In Development this week (Vol. 142, Issue 7)
Posted by Seema Grewal, on 24 March 2015
Here are the highlights from the current issue of Development:
ActivinA-ting spiny neuron production from hPSCs
 The medium-sized spiny neurons, the main projection neurons of the striatum, are generated in the lateral ganglionic eminence (LGE) and degenerate in the early stages of Huntington’s disease (HD) – for which no pharmacological treatment is yet available. Hence, an efficient way to derive striatal neurons is crucial for disease modelling, drug development and cell-replacement therapy. Striatal neurons have previously been generated from human pluripotent stem cell (hPSC)-derived neural progenitors treated with sonic hedgehog (SHH), or SHH plus Wnt pathway inhibition. Now, Meng Li and co-workers (p. 1375) report a more robust and efficient way to generate functional striatal neurons from hPSCs. They show that activin A induces LGE characteristics in hPSC-derived neural progenitors. This is independent of SHH but requires CTIP2, a transcription factor required for striatal neuron development. Furthermore, the activin-patterned neural precursors efficiently generate functional DARPP32+ GABAergic striatal neurons in vitro, and acquire striatal spiny neuron properties without overgrowth or teratoma formation upon engraftment in a rat HD model. Altogether, these findings uncover a novel role for activin A in striatal projection neuron specification and establish a robust protocol for deriving these neurons.
The medium-sized spiny neurons, the main projection neurons of the striatum, are generated in the lateral ganglionic eminence (LGE) and degenerate in the early stages of Huntington’s disease (HD) – for which no pharmacological treatment is yet available. Hence, an efficient way to derive striatal neurons is crucial for disease modelling, drug development and cell-replacement therapy. Striatal neurons have previously been generated from human pluripotent stem cell (hPSC)-derived neural progenitors treated with sonic hedgehog (SHH), or SHH plus Wnt pathway inhibition. Now, Meng Li and co-workers (p. 1375) report a more robust and efficient way to generate functional striatal neurons from hPSCs. They show that activin A induces LGE characteristics in hPSC-derived neural progenitors. This is independent of SHH but requires CTIP2, a transcription factor required for striatal neuron development. Furthermore, the activin-patterned neural precursors efficiently generate functional DARPP32+ GABAergic striatal neurons in vitro, and acquire striatal spiny neuron properties without overgrowth or teratoma formation upon engraftment in a rat HD model. Altogether, these findings uncover a novel role for activin A in striatal projection neuron specification and establish a robust protocol for deriving these neurons.
Cofilin the gap in neural tube closure
 Neural tube closure occurs through highly orchestrated cell shape changes mediated by actin dynamics. Its failure results in some of the most common and severe human congenital malformations. Cofilin 1, an actin-depolymerising protein, is known to be involved in neural tube closure but its precise functions had not been elucidated. In this study (p. 1305), Joaquim Grego-Bessa and colleagues show that the absence of cofilin 1 in mouse leads to defective neural tube closure, reduced cell number, altered cell shape and cell cycle kinetics. The protein is enriched at both apical and basal domains of the neuroepithelium but, intriguingly, has opposing activities on either side of the cell. Apically, although localisation of the apical polarity complexes remains unchanged, phosphorylation of myosin light chain is impaired in cofilin 1 mutants. By contrast, basally, the absence of cofilin 1 leads to an accumulation of actin and phosphorylated myosin light chain, ectopic tight junction-like structures and disruption of the basement membrane and hence of epithelial organisation. Altogether, these results shed light on the cellular mechanisms of neural tube closure and reveal a dual role for cofilin that is presumably dependent on the intracellular context.
Neural tube closure occurs through highly orchestrated cell shape changes mediated by actin dynamics. Its failure results in some of the most common and severe human congenital malformations. Cofilin 1, an actin-depolymerising protein, is known to be involved in neural tube closure but its precise functions had not been elucidated. In this study (p. 1305), Joaquim Grego-Bessa and colleagues show that the absence of cofilin 1 in mouse leads to defective neural tube closure, reduced cell number, altered cell shape and cell cycle kinetics. The protein is enriched at both apical and basal domains of the neuroepithelium but, intriguingly, has opposing activities on either side of the cell. Apically, although localisation of the apical polarity complexes remains unchanged, phosphorylation of myosin light chain is impaired in cofilin 1 mutants. By contrast, basally, the absence of cofilin 1 leads to an accumulation of actin and phosphorylated myosin light chain, ectopic tight junction-like structures and disruption of the basement membrane and hence of epithelial organisation. Altogether, these results shed light on the cellular mechanisms of neural tube closure and reveal a dual role for cofilin that is presumably dependent on the intracellular context.
Preserving progenitor pools in the kidney: a balancing act
The nephrons are the filtration units of the kidney that excrete toxins, balance salt and water content in the blood and regulate blood pressure. Their number is determined during kidney development by the size of the nephron progenitor cell (NPC) pool, which exhausts in early postnatal life in mouse. Understanding the mechanisms that regulate the balance between NPC self-renewal and differentiation is a crucial endeavour. In this issue, two papers provide insights into the molecular cues controlling NPC self-renewal.
 On p. 1228, Zubaida Saifudeen and colleagues report that the specific deletion of p53 in mouse NPCs leads to hypoplastic kidneys, reduced nephron number and elevated blood pressure. p53 is classically associated with restraining proliferation, but the observed phenotype suggests a positive role for p53 in progenitor renewal: in mutants, NPC proliferation is reduced while senescence, apoptosis and the levels of known regulators of NPC survival remain unchanged. Furthermore, using functional genomics, the authors find that p53 regulates factors involved in cell-matrix interactions and metabolism. They then show that mutants display aberrant ATP and reactive oxygen species levels in NPCs. Altogether, these results uncover an unexpected contribution of p53 to NPC self-renewal capacity, energy metabolism and niche architecture.
On p. 1228, Zubaida Saifudeen and colleagues report that the specific deletion of p53 in mouse NPCs leads to hypoplastic kidneys, reduced nephron number and elevated blood pressure. p53 is classically associated with restraining proliferation, but the observed phenotype suggests a positive role for p53 in progenitor renewal: in mutants, NPC proliferation is reduced while senescence, apoptosis and the levels of known regulators of NPC survival remain unchanged. Furthermore, using functional genomics, the authors find that p53 regulates factors involved in cell-matrix interactions and metabolism. They then show that mutants display aberrant ATP and reactive oxygen species levels in NPCs. Altogether, these results uncover an unexpected contribution of p53 to NPC self-renewal capacity, energy metabolism and niche architecture. In the second study (p. 1254), Martin Kann and co-workers identify growth arrest-specific 1 (Gas1) as a direct target of Wilms’ tumor suppressor protein 1 (WT1), a transcription factor required for NPC self-renewal and differentiation. Phenotypically, the loss of GAS1 is similar to p53 depletion, with mutant mice displaying hypoplastic kidneys and decreased nephron numbers, stemming from reduced NPC proliferation. The authors further analyse the mechanism by which GAS1 acts in NPCs, finding that it modulates the response to fibroblast growth factor (FGF) signalling, a known regulator of NPC growth and proliferation, by specifically promoting the AKT pathway branch downstream of receptor activation. This study therefore links WT1 to FGF-mediated regulation of NPC proliferation, providing additional insights into the mechanisms by which this key transcription factor functions.
In the second study (p. 1254), Martin Kann and co-workers identify growth arrest-specific 1 (Gas1) as a direct target of Wilms’ tumor suppressor protein 1 (WT1), a transcription factor required for NPC self-renewal and differentiation. Phenotypically, the loss of GAS1 is similar to p53 depletion, with mutant mice displaying hypoplastic kidneys and decreased nephron numbers, stemming from reduced NPC proliferation. The authors further analyse the mechanism by which GAS1 acts in NPCs, finding that it modulates the response to fibroblast growth factor (FGF) signalling, a known regulator of NPC growth and proliferation, by specifically promoting the AKT pathway branch downstream of receptor activation. This study therefore links WT1 to FGF-mediated regulation of NPC proliferation, providing additional insights into the mechanisms by which this key transcription factor functions.
PLUS…
Positional information and reaction-diffusion: two big ideas in developmental biology combine
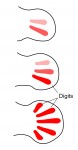 The two most influential ideas in the field of pattern formation are those of Alan Turing’s ‘reaction-diffusion’ and Lewis Wolpert’s ‘positional information’. Much has been written about these two concepts but some confusion still remains, in particular about the relationship between them. Here, Jeremy Green and James Sharpe address this relationship and propose a scheme of three distinct ways in which these two ideas work together to shape biological form. See their Hypothesis article on p. 1203
The two most influential ideas in the field of pattern formation are those of Alan Turing’s ‘reaction-diffusion’ and Lewis Wolpert’s ‘positional information’. Much has been written about these two concepts but some confusion still remains, in particular about the relationship between them. Here, Jeremy Green and James Sharpe address this relationship and propose a scheme of three distinct ways in which these two ideas work together to shape biological form. See their Hypothesis article on p. 1203
Cellular and molecular insights into Hox protein action
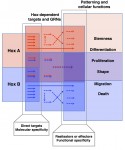 Hox genes encode homeodomain transcription factors that control morphogenesis and have established functions in development and evolution.Here, Yacine Graba and colleagues discuss the molecular and cellular mechanisms underlying the diverse and context-dependent functions of Hox transcription factors during morphogenesis and organogenesis. See the Review article on p. 1212
Hox genes encode homeodomain transcription factors that control morphogenesis and have established functions in development and evolution.Here, Yacine Graba and colleagues discuss the molecular and cellular mechanisms underlying the diverse and context-dependent functions of Hox transcription factors during morphogenesis and organogenesis. See the Review article on p. 1212


 (1 votes)
(1 votes)

 (No Ratings Yet)
(No Ratings Yet)
 Dmrt1 and its related genes play a key role in sex determination in a broad range of metazoan species. However, Dmrt1 has become dispensable for testis determination in mammals, and this function is instead carried out by Sry, which is a newly evolved gene found on the Y chromosome. Now, Peter Koopman and colleagues show that, even though its function is not normally required, Dmrt1 is able to drive female-to-male sex reversal in mice (p.
Dmrt1 and its related genes play a key role in sex determination in a broad range of metazoan species. However, Dmrt1 has become dispensable for testis determination in mammals, and this function is instead carried out by Sry, which is a newly evolved gene found on the Y chromosome. Now, Peter Koopman and colleagues show that, even though its function is not normally required, Dmrt1 is able to drive female-to-male sex reversal in mice (p.  In plants, stem cell proliferation is negatively regulated by the receptor kinase CLAVATA1 (CLV1) and its peptide ligand CLAVATA3 (CLV3). Previous studies have suggested that CLV1 acts redundantly with other receptor kinases, such as BAM1, 2 and 3, but the molecular mechanisms underpinning this redundancy have been unclear. Now, Elliot Meyerowitz and co-workers interrogate the role of CLV1-CLV3 signalling in the Arabidopsis shoot apical meristem (p.
In plants, stem cell proliferation is negatively regulated by the receptor kinase CLAVATA1 (CLV1) and its peptide ligand CLAVATA3 (CLV3). Previous studies have suggested that CLV1 acts redundantly with other receptor kinases, such as BAM1, 2 and 3, but the molecular mechanisms underpinning this redundancy have been unclear. Now, Elliot Meyerowitz and co-workers interrogate the role of CLV1-CLV3 signalling in the Arabidopsis shoot apical meristem (p.  Hematopoietic stem cells (HSCs) give rise to all cells of the adult blood system, and understanding how these cells first arise during embryogenesis is important for developing regenerative medicine-based strategies for producing HSCs in vitro. Here, David Traver and colleagues demonstrate that Gata2b acts as an early regulator of zebrafish hematopoietic precursors (p.
Hematopoietic stem cells (HSCs) give rise to all cells of the adult blood system, and understanding how these cells first arise during embryogenesis is important for developing regenerative medicine-based strategies for producing HSCs in vitro. Here, David Traver and colleagues demonstrate that Gata2b acts as an early regulator of zebrafish hematopoietic precursors (p. Adherens junctions (AJs), which are specialised E-cadherin-based cell contacts, are continuously remodelled during tissue morphogenesis, as cells change shape and position. The accumulation of Bazooka (Baz), the Drosophila PAR3 homologue, is thought to specify where new E-cadherin complexes are deposited during AJ remodelling, but what regulates Baz localisation? Here, Alexandre Djiane and colleagues show that the scaffold protein Magi regulates Baz localization and hence AJ remodelling inDrosophila eye epithelial cells (p.
Adherens junctions (AJs), which are specialised E-cadherin-based cell contacts, are continuously remodelled during tissue morphogenesis, as cells change shape and position. The accumulation of Bazooka (Baz), the Drosophila PAR3 homologue, is thought to specify where new E-cadherin complexes are deposited during AJ remodelling, but what regulates Baz localisation? Here, Alexandre Djiane and colleagues show that the scaffold protein Magi regulates Baz localization and hence AJ remodelling inDrosophila eye epithelial cells (p.  Skeletal stem cells (SSCs) reside in the postnatal bone marrow and give rise to cartilage, bone, hematopoiesis-supportive stroma and marrow adipocytes. Here, Paolo Bianco and Pamela Robey discuss the biology of SSCs in the context of the development and postnatal physiology of skeletal lineages, to which their use in medicine must remain anchored. See the Development at a Glance poster article on p.
Skeletal stem cells (SSCs) reside in the postnatal bone marrow and give rise to cartilage, bone, hematopoiesis-supportive stroma and marrow adipocytes. Here, Paolo Bianco and Pamela Robey discuss the biology of SSCs in the context of the development and postnatal physiology of skeletal lineages, to which their use in medicine must remain anchored. See the Development at a Glance poster article on p.  The mammary gland provides an excellent model for studying ‘stem/progenitor’ cells, which – in this context – allow for the repeated expansion and renewal of the gland during adult life. Here, Mina Bissell and colleagues discuss the various cell types that constitute the mammary gland, highlighting how they arise and differentiate, and how the microenvironment influences their development. See the Review on p.
The mammary gland provides an excellent model for studying ‘stem/progenitor’ cells, which – in this context – allow for the repeated expansion and renewal of the gland during adult life. Here, Mina Bissell and colleagues discuss the various cell types that constitute the mammary gland, highlighting how they arise and differentiate, and how the microenvironment influences their development. See the Review on p.  Today, a few button clicks gives access to vast troves of knowledge, and a few dollars buys technologies that even well-funded labs could not get a few decades ago. So, it should be much easier for amateurs and hobbyists to do scientific research now than it was in Darwin’s era. The
Today, a few button clicks gives access to vast troves of knowledge, and a few dollars buys technologies that even well-funded labs could not get a few decades ago. So, it should be much easier for amateurs and hobbyists to do scientific research now than it was in Darwin’s era. The  Facilitating amateur-professional interactions would also improve public understanding of science. This is especially important in areas that intersect with developmental biology; voters are routinely called upon to make decisions related to stem cells, genetics, or evolution. The premise of every graduate school is that the best way to learn how science works is to do it, yet there are few opportunities for adult non-scientists to experience the creative and intellectual side of research. The success of the citizen science movement shows that many people are interested in participating in science. However, most citizen science projects are designed to get a large number of volunteers to do a defined task, rather than to help non-scientists plan research and interpret results. This leaves a big gulf between non-scientists and professionals.
Facilitating amateur-professional interactions would also improve public understanding of science. This is especially important in areas that intersect with developmental biology; voters are routinely called upon to make decisions related to stem cells, genetics, or evolution. The premise of every graduate school is that the best way to learn how science works is to do it, yet there are few opportunities for adult non-scientists to experience the creative and intellectual side of research. The success of the citizen science movement shows that many people are interested in participating in science. However, most citizen science projects are designed to get a large number of volunteers to do a defined task, rather than to help non-scientists plan research and interpret results. This leaves a big gulf between non-scientists and professionals. Developmental biology is ripe for this. Although a lot of developmental biology depends on expensive reagents and high-tech equipment, plenty of high-value, low-tech research remains to be done. Two of my all-time favorite papers (1, 2) used nothing more than glass needles and intelligence to identify, and partially solve, a paradox of ctenophore development: when an embryo is split in two, each half develops into half an embryo; yet the adults can regenerate an entire half of their body. The authors documented ontogenetic transitions in these phenomena, and then deciphered the roles of specific cell lineages in patterning and regeneration. In my own work, I’ve found that the most useful biomechanical techniques for working with embryos are things like micropipette aspiration, which would be easily accessible to amateur microscopists (it was developed in the 1950’s (3)). There are myriad questions in developmental biology that could be investigated with low-budget techniques.
Developmental biology is ripe for this. Although a lot of developmental biology depends on expensive reagents and high-tech equipment, plenty of high-value, low-tech research remains to be done. Two of my all-time favorite papers (1, 2) used nothing more than glass needles and intelligence to identify, and partially solve, a paradox of ctenophore development: when an embryo is split in two, each half develops into half an embryo; yet the adults can regenerate an entire half of their body. The authors documented ontogenetic transitions in these phenomena, and then deciphered the roles of specific cell lineages in patterning and regeneration. In my own work, I’ve found that the most useful biomechanical techniques for working with embryos are things like micropipette aspiration, which would be easily accessible to amateur microscopists (it was developed in the 1950’s (3)). There are myriad questions in developmental biology that could be investigated with low-budget techniques.

