further show that the mutant form of Xist is able to coat the X chromosome but fails to silence it in embryonic and extraembryonic tissues. Surprisingly, however, mutant Xist RNA is still able to silence a subset of genes in the trophoblast. Finally, the authors reveal that the failure of imprinted XCI has a more significant impact on genome-wide gene expression than expected; changes in the expression of both X-linked and autosomal genes are observed. Together these findings provide new insights into Xist-mediated gene silencing but also raise the intriguing possibility that dosage compensation regulates X-linked genes as well as gene expression more globally.
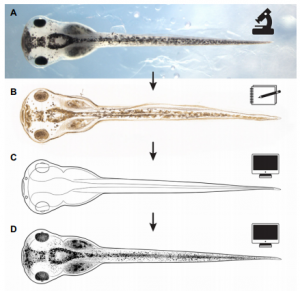
The 5′ region of Xist RNA contains a conserved element termed the A-repeat that is required for the silencing function of Xist in embryonic stem cells, but how this region functions during development is unclear. Now, Takashi Sado and co-workers explore this by introducing into mice a mutated Xist allele that produces Xist RNA lacking the A-repeat region (p. 2784). They first report that imprinted XCI is compromised upon paternal transmission of this allele. The authors further show that the mutant form of Xist is able to coat the X chromosome but fails to silence it in embryonic and extraembryonic tissues. Surprisingly, however, mutant Xist RNA is still able to silence a subset of genes in the trophoblast. Finally, the authors reveal that the failure of imprinted XCI has a more significant impact on genome-wide gene expression than expected; changes in the expression of both X-linked and autosomal genes are observed. Together these findings provide new insights into Xist-mediated gene silencing but also raise the intriguing possibility that dosage compensation regulates X-linked genes as well as gene expression more globally.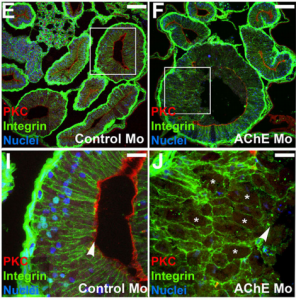 Now, on p. 2764, Nanette Nascone-Yoder and colleagues reveal that AChE plays an essential non-classical role in Xenopus gut morphogenesis. By exposing tailbud stage Xenopus embryos to AChE inhibitors, or by injecting embryos with morpholinos to knock down AChE in the intestinal endoderm, they show that AChE is required for proper intestinal morphogenesis; in the absence of AChE function, intestines are short/malrotated and exhibit a disorganised epithelium. This function of AChE , they report, is independent of its cholinesterase activity. Further analyses demonstrate that AChE is required for endoderm cell rearrangement and polarisation – events that drive gut lengthening and morphogenesis – as well as endoderm cell differentiation. Finally, the researchers demonstrate that AChE regulates cell-substrate but not cell-cell adhesion. Overall, these results provide direct in vivo evidence for a morphogenetic function for AChE in non-neuronal tissues and suggest that AChE may function in other aspects of development and physiology, a find that has important implications given the widespread use of cholinesterase inhibitors in the treatment of human diseases.
Now, on p. 2764, Nanette Nascone-Yoder and colleagues reveal that AChE plays an essential non-classical role in Xenopus gut morphogenesis. By exposing tailbud stage Xenopus embryos to AChE inhibitors, or by injecting embryos with morpholinos to knock down AChE in the intestinal endoderm, they show that AChE is required for proper intestinal morphogenesis; in the absence of AChE function, intestines are short/malrotated and exhibit a disorganised epithelium. This function of AChE , they report, is independent of its cholinesterase activity. Further analyses demonstrate that AChE is required for endoderm cell rearrangement and polarisation – events that drive gut lengthening and morphogenesis – as well as endoderm cell differentiation. Finally, the researchers demonstrate that AChE regulates cell-substrate but not cell-cell adhesion. Overall, these results provide direct in vivo evidence for a morphogenetic function for AChE in non-neuronal tissues and suggest that AChE may function in other aspects of development and physiology, a find that has important implications given the widespread use of cholinesterase inhibitors in the treatment of human diseases.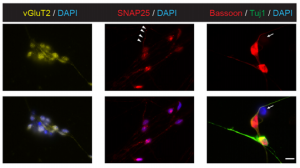 On p. 2748, Ge Guo, Austin Smith and colleagues provide a simple and efficient method for resetting human pluripotency based on transient inhibition of histone deacetylases (HDACs) with chemical inhibitors. HDAC inhibition leads to increased expression of naïve markers in a variety of different hPSC lines, and cells can be expanded in naïve culture conditions without requiring feeders. Chemically reset cells show a marked transcriptional difference to primed hPSCs and a similarity to epiblast cells of the preimplantation inner cell mass. Reset cells undergo global reduction in DNA methylation, have two active X chromosomes, and can differentiate into multiple lineages. This work provides a protocol for efficient resetting of hPSC pluripotency, and a transcription and methylation resource for further interrogation of the human naïve state.
On p. 2748, Ge Guo, Austin Smith and colleagues provide a simple and efficient method for resetting human pluripotency based on transient inhibition of histone deacetylases (HDACs) with chemical inhibitors. HDAC inhibition leads to increased expression of naïve markers in a variety of different hPSC lines, and cells can be expanded in naïve culture conditions without requiring feeders. Chemically reset cells show a marked transcriptional difference to primed hPSCs and a similarity to epiblast cells of the preimplantation inner cell mass. Reset cells undergo global reduction in DNA methylation, have two active X chromosomes, and can differentiate into multiple lineages. This work provides a protocol for efficient resetting of hPSC pluripotency, and a transcription and methylation resource for further interrogation of the human naïve state. On p. 2771, Emily Anne Bates and colleagues describe a role for Irk2, an inwardly rectifying potassium channel in Drosophilaorthologous to Kir2.1 in vertebrates, in regulating the release of the BMP family morphogen Dpp during development of the wing epithelium. Building on their previous finding that a dominant-negative Irk2 reduces Dpp signalling and causes wing defects, they now show that human KIR2.1 can substitute for inhibited DrosophilaIrk2, and that Irk2’s role is restricted to the cells that produce, rather than just transduce, the Dpp signal. Surprisingly, inhibiting Irk2 broadens the distribution of Dpp in the wing, but also alters the dynamics of Dpp release from cells, suggesting that Irk2 controls the timing of Dpp secretion. Irk2 inhibition reduces the amplitude of calcium spikes in wing cells, and depolarising the membrane with extracellular potassium leads to an overall increased release of Dpp, implicating Irk2’s regulation of membrane potential in Dpp release. Together, these results suggest that ion channels influence tissue morphogenesis by regulating the release of morphogens from the membrane.
On p. 2771, Emily Anne Bates and colleagues describe a role for Irk2, an inwardly rectifying potassium channel in Drosophilaorthologous to Kir2.1 in vertebrates, in regulating the release of the BMP family morphogen Dpp during development of the wing epithelium. Building on their previous finding that a dominant-negative Irk2 reduces Dpp signalling and causes wing defects, they now show that human KIR2.1 can substitute for inhibited DrosophilaIrk2, and that Irk2’s role is restricted to the cells that produce, rather than just transduce, the Dpp signal. Surprisingly, inhibiting Irk2 broadens the distribution of Dpp in the wing, but also alters the dynamics of Dpp release from cells, suggesting that Irk2 controls the timing of Dpp secretion. Irk2 inhibition reduces the amplitude of calcium spikes in wing cells, and depolarising the membrane with extracellular potassium leads to an overall increased release of Dpp, implicating Irk2’s regulation of membrane potential in Dpp release. Together, these results suggest that ion channels influence tissue morphogenesis by regulating the release of morphogens from the membrane.


 (No Ratings Yet)
(No Ratings Yet) (1 votes)
(1 votes) (4 votes)
(4 votes)





