February in preprints
Posted by the Node, on 31 March 2026
Welcome to our monthly trawl for developmental and stem cell biology (and related) preprints.
The preprints this month are hosted on bioRxiv – use these links to get to the section you want.
- Patterning & signalling
- Morphogenesis & mechanics
- Genes & genomes
- Stem cells, regeneration & disease modelling
- Plant development
- Evo-devo
Developmental biology
| Patterning & signalling
Lineage domains and cytoskeletal cables organize a cellular square grid in a crustacean
Beatrice L. Steinert, Leo Blondel, Chandrashekar Kuyyamudi, Evangelia Stamataki, Anastasios Pavlopoulos, Cassandra G. Extavour
An optogenetic toolkit for robust activation of FGF, BMP, & Nodal signaling in zebrafish
Leanne E. Iannucci, Velanganni Selvaraj Maria Thomas, William K. Anderson, Micaela R. Murphy, Caitlin E.T. Donahue, Catherine E. Campbell, Matthew T. Monaghan, Allison J. Saul, Katherine W. Rogers
Directed conversion of porcine extended pluripotent stem cells into trophoblast-like stem cells through modulation of conserved TGF-β and ERK signaling pathways
Chi-Hun Park, Young-Hee Jeoung, JiTao Wang, Bhanu P. Telugu
Synthetic germ granules reveal a direct role of Vasa/DDX4 in RNA localization and translational activation
Ruoyu Chen, Henoc Zinga, Jay S. Goodman, Ruth Lehmann
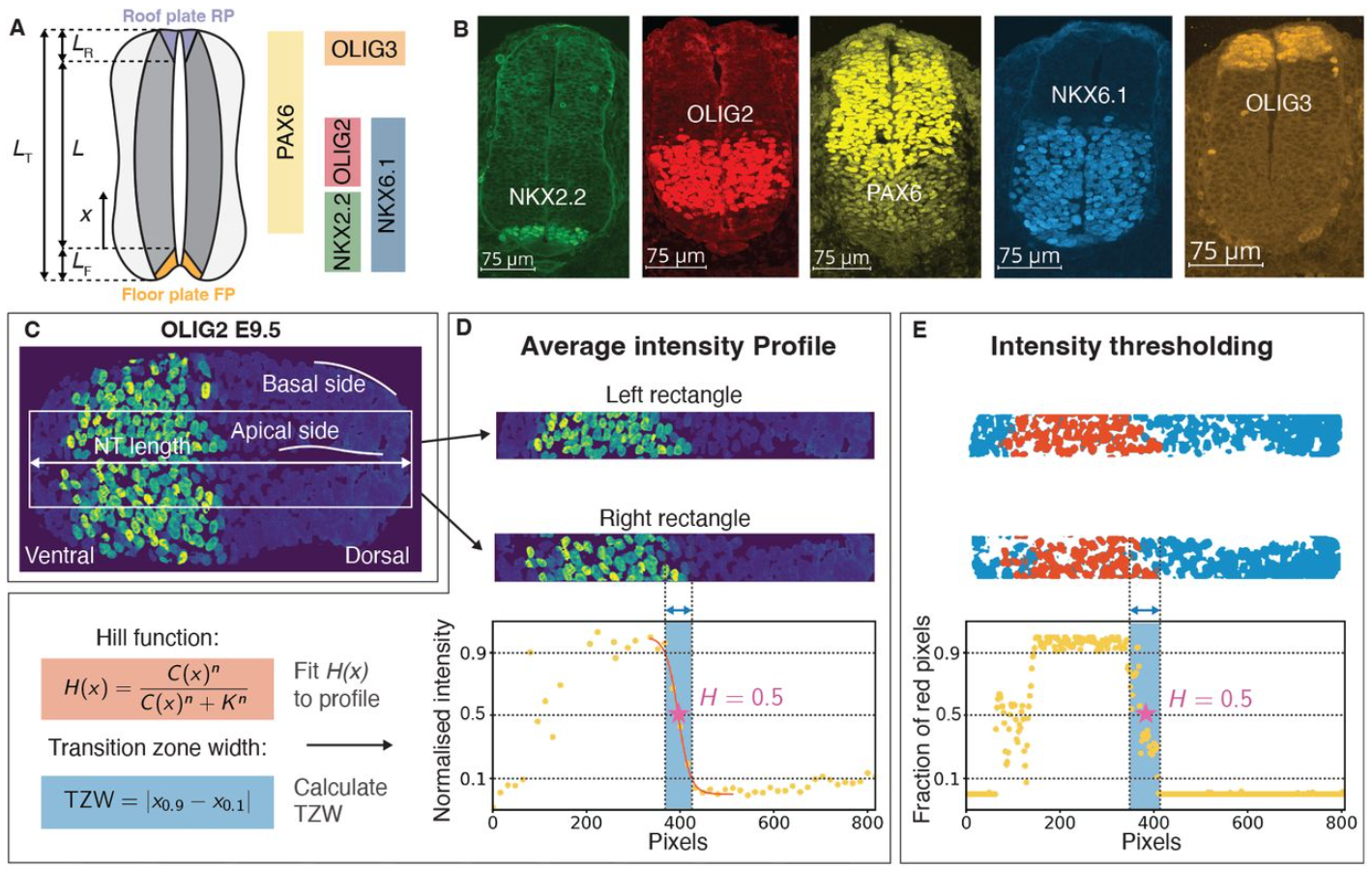
Determinants of the Transition Zone Width of Morphogen Readouts
Jan A. Adelmann, Roman Vetter, José M. Dias, Johan Ericson, Dagmar Iber
Mechanistic tradeoffs between local and long-range signaling activity in natural and synthetic morphogens
Gavin Schlissel, Anders S. Hansen, Pulin Li
Inflammatory IL-1 signaling remodels epidermal stem cell compartments by suppressing Wnt activity
Hung Manh Phung, Ikuto Nishikawa, Nguyen Thi Kim Nguyen, Aiya K. Yesbolatova, Ahmed M. Hegazy, Tomson Kosasih, Jun Aoi, Satoshi Fukushima, Sho Hiroyasu, Hitoshi Takizawa, Aiko Sada
Cajal-Retzius fate specification is disrupted by constitutive activation of β-Catenin in hem progenitors
Amrita Singh, Arpan Parichha, Debarpita Datta, Mallika Chatterjee, Shubha Tole
Kinetic Control of Out-Of-Equilibrium Dynamics in the RhoA Signaling Cascade Shapes Actomyosin Contractility
Serena Prigent Garcia, Étienne Pinard, Camille N. Plancke, Jing Li, Shashi Kumar Suman, Loan Bourdon, Christelle Gally, Taeyoon Kim, François B. Robin
Dorsal/NF-κB exhibits a dorsal-to-ventral mobility gradient in the Drosophila embryo
Hadel Al Asafen, Natalie M. Clark, Etika Goyal, Sadia Siddika Dima, Hung-Yuan Chen, Thomas Jacobsen, Rosangela Sozzani, Gregory T. Reeves
Position Dependent Feedback Drives Scaling and Robustness of Morphogen Gradients
Lewis Scott Mosby, Zena Hadjivasiliou
TGF-β signaling regulates epithelial permeability in Drosophila ovaries by modulating adhesion independent of actomyosin contractility
Harshath Amal, Thea Jacobs, Max Lohrberg, Stefan Luschnig
Reconstructing signaling histories of single cells via perturbation screens and transfer learning
Nicholas T. Hutchins, Miram Meziane, Claire Lu, Maisam Mitalipova, David Fischer, Pulin Li
Direct cell-to-cell transport of Hedgehog morphogen is aided by the diffusible carrier Shifted/DmWif1
Carlos Jiménez-Jiménez, Gustavo Aguilar, Clara Fernández-Pardo, Markus Affolter, Isabel Guerrero
Arginine Kinase 1 regulates energy homeostasis in Drosophila muscle development
Maria Paula Zappia, Anton Westacott, Hannah Cooke, Rhianna Geary, Libby Travers, Lucia de Castro, Oliver Carty, Maxim V Frolov
Weckle is a molecular switch that diverts Toll signalling from innate immunity towards growth by engaging Yki
Maria Dolores Perez-Sanchez, Guiyi Li, Martin Moncrieffe, Francisca Rojo-Cortés, Karina Malinovska, Emily Sample, Myles Maddick, Marta Moreira, Elizabeth Connolly, Anna Parsons, Roberto Feuda, Nick J. Gay, Alicia Hidalgo
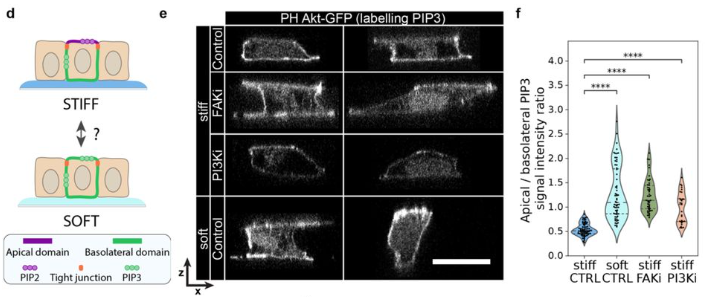
Basement membrane mechanics drives patterned response to developmental signalling
Ana Raffaelli, Tom P.J. Wyatt, Claire S. Simon, Léa M.D. Wenger, Kathy K. Niakan, Ewa K. Paluch, Kevin J. Chalut
A single-cell transcriptomic map of the Xenopus mesonephros reveals conserved nephron patterning across vertebrate kidney forms
Mark E. Corkins, Adrian Romero, MaryAnne A. Achieng, Nils O. Lindström, Rachel K. Miller
Sharp cell-type boundaries emerge from temporal coordination between morphogen signals
Ruiqi Li, Yiqun Jiang, Sarah Platt, Tianchi Xin, Ryan Driskell, Kevin A. Peterson, Sarah Van, Hainan Lam, Shagun Lukkad, Eva-LaRue Barber, Chae Ho Lim, M. Mark Taketo, Yuval Kluger, Peggy Myung
Synthetic reconstitution of planar polarity initiation reveals collective migration as a symmetry-breaking cue
Leah A Wallach, Connor D Thomas, Pulin Li
Epithelial-Mesenchymal Wnt Crosstalk Directs Planar Cell Polarity in the Developing Cochlea
Ippei Kishimoto, Abel P. David, Kevin P. Rose, Balasubramanian Narasimhan, Bradley Efron, Sara E. Billings, Erin L. Su, Wuxing Dong, Taha A. Jan, Ronna Hertzano, Alan G. Cheng
Notch-driven fate asymmetry dictates hair cell behavior via a fate-specific kinase
Emily Atlas, Caleb C. Reagor, Brian Frost, Sapna Krishnakumar, A. J. Hudspeth, Adrian Jacobo
A synNotch-based morphogen detection system reveals sFRP2 enhances Wnt3a signaling
Kosuke Mizuno, Satoshi Toda
| Morphogenesis & mechanics
Tenascin N contributes to spinal motor nerve morphogenesis during development
Charles G. Marcucci, Marieke Jones, Coleman Blanton, Sarah Kucenas
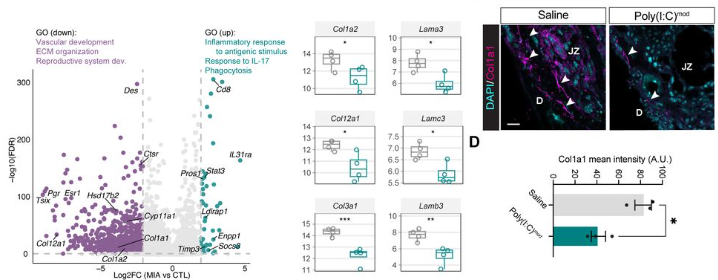
A stress-responsive morphogenetic program of the uterine epithelium safeguards the establishment of early pregnancy
Chihiro Ishizawa, Shizu Aikawa, Yamato Fukui, Xueting He, Ryoko Shimizu-Hirota, Daiki Hiratsuka, Mitsunori Matsuo, Takehiro Hiraoka, Yasushi Hirota
Cell-intrinsic compliance mechanism enables release of tensile stress to prevent tissue rupture
Chun Wai Kwan, Shunta Sakaguchi, Michiko Takeda, Takefumi Kondo, Yu-Chiun Wang
The mechanosensitive protein Zyxin influences Hippo signalling and tissue growth via adherens junctions and basal spot junctions in Drosophila
Harmanjeet Singh, Elliot Brooks, Kyoko Jinnai, Shu Kondo, Samuel A. Manning, Benjamin Kroeger, Kieran F. Harvey
Inverted Assembly of the Lens Within Ocular Organoids Reveals Alternate Paths to Ocular Morphogenesis
Elin Stahl, Miguel Angel Delgado-Toscano, Ishwariya Saravanan, Anastasija Paneva, Joachim Wittbrodt, Lucie Zilova
Long-Range Coupling of Posterior Cell Addition and Anterior Vacuolation Provides Robustness in Notochord Elongation
Carlos Camacho-Macorra, Alberto Ceccarelli, Dillan Saunders, Guillermo Serrano Nájera, Osvaldo Chara, Benjamin Steventon
Pre-cuticle DPY-6 acts as a blueprint for aECM periodic organization in C. elegans
Sophie Mazzoli, Thomas Sonntag, Emma Cadena, Claire Valotteau, Susanna K. Birnbaum, Meera V. Sundaram, Nathalie Pujol
Unified Transcriptome and Mechanics Map of the Intact Mammalian Preimplantation Embryo In Situ
Ehsan Habibi, Anubhav Sinha, Haiqian Yang, Payman Yadollahpour, Yiwei Li, Lani Lee, David A. Wollensak, Zachary D. Chiang, Denny Sakkas, Edward S. Boyden, Ming Guo, Aviv Regev, Fei Chen
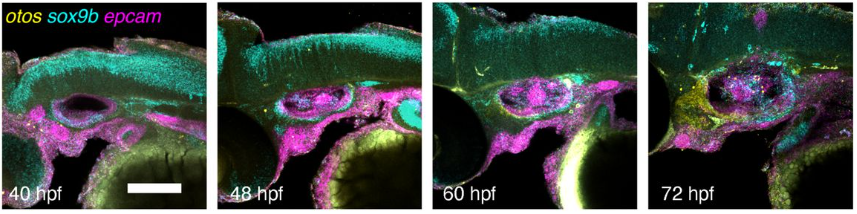
A single-cell transcriptomic atlas of inner ear morphogenesis in zebrafish
Akankshi Munjal, Kalki Kukreja, Samara Williams, Toru Kawanishi, Natasha M. O’Brown, Kana Ishimatsu, Allon Klein, Sean G. Tsung-Megason, Ian A. Swinburne
Kindlin-2-Moesin interaction orchestrates sprouting angiogenesis via modulating endothelial membrane mechanics and VEGF signaling
Lu Wang, Yuxin Fu, Zeyang Yu, Yi Lei, Tianjing Yang, Jiayu Liu, Nina Ma, Yuming Liu, Kunfu Ouyang, Kai Zhang, Junhao Hu, Xi Fang, Ying Shen, Jing Zhou, Xiaohong Wang
Tissue-scale mechanics controls differentiation strategy and dynamics of epithelial multilayering
Clémentine Villeneuve, Somiealo Azote Epse Hassikpezi, Marga Albu, Matthias Rübsam, Leah C. Biggs, Sabrina Vinzens, Kai Kruse, Anubhav Prakash, Peter Zentis, Elizabeth Lawson-Keister, Gautier Follain, Johanna Ivaska, Carien M. Niessen, M. Lisa Manning, Sara A. Wickström
Mechanical memory of confinement pressure governs expansion size in epithelial monolayers
Linn Engström, Simon K Schnyder, Johannes K Ahnlide, Valeriia Grudtsyna, Martijn Gloerich, Pontus Nordenfelt, Amin Doostmohammadi, Vinay Swaminathan
| Genes & genomes
Light-entrained chromatin priming poises rapid metamorphosis in a marine sponge
Huifang Yuan, Oceane Blard, Zac Pujic, Bernard M. Degnan, Sandie M. Degnan
Single-cell multiomics identifies key nodes and cis-regulatory elements of the networks specifying the eye domains in zebrafish
Javier Macho Rendón, Rocío Polvillo, Álvaro Gónzalez-Cid, Jorge Corbacho, Silvia Naranjo, Sofia Manzo, Ana Sousa-Ortega, Ana Fernández-Miñán, Juan Tena, Juan Ramón Martínez-Morales
Reciprocal zebrafish-medaka hybrids reveal maternal control of zygotic genome activation timing
Krista R. Briedis-Gert, Gunnar Schulze, Maria Novatchkova, Karin Panser, Luis Enrique Cabrera Quio, Anja Koller, Yixuan Guo, Bradley R. Cairns, Eivind Valen, Andrea Pauli
A Multi-tissue Transcriptomic-Metabolomic Map Linking Maternal High-Fiber Diet to Reduced Offspring Type 2 Diabetes
Tetsuto Katsura, Oluwagbotemi Omojola, Antwi-Boasiasko Oteng, Peng Jiang, Katherine A. Overmyer, Josh Coon, Amadou Gaye, Huishi Toh
A transcriptional code controlling fluid shear stress-induced gene expression
Lucija Fleisinger, Susann Bruche, Hyewon Lim, Anna Rataj, Helena Rodriguez-Caro, Amaury Genovese, Vinesh Vinayachandran, Svanhild Nornes, Dorota Szumska, Dhruv S Gupta, Indrika Ratnayaka, Kira Chouliaras, Marek Giers, Simon J Conway, Alice Neal, Sophie Payne, Martin A Schwartz, Mukesh K Jain, Brian G Coon, Sarah De Val
Single-cell multiomic approaches define a gradual, spatially-regulated epigenetic and transcriptional transition from embryonic to adult neural stem cells
Beatrix S. Wang, Konstantina Karamboulas, Nareh Tahmasian, Daniel J. Dennis, David R. Kaplan, Freda D. Miller
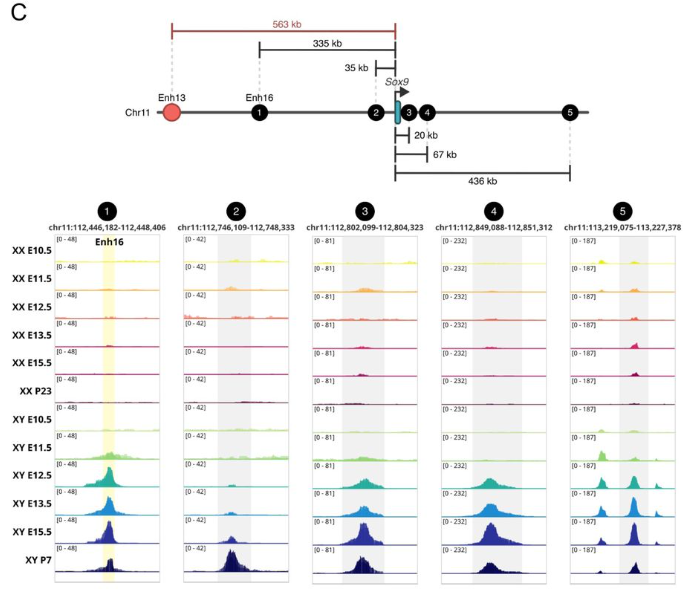
Male fertility is independent of Enh13 control of Sox9 testicular expression
Maor Lubman, Meshi Ridnik, Isabelle Stévant, Yael Kimchi Djanshvili, Elisheva Abberbock, Shelly Ziv Lhermann, Nitzan Gonen
A dual-phase enhancer couples progenitor maintenance and pancreatic lineage stability
Marta Duque, João Amorim, Joana Teixeira, Beatriz Custódio, Mafalda Galhardo, Francisco Camões Magalhães, Joana Marques, Ana Paula Pêgo, José Bessa
Enhancer-mediated metabolic pre-patterning defines trabecular cardiomyocyte identity prior to morphogenesis
Costantino Parisi, Shikha Vashisht, Mohammad Salar Ghasemi Nasab, Kandhadayar Gopalan Srinivasan, Katarzyna Misztal, Marcin Zagorski, Cecilia Winata
ERH elicits cell lineage restriction in mammalian preimplantation development and differentiation from pluripotency via H3K9me3-heterochromatin
Andrew Katznelson, Blake Hernandez, Kylea Tapia, Holly Fahning, Adam Burton, Jingchao Zhang, Maria-Elena Torres-Padilla, Nicolas Plachta, Kenneth S. Zaret, Ryan L. McCarthy
A conserved C. elegans zinc finger-homeodomain protein, ZFH-2, continuously required for structural integrity and function of alimentary tract and gonad
Antoine Sussfeld, Berta Vidal, Surojit Sural, Daniel M. Merritt, G. Robert Aguilar, Yasmin Ramadan, Oliver Hobert
The Nkx2.3–Nr5a1 gene cascade plays a crucial role in spleen-specific vascular architecture and marginal zone formation
Kanako Miyabayashi, Koji Ono, Tetsuya Sato, Ayano Yahagi, Masanori Iseki, Katsuhiko Ishihara, Takami Mori, Miki Inoue, Ryuki Shimada, Kei-ichiro Ishiguro, Tomohiro Ishii, Jongsung Noh, Man Ho Choi, Takashi Baba, Yasuyuki Ohkawa, Emi Kiyokage, Kazunori Toida, Yuichi Shima
A Phospho-Switch for Cell Fate Control
Jin Ming, Xianzhuang Liu, Zexiao Jia, Wei Shi, Jiajun Li, Shikun Wang, Yulin Chen, Shixian Lin, Yu Liang, Peng Guo, Hanqing Zhao, Yuxiang Yao, Ruona Shi, Xiaofei Zhang, Yuanyue Shan, Yu Fu, Bo Wang, Chengchen Zhao, Duanqing Pei
Mitotic bookmarking by Prox1 preserves mammalian neuronal lineage identity memory via promoting timely H3K27me3 restoration
Chouin Wong, Jie Liu, Haoran Yang, Haotian Li, Xiaoqi Luo, Tingyi Li, Zili Chen, Jingyi Chu, Yuying Shen, Shuai Long, Yong Zhang, Yan Song
A Mediator-dependent hypertranscriptional program governs neural stem cell fate decisions in vivo
Tiago Baptista, Daniela Lopes, Ana Rita Rebelo, Catarina CF Homem
Single-Cell Atlas of Transcription and Chromatin States Reveals Regulatory Programs in the Human Brain
Yang Xie, Lei Chang, Guojie Zhong, Jonathan A. Rink, Tatiana Báez-Becerra, Ethan Armand, Wubin Ding, Kai Li, Eric Bonne, Audrey Lie, Hannah S Indralingam, Keyi Dong, Timothy Loe, Bohan Huang, Zhaoning Wang, Ariana S. Barcoma, Jackson K. Willier, Kyle W. Knutson, Jiayi Liu, Silvia Cho, Stella Cao, Kaitlyn G. Russo, Carissa K. Young, Jessica Arzavala, Yareli Sanchez, Aleksandra Bikkina, Natalie Schenker-Ahmed, Colin Kern, Zoey Zhao, Amit Klein, Jesus Flores, Chu-Yi Tai, Jacqueline Olness, Alexander Monell, Siavash Moghadami, Cesar Barragan, Chumo Chen, William Owens, Carolyn O’Connor, Michelle Liem, Mikayla V. Marrin, Cynthia Rose, Shane N. Alt, Nora Emerson, Julia Osteen, Jacinta Lucero, Daofeng Li, Rebecca D. Hodge, Ting Wang, C. Dirk Keene, Xiangming Xu, Quan Zhu, Joseph R. Ecker, M. Margarita Behrens, Bing Ren
DNA supercoiling links transcription and chromatin architecture during human stem cell differentiation
Consuelo Perez, Pierre Murat, Andrew Zeller, Kim C. Liu, Alastair Crisp, Julian E. Sale
Matched single-cell chromatin, transcriptome, and surface marker profiling captures in vivo epigenomic reprogramming during basal-to-luminal transition in the mammary gland
Anna Schwager, Eve Moutaux, Adeline Durand, Alexandra Van Keymeulen, Amélie Viaene, Mélanie Miranda, Louisa Hadj Abed, Simon Besson-Girard, Marion Lambault, Délia Dupré, Grégoire Jouault, Mélissa Saichi, Juliette Bertorello, Simon Dumas, Mathias Schwartz, Marthe Laisné, Justine Marsolier, Manuel Guthmann, Lorraine Bonneville, Urvashi Chitnavis, Déborah Bourc’his, Elisabetta Marangoni, Nicolas Servant, Cédric Blanpain, Leïla Perié, Céline Vallot
| Stem cells, regeneration & disease modelling
Neonatal diethylstilbestrol exposure disrupts uterine epithelial apical-basal polarity and partial EMT state
Rachel E. Bainbridge, Wendy N. Jefferson, Tianyuan Wang, Sara A. Grimm, Carmen J. Williams
PIK3CA-related overgrowth spectrum (PROS) zebrafish models reveal pan-lineage developmental dysregulation
Hannah Brunsdon, Nuoya Wang, Micha Sam Brickman Raredon, Ralitsa R Madsen, Robert K Semple, E Elizabeth Patton
Maternal-fetal immune conflict contributes to male-specific impairments in a mouse model of neurodevelopmental disorders
Irene Sanchez-Martin, Bharti Kukreja, Paige Henderson, Qianyu Lin, Daniel DiMartino, Valerie Bagan, Justin Park, Brian T. Kalish, Lucas Cheadle
PRDM16 is necessary for sensory neuronal development in the Trigeminal Ganglion
Fahmida Raha, Qiman Gao, Lomeli C. Shull, Kristin B. Artinger
Leucettinib-21 decreases dosage effects of DYRK1A in human trisomy 21 iPSC-derived neural cells
Nicole R. West, Mattias F. Lindberg, Julien Dairou, Shawn MacGregor, Sahith Puthireddy, Laurent Meijer, Anita Bhattacharyya
Mutant FGFR3 restricts bone yet expands cortex via ERK-mediated self-repression
Zhuangzhi Zhang, Zhejun Xu, Tongye Fu, Wenhui Zheng, Zizhuo Sha, Chuannan Yang, Feihong Yang, Jialin Li, Jing Ding, Zhengang Yang
Hypoxia differentially affects coronary vessel formation during heart development
Sophie Payne, Susann Bruche, Dorota Szumska, Alice Neal, Mark D Preston, Sarah De Val
Pancreatic Duct Cells as a Potential Source for Human Islet Neogenesis: Insights from Imaging Mass Cytometry
Rui Liang, Tengli Liu, Lanqiu Zhang, Wenmiao Ma, Huixia Ren, Shusen Wang
Rp-vasa: a bona fide Primordial Germ Cell marker that drives embryonic expression in the Chagas disease vector Rhodnius prolixus
G. Martins, M. Berni, T. Guedes-Silva, D. Bressan, J. Vieira, M. Cardoso, A. Pane, V. Gantz, E. Bier, H.M Araujo
Two axolotl-adapted cell-ablation platforms reveal macrophage-dependent processes essential for spinal-cord and skeletal regeneration
Gabriela Johnson, Andrew Hart, Markus Sujansky, Joel H. Graber, James W. Godwin
The regulatory landscape of optic fissure closure in the vertebrate eye
Brian Ho Ching Chan, Mariya Moosajee, Holly Hardy, James Prendergast, Joe Rainger
Evidence that injury can cause Drosophila gut differentiated, polyploid enterocytes to be recruited as stem cells via paligenosis
Dongkook Park, Robert M. Lawrence, Tyler Jackson, Hongjie Li, Jason C. Mills
The abnormal C-terminus in DVL1 impacts Robinow Syndrome phenotypes
Shruti S. Tophkhane, Gamze Akarsu, Sarah J. Gignac, Katherine Fu, Stephanie Xie, Esther M. Verheyen, Joy M. Richman
HSD17B7 is required for the function of sensory hair cells by regulating cholesterol synthesis
Yuqian Shen, Ziyang Wang, Xun Wang, Fuping Qian, Mingjun Zhong, Xin Wang, Jing Cheng, Dong Liu
Regeneration can take place across Drosophila compartments or segments with different Hox gene expression
Rafael Alejandro Juárez-Uribe, Paloma Martín, Laura Utiel, Blanca L. Arrabal, Marina Blanco, Roberto Yagüe-Serrano, Eduardo Cazalla, Ernesto Sánchez-Herrero
Intellectual disability risk gene RFX4 regulates cortical neurogenesis by restraining neuronal differentiation
Julianna J. Determan, Gareth Chapman, Sydney R. Crump, Faiza Batool, Sofia Malik, Taranjit S. Gujral, William Buchser, Caleb Valentine, Serena Elia, Monica Sentmanat, Xiaoxia Cui, Haley Jetter, Kristen L. Kroll
Brain morphological pattern is associated with the presence, severity, and transition of transdiagnostic psychiatric disorders in preadolescents
Nanyu Kuang, Christopher J Hammond, Betty Jo Salmeron, Xiang Xiao, Danni Wang, Laura Murray, Hong Gu, Tianye Zhai, Hui Zheng, Justine Hill, Maria Scavinicky, Hanbing Lu, Amy Janes, Thomas J Ross, Yihong Yang
4D Single-Cell Spatial Transcriptomics Reveals Dynamic Morphogenetic Gradients and Regenerative Domains in Planarians
Kai Han, Yue Chen, Yao Li, Lidong Guo, Yuxiaofei Wang, Xiawei Liu, Yaru Lin, Zhi Huang, Qun Liu, Wenjie Guo, Rui Zhang, Wandong Zhao, Langchao Liang, Xiaoyu Wei, Li Zhou, Xuebin Mao, Jiaqi Wang, Weijian Wu, Hongwei Pan, Tao Yang, He Zhang, Xiaoshan Su, Shanshan Liu, Wenwei Zhang, Longqi Liu, Søren Tvorup Christensen, Jifeng Fei, Xin Liu, Guangyi Fan, Hanbo Li, Ying Gu, Jian Wang, Huanming Yang, Gang Pei, Xun Xu, An Zeng, Mengyang Xu
Drosophila ryanodine receptor gene triggers functional and developmental muscle properties and could be used to assess the impact of human RYR1 mutations
Monika Zmojdzian, Teresa Jagla, Florian Cherik, Magda Dubinska-Magiera, Marta Migocka-Patrzalek, Malgorzata Daczewska, John Rendu, Krzysztof Jagla, Catherine Sarret
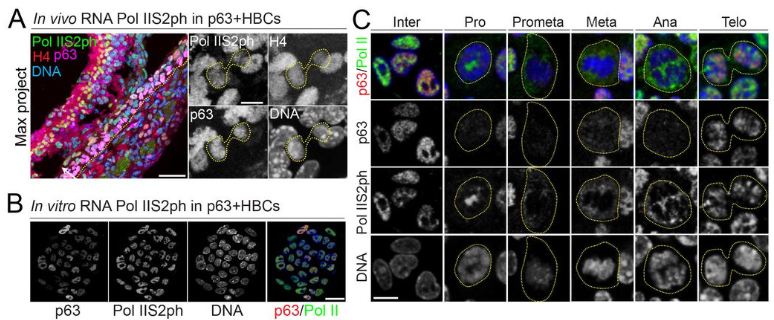
Asymmetric Histone Inheritance Regulates Olfactory Stem Cell Fates During Regeneration
Binbin Ma, Guanghui Yang, Jonathan Yao, Charles Wu, Jean Pinckney Vega, Gabriel Manske, Saher Sue Hammoud, Satrajit Sinha, Abhyudai Singh, Haiqing Zhao, Xin Chen
A knock-in Six2Cre line reveals transient interstitial potential in nephron progenitors
Azadeh Haghighitalab, Fariba Nosrati, Zeinab Dehghani-Ghobadi, Mohammed Sayed, Christopher Ahn, Yueh-Chiang Hu, Eunah Chung, Hee-Woong Lim, Joo-Seop Park
Lack of specificity of progenitor responses to injury in regeneration
Cecilia E. Pellegrini, Peter W. Reddien
Unique mineralization pattern revealed in TBCK syndrome mouse model
Kaitlin A. Katsura, Yuchen Jiang, Marius Didziokas, Nir Z. Badt, Sonia Dougherty, Kyle H. Vining, Elizabeth J. Bhoj
Pre-clinical models of idiopathic scoliosis implicate sex-specific roles for complement activity in modulating spinal curve severity
Vida Erfani, Brian Ciruna
Adipocyte-Derived Amino Acid Storage Proteins are Required for Germline Stem Cell Maintenance in Adult Drosophila Females
Anna B. Zike, Mekenzi O. Hazen, Madison G. Abel, Eleanor B. Goldstone, Robert C. Eisman, Lesley N. Weaver
Cell-autonomous Wnt activity promotes transient re-programming and cell cycle re-entry of coronary artery endothelial cells
Bhavnesh Bishnoi, Alfia Nirguni Saini, Vinay Rao, Omkar Golatkar, Ravindra Kailasrao Zirmire, Shruthi Viswanath, Perundurai Subramaniam Dhandapany, Soumyashree Das
Niche-dependent modular regulation of the stem cell transcriptome separates cell identity and potential
Amelie Raz, Hafidh Hassan, Yukiko Yamashita
TNAP and PHOSPHO1 function synergistically to afford critical control over the mineralisation of the postnatal murine skeleton
Lucie E Bourne, Aikta Sharma, Scott Dillon, Jacob Keen, Soher N Jayash, Natalie Crump, Lucinda AE Evans, Maya Karmali, Worachet Promruk, Claire E Clarkin, Sonoko Narisawa, Louise Stephen, Brian L Foster, José Luis Millán, Colin Farquharson, Katherine A Staines
Spinal cord regeneration deploys adult molecular programs that do not recapitulate embryonic development
Yuxiao Xu, Amulya Saini, Wenda Zhang, Lili Zhou, Mayssa H. Mokalled
Tissue composition shapes differential skeletal integration strategies during axolotl limb regeneration
Rita Aires, Sean D. Keeley, Kerstin Brandt, Mário Carreira, Doğa Berşan Güneş, Yagiz Savci, Ulrike Anne Friedrich, Andreas Dahl, Can Aztekin, Tatiana Sandoval-Guzmán
Endocardial TIE1 synergizes with TIE2 to regulate the atrial internal muscular network assembly
Kai Ding, Beibei Xu, Xinhao Yu, Xiwen Jia, Taotao Li, Xin Shen, Junda Li, Xudong Cao, Yahui Liu, Zhen Zhang, Yulong He
Hypoxia-activated scleraxis a mediates epicardial progenitor differentiation into a unique cardiac perivascular cell type
Björn Perder, Yu Xia, Jun Yao, Miaoyan Qiu, Alvin Gea Chen Yao, Muhammad Naeem, Paul Zumbo, Ignace Van der Wee, Avi Yakubov, Kazu Kikuchi, Doron Betel, Todd Evans, Michael R. Harrison, Jingli Cao
| Plant development
A Functional Basis for the Developmental Sequence of the Macrostructure of the Venus Flower Basket (Euplectella aspergillum)
Y. Mistry, S. Morankar, D. Kingsbury, N. Chawla, C. A. Penick, D. Bhate
Naturally occurring variation in gene-associated transposable elements impacts gene expression and phenotypic diversity in woodland strawberry
Santiago Priego-Cubero, Rocio Tolley, Julia Llinares-Gómez, Camila Zlauvinen, Tuomas Toivainen, Timo Hytönen, Carmen Martín-Pizarro, Ileana Tossolini, Pablo A. Manavella
Toxic metals increase root hair density by reducing epidermal cell length
Julia Zheku, Thea Do, M. Arif Ashraf
Extracellular calcium modulates pollen tube growth and guidance in Arabidopsis thaliana
Kumi Matsuura-Tokita, Yoko Mizuta, Daisuke Kurihara, Tetsuya Higashiyama
WUSCHEL modulates jasmonate signaling to control the balance between growth and defense in the shoot apical meristem
Pengfei Fan, Panagiotis Boumpas, Christian Wenzl, Yanfei Ma, Gernot Poschet, Jiao Zhao, Thomas Greb, Jan U. Lohmann
Mechanical coordination of counter-gradient growth maintains organ curvature in the apical hook
Sara Raggi, Hemamshu Ratnakaram, Adrien Heymans, Loitongbam Lorinda Devi, Özer Erguvan, Siamsa M. Doyle, François Jobert, Asal Atakhani, Sijia Liu, Manuel Petit, Jürgen Kleine-Vehn, Krzysztof Wabnik, Stéphane Verger, Stéphanie Robert
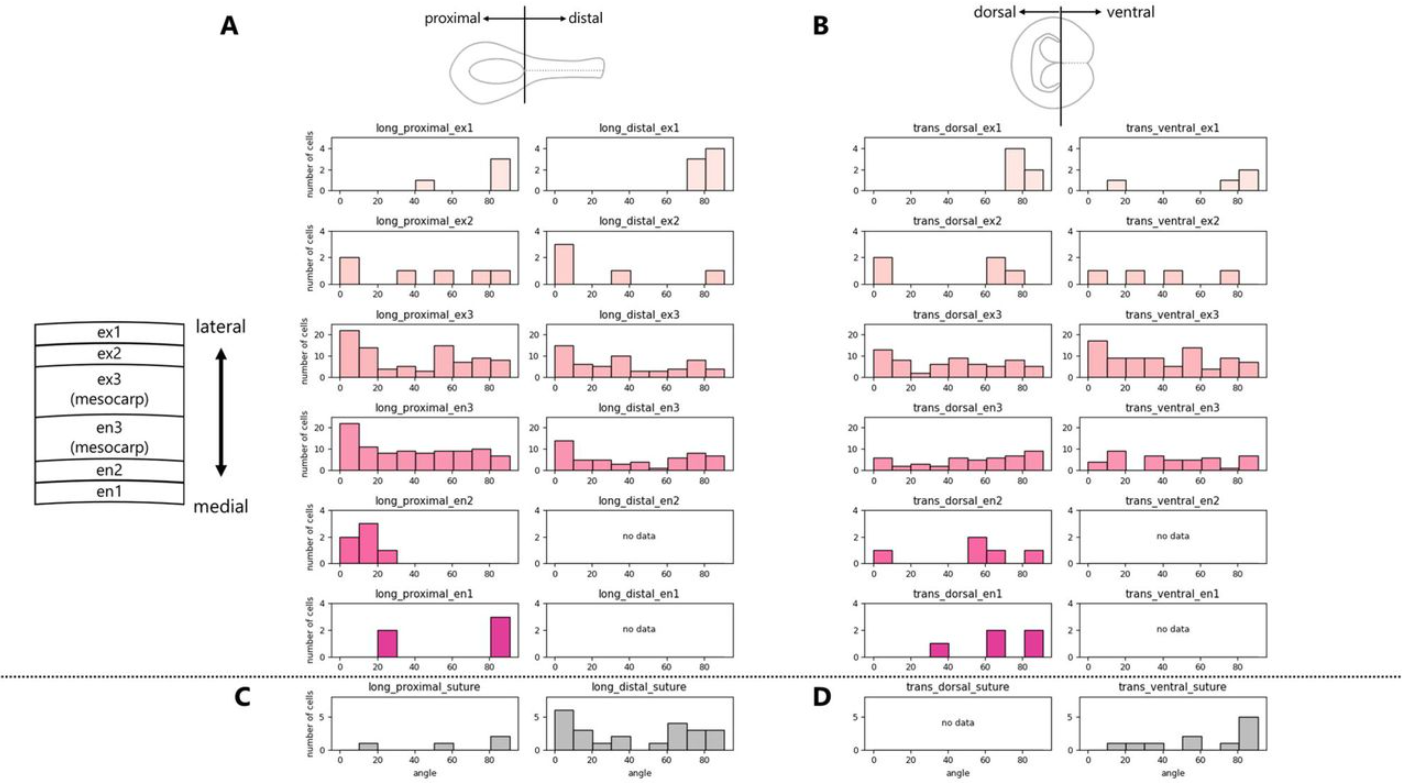
A visualization framework for cell division activity and orientation in pre-anthesis ovaries of Prunus species
Ayame Shimbo, Soichiro Nishiyama, Tatsuya Katsuno, Akane Kusumi, Hisayo Yamane, Masahiro M. Kanaoka, Ryutaro Tao
Transformers Outperform ConvNets for Root Segmentation: A Systematic Comparison Across Nine Datasets
Abraham George Smith, Sotiris Lamprinidis, Anand Seethepalli, Larry M. York, Eusun Han, Patrick Möhl, Kyriaki Boulata, Kristian Thorup-Kristensen, Jens Petersen
Evolution of moss leaf-like organs through variations in deeply conserved developmental principles
Wenye Lin, Loann Collet, Laure Mancini, Mandar Deshpande, Brendan Lane, Benjamin P. Lapointe, Agnieszka Bagniewska-Zadworna, Anne-Lise Routier-Kierzkowska, Richard S. Smith, Yoan Coudert, Daniel Kierzkowski
Ca²⁺ oscillations promote microtubule-band turnover and support tip growth in Arabidopsis zygotes
Hikari Matsumoto, Zichen Kang, Tomonobu Nonoyama, Yusuke Kimata, Satoru Tsugawa, Minako Ueda
Effects of lovastatin on auxin transport and root development in Arabidopsis thaliana
Veronica Giourieva, Christos Tersenidis, Alkiviadis Athanasiadis, Stylianos Poulios, Anna Kouskouveli, Konstantinos Vlachonasios, Emmanuel Panteris, George Komis
| Eco-evo-devo
Ancient origin of the dorso-ventral patterning system of vertebrate paired fins
Rebecca E. Dale, Silke Berger, Sara Alaei, Adele Barugahare, Marcus C. Davis, Laura Perlaza-Jimenez, Frank J. Tulenko, Peter D. Currie
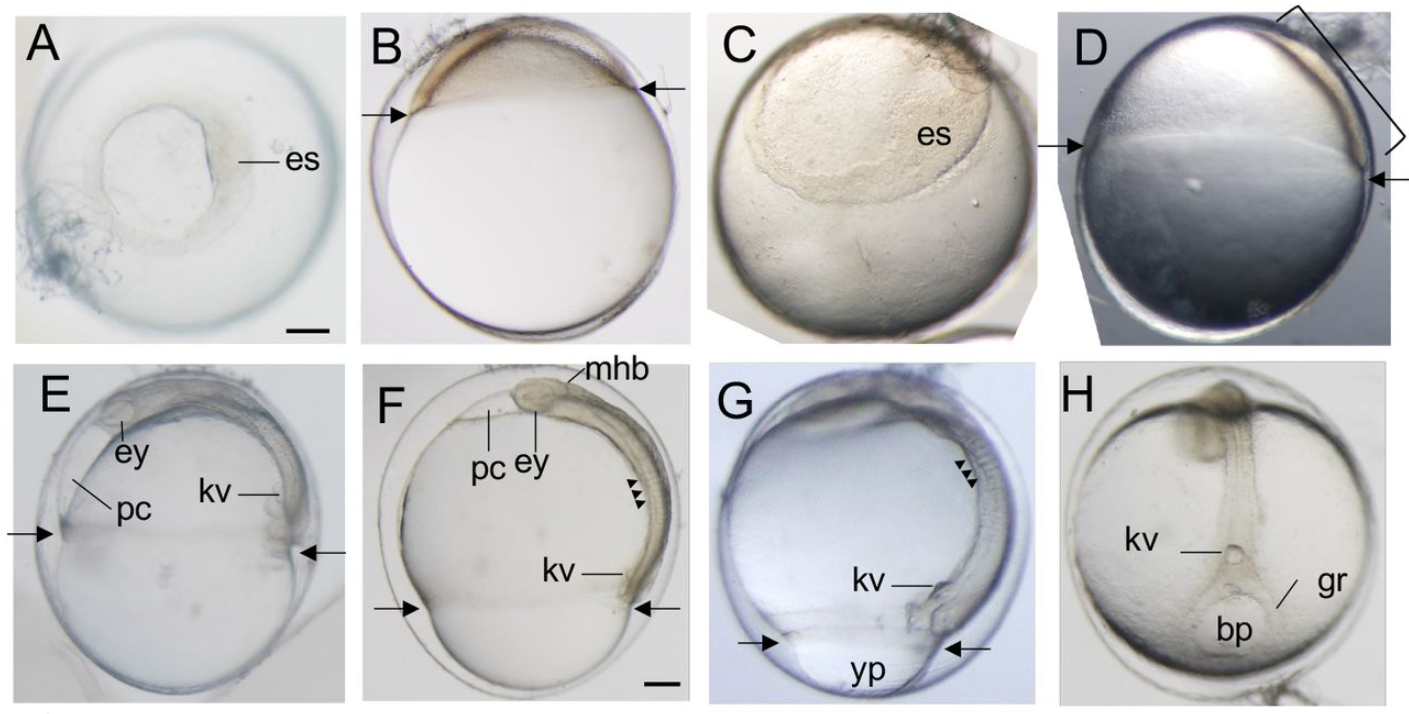
Embryonic and larval development of the Pacific saury Cololabis saira: Distinctive characteristics of a rapidly growing beloniform fish
Rie Kusakabe, Shinya Yamauchi, Shigehiro Kuraku

Age- and Light-Dependent Changes in the Zebrafish Olfactory Epithelium
George B. Chapman, Rania Abutarboush, Victoria Connaughton
Thrifty phenotypes in ants: Extending a human developmental hypothesis to a superorganism
Érik Plante, Ehab Abouheif, Jean-Philippe Lessard
Thermal Plasticity of Stage-specific Development Time and Adult Body Size under Temperature Shifts: A Case Study Using Drosophila melanogaster
Aradhya Chattopadhyay, Rishav Roy, Payel Biswas, Shampa M. Ghosh
The dynamic evolution of panarthropod germ cell specification mechanisms
Jonchee A. Kao, Emily L. Rivard, Rishabh R. Kapoor, Cassandra G. Extavour
Annelid eye evolution revealed by developmental, ultrastructural, and connectome analyses of cerebral eyes in Malacoceros fuliginosus
Suman Kumar, Anna Seybold, Oleg Tolstenkov, Sharat Chandra Tumu, Harald Hausen
Tracking morphological development in stony corals
Garrett J. Fundakowski, Viviana Brambilla, Kyle J. A. Zawada, Cher F Y Chow, Emily Croasdale, Amelia J. F. Errington, Luisa Fontoura, Wilhelm J Marais, Rachael M. Woods, Pim Edelaar, Kevin Lala, Joshua S. Madin, Maria Dornelas
Cell Biology
Whole-Cell Proteomics Identifies Novel Regulators of Ciliogenesis Beyond the Axoneme
Xiaolu Xu, Yanbao Yu, Tony Zheng, Fiona Clark, Jean Ross, Neha Sindhu, Andre L P Tavares, John B Wallingford, Shuo Wei, Jian Sun
Developmentally programmed nuclear pore complex replacement enables oocyte specification
Shruti Venkat, Tram Nguyen, Cecilia Blangini, Michelle Pollak, Karen Schindler, Maya Capelson, Prashanth Rangan
PKN is a sex- and species-specific fertilization factor in brown algae
Masakazu Hoshino, Meri Nehlsen, Rita A. Batista, Morgane Raphalen, Toshiyuki Wakimoto, Shinya Uwai, Kazuhiro Kogame, Vikram Alva, Susana Coelho
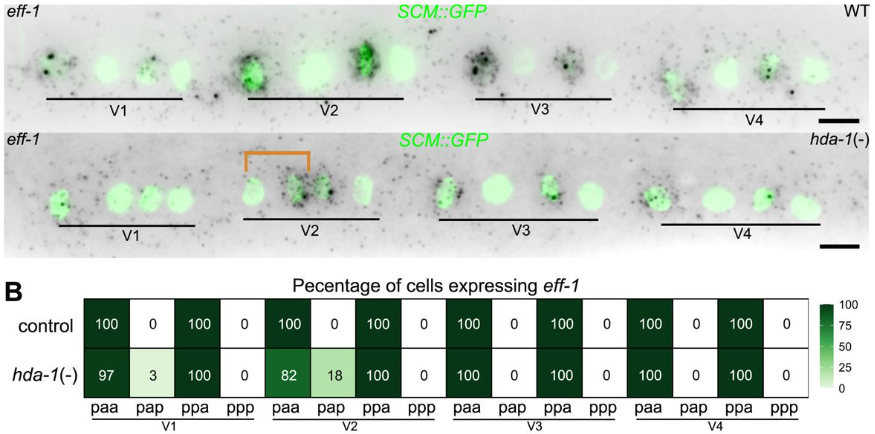
HDAC1/2-mediated repression of Wnt receptor expression orients asymmetric division polarity in C. elegans
Mar Ferrando-Marco, Beatriz Garcia del Valle, Mark Hintze, Lucy Narunsky, Shuxiao Lin, Junyue Huang, Shannon Edwards, Michalis Barkoulas
Forward Programming Identifies Inducers of Blood-Brain Barrier Properties in Human Pluripotent Stem Cell-Derived Endothelial Cells
Soniya Tamhankar, Yunfeng Ding, Fatemeh Yaghoobi Hashjin, Sarah M. Boutom, Richard Daneman, Sean P. Palecek, Eric V. Shusta
Restoration of Spermatogenesis is Dependent on Activation of a SPRY4-ERK Checkpoint Following Germline Stem Cell Damage
Ying Liu, Tansol Choi, Brad Pearson, Ryan Nachman, Whitney Woo, Na Xu, Ryan Schreiner, Romulo Hurtado, Marco Seandel, Shahin Rafii, Todd Evans
Microtubules sustain the fidelity of cellularization in a coenocytic relative of animals
Margarida Araújo, Marine Olivetta, Paolo Ronchi, Viola Oorschot, Arif Khan, Christian Tischer, Hiral Shah, Gautam Dey, Omaya Dudin
Mechanosensing and IL-13 Signaling Synergistically Modulate Intestinal Stem Cell Differentiation via STAT6 and YAP
Sarbari Saha, Thao Nguyen, Cornelis Mense, Marie Touzet-Robin, Karen Kresbach, Stephan A. Eisler, Ulrich S. Schwarz, Andrew G. Clark
Altered stem cell properties of human hematopoietic stem and progenitor cells based on bone region location
Christopher J. Wells, Christine Hall, Samantha M. Holmes, Isabelle J. Grenier-Pleau, John F. Rudan, Steve Mann, Sheela A. Abraham
Modelling
Segmented wavetrains and sites of reversal in the mouse seminiferous tubules
Kei Sugihara, Ayuki Sekisaka, Toshiyuki Ogawa, Takashi Miura
Early development of male germ cell clones shapes their reproductive success
Tatsuro Ikeda, Maurice Langhinrichs, Tamar Nizharadze, Chieko Koike, Yuzuru Kato, Katsushi Yamaguchi, Shuji Shigenobu, Kana Yoshido, Shinnosuke Suzuki, Toshinori Nakagawa, Ayumi Maruyama, Seiya Mizuno, Satoru Takahashi, Nils B. Becker, Hans-Reimer Rodewald, Thomas Höfer, Shosei Yoshida
A mathematical synthesis of genetics, development, and evolution
Mauricio González-Forero
Tools & Resources
Cellular diversity of the developing chick trigeminal ganglion at single-cell resolution
Arvind Arul Nambi Rajan, Erica J. Hutchins
A quantitative in vivo CRISPR-imaging platform identifies regulators of hyperplastic and hypertrophic adipose morphology in zebrafish
Rebecca Wafer, Panna Tandon, James E. N. Minchin
A robust human airway organoid platform enables scalable expansion and trajectory mapping of pulmonary neuroendocrine cells
Noah Candeli, Lisanne den Hartigh, Nicholas Hou, Andrés Marco, José Antonio Sánchez-Villicaña, Andrea García-González, Shashank Gandhi, Francesca Sgualdino, Alyssa J. Miller, Jason Spence, Susana Chuva de Sousa Lopes, José L. McFaline-Figueroa, Hans Clevers, Talya L. Dayton
Nuclear Histone 3 Post-Translational Modification Profiling in Whole Cells using Spectral Flow Cytometry
Carly S. Golden, Saylor Williams, Sandeep Sreerama, Sophia Blankevoort, H. Joseph Yost, Martin Tristani-Firouzi, Anna Belkina, Maria A. Serrano
Epitope-based labeling for improved live-imaging of endogenous proteins in C. elegans
Elise van der Salm, Mette H. Schroeder, Loes B. Steller, Stephanie I. Miller, Amelie Scheper, Gwen Nowee, Erik. E. Griffin, Suzan Ruijtenberg
A facile method for fluorescent visualization of newly synthesized fibrous collagen by capturing allysine aldehyde groups as cross-link precursors
Junpei Kuroda, Kazunori K. Fujii, Sugiko Futaki, Azumi Hirata, Yuki Taga, Takaki Koide
Electrical stimulation combined with p27Kip1 inactivation drives proliferative neurogenic reprogramming of Mueller glia in the adult mouse retina
Megan L. Stone, Joel Jovanovic, Edward M. Levine
Arrayed single-gene perturbations identify drivers of human anterior neural tube closure
Roya E. Huang, Giridhar M. Anand, Heitor C. Megale, Jason Chen, Chudi Abraham-Igwe, Sharad Ramanathan
Conserved cellular architecture and developmental mechanisms of the zebrafish meninges
Ashley L. Arancio, Kathryn Wilhem, Hung-Jhen Chen, Brandon M. Hernandez, Percy J. Raggi, D’Juan T. Farmer
A Single-Cell Temporal Atlas of Mouse Nasal Embryonic Development
Huan Chen, Yingxiu Chen, Mengjie Pan, Ziyu Feng, Baomei Cai, Yiyi Cheng, Sihao Chen, Jiehong Deng, Xia Yao, Chunhua Zhou, Yunjing Du, Wei He, Ruifang Zhang, Yudong Fu, Shujuan Liu, Lihui Lin, Shengyong Yu, Yuehong Yan, Duanqing Pei, Dajiang Qin, Jiekai Chen, Shangtao Cao
Cell Type Architecture and Positional Gene Gradients in an Adult Animal at Subcellular Resolution
Maoqin Sun, Yuxiaofei Wang, Kai Han, Lidong Guo, Yue Chen, Yao Li, Yaru Lin, Xiawei Liu, Zhi Huang, Qun Liu, Wenjie Guo, Rui Zhang, Wandong Zhao, Langchao Liang, Xiaoyu Wei, Li Zhou, Xuebin Mao, Jiaqi Wang, Weijian Wu, Hongwei Pan, Tao Yang, He Zhang, Xiaoshan Su, Shanshan Liu, Wenwei Zhang, Longqi Liu, Søren Tvorup Christensen, Jifeng Fei, Xin Liu, Ying Gu, Jian Wang, Huanming Yang, Gang Pei, Guangyi Fan, Xun Xu, Hanbo Li, Mengyang Xu, An Zeng
Husbandry and Maintenance of Carausius morosus Laboratory Populations
Macy Ingersoll, Petra Kovacikova, Yousuf Hashmi, Cassandra G. Extavour
A molecular and spatial resource defining tubulin isotype organization during corneal development
R Ramarapu, WR Stoehr, M Miesen, S. Border, SM Thomasy, CD Rogers
Efficient derivation and transcriptional characterization of mouse extra-embryonic endoderm stem cell lines generated by somatic cell nuclear transfer
Shuaipeng Li, Shu Wei, Guomeng Li, Mei Hu, Jiangwei Lin, Wandong Bao
Integrative Inference of Spatially Resolved Cell Lineage Trees using LineageMap
Xinhai Pan, Yiru Chen, Xiuwei Zhang
Observing concurrent subcellular dynamics in large living tissues
Charles S Wright, Sanjeev Uthishtran, Laura Z Kreplin, Hetvi R Gandhi, Abhishek Patil, Harrison M York, Samyukta Sita, Samuel A Manning, Elliot Brooks, Guizhi Sun, In-won Lee, Wing Hei Chan, Sara Hlavca, Samuel Crossman, Helen E Abud, Jan Kaslin, Avnika A Ruparelia, Peter D Currie, Kieran F Harvey, Jose M Polo, John Carroll, Senthil Arumugam
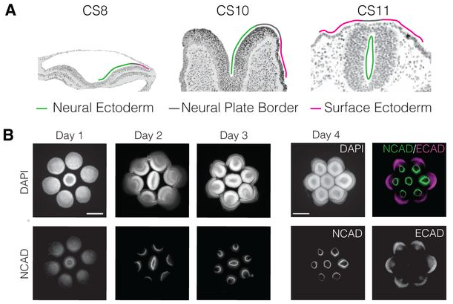
High resolution spatial transcriptomic and proteomic profiling of early primate gastrulation in utero
Nikola Sekulovski, Maliha Kabir, Anusha Rengarajan, Amber E. Carleton, Jenna K. Schmidt, Chien-Wei Lin, Kenichiro Taniguchi
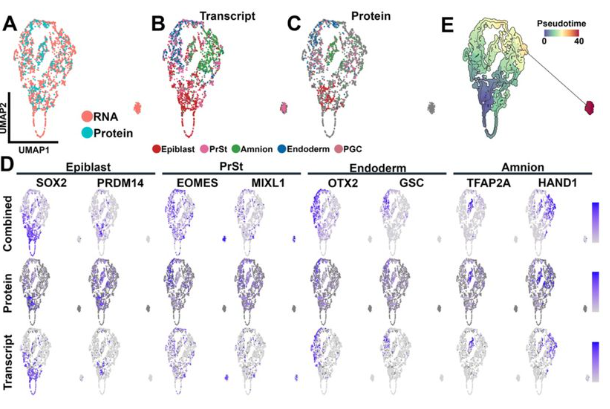
Single-cell-scale spatial transcriptome reveals early regional priming of the developing mouse ovary
Anthony S. Martinez, Tyler J. Gibson, Courtney Diamond, Jennifer Jaime, Jennifer McKey
Efficient multi-lineage cardiovascular differentiation of human pluripotent stem cells in animal serum-free conditions
Nguyen T N Vo, Kelvin Chung, Aishah Nasir, Davor Pavlovic, Chris Denning
Adapting the OpenFlexure Microscope for Affordable Live-Cell Imaging
Jodie R Malcolm, Olympia Physouni, Stuart Lacy, Mark Bentley, Stephen P Howarth, Sandy MacDonald, Alastair P Droop, Benedict Powell, Laura Wiggins, William J Brackenbury, Peter J O’Toole
Direct, high-throughput linking of single-cell imaging and gene expression
Catherine K Xu, Georg Meisl, Nikita Moshkov, Niklas A Schmacke, Karolis Goda, Alexey Shkarin, Maximilian F Schlögel, Tuomas PJ Knowles, Fabian J Theis, Linas Mazutis, Jochen Guck
A live-imaging system for Arabidopsis leaf primordia at early stages
Yujie Zhao, Hokuto Nakayama, Satohiro Okuda, Tetsuya Higashiyama, Hirokazu Tsukaya
NucVerse3D: Generalizable 3D nuclear instance segmentation across heterogeneous microscopy modalities
Jorge Vergara, Cristian Perez-Gallardo, Ricardo Velasco, Dilan Martinez, Diego Badilla, Esteban G. Contreras, Pamela Guevara, Fabián Segovia-Miranda, Hernán Morales-Navarrete
Super-resolution single-cell spatial atlas of plant de novo regeneration
Xiehai Song, Shaoman Zhang, Zhiliang Yue, Yongqi Liu, Shanshan Chen, Yani Niu, Yan Shi, Hengjia Yang, Li Xu, Naixu Liu, Yuanyuan Miao, Man Lv, Jinshan Li, Tong Wang, Meizhi Xu, Binmei Sun, Chuan Qiu, Ruirui Xu, Jizong Wang, Huawei Zhang, Shuguo Hou, Gang Li, Haodong Chen, Xing Wang Deng, Bosheng Li
Long-term ex ovo culture of Caenorhabditis elegans embryos
Clover Ann Stubbert, Cherry Soe, Pavak Kirit Shah
A Data-Driven Image Extraction and Analysis Pipeline for Plant Phenotyping in Controlled Environments
Fahimeh Orvati Nia, Joshua Peeples, Seth C. Murray, Andrew McFarland, Troy Vann, Shima Salehi, Robert Hardin, David D. Baltensperger, Amir Ibrahim, J. Alex Thomasson, Henry Fadamiro, Nithya K Subramanian, Nazar Oladepo, Uday Vysyaraju
Research practice & education
Set-up, validation, evaluation, and cost-benefit analysis of an AI-assisted assessment of responsible research practices in a sample of life science publications
Silke Kniffert, Ben Katthöfer, Robert Emprechtinger, Pasquale Pellegrini, Eva Maria Funk, Ishminder Singh Dhamrait, Yalei Zang, Ailyn Bornmüller, Ulf Toelch
Science should be machine-readable
A. Sina Booeshaghi, Laura Luebbert, Lior Pachter
bioRxiv: the preprint server for biology
Richard Sever, Samantha Hindle, Ted Roeder, Sol Fereres, Olaya Fernández Gayol, Sanchari Ghosh, Martina Proietti Onori, Emma Croushore, Kevin-John Black, Linda Sussman, Janet Argentine, Wayne Manos, Marisol Muñoz, Josh Sinanan, Tracy K. Teal, John R. Inglis
Benefits and Challenges of Integrating a Generative AI Assisted Reading Guide in an Undergraduate Journal Club Assignment
Ashley Ringer McDonald, Anne V. Vázquez


 (No Ratings Yet)
(No Ratings Yet)







 (1 votes)
(1 votes)



