For both young and established developmental biologists considering their next career move, choosing a model system with which to answer one’s research questions is a big decision. Of course, the most important thing to consider is whether or not a particular system is compatible with your research goals. But for a young scientist looking to make the move from one system to another, the ease of such a transition is an important concern. Having transitioned from sea urchin to zebrafish and then to chicken, I am often asked about my experience working with each of these three organisms, and what the practical, hands-on similarities and differences are among these creatures as model systems for development.
It began with the sea urchin
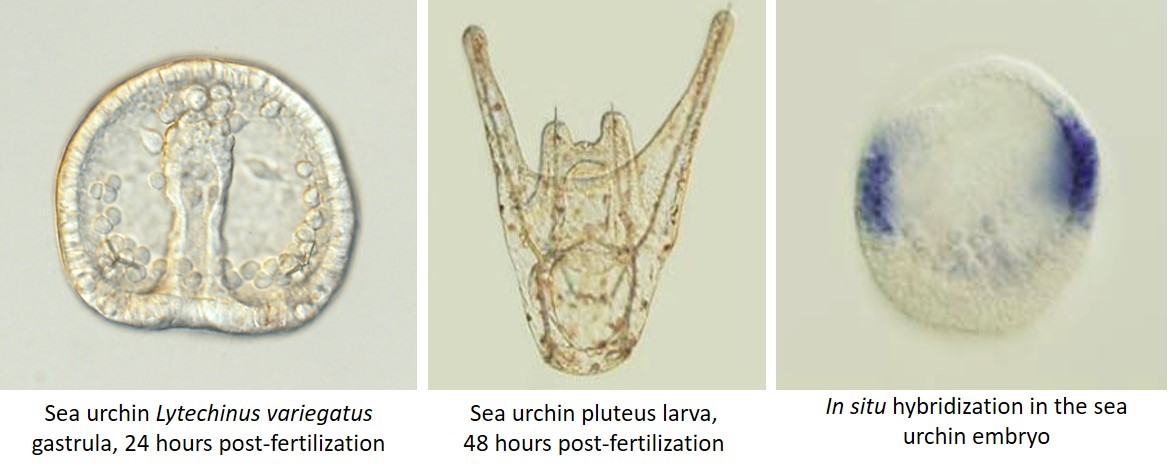
For my graduate research I studied skeleton formation in the sea urchin embryo. Most laboratories get adult sea urchins either by lab-organized diving trips if the lab is close enough to the ocean, or much more commonly by purchasing them from a supplier. When the sea urchins get to the lab, they need to be carefully acclimatized to the artificial sea water and the temperatures of lab aquaria, as any stress can make them spawn out their precious gametes prematurely. The maintenance of these aquaria and the monitoring of a new shipment of animals is an important and time-consuming job. Harvesting sea urchin sperm and eggs is as simple as injecting potassium chloride into the adults, after which they usually die (sad face!). In some species, however, adults can be shaken vigorously to make them spawn, and these animals can be used a few more times.
A major disadvantage of the sea urchin as a model system is the inability to establish stable transgenic lines or to easily create knockouts. Most analyses of development therefore depend on knockdown and overexpression experiments, which though easy to perform by microinjections, are not always clean. On the other hand, the ability to produce very large numbers of embryos from each round of fertilization gives the sea urchin a strong advantage for large-scale experiments. Since in situ hybridization and immunohistochemistry can be done on whole embryos, no time is spent sectioning embryos prior to analysis. Also, these embryos are very easy to analyze with several types of microscopy. Sea urchin embryonic development lasts only 2-4 days depending on the species, a definite benefit for moving a project along. An unexpected advantage of the sea urchin is the ability to replicate results in several closely-related species, which makes for a stronger case when putting together a story for publication. This also makes it an interesting model for evo-devo research especially when comparing multiple urchin species to other echinoderms.
Fish, fish and more fish
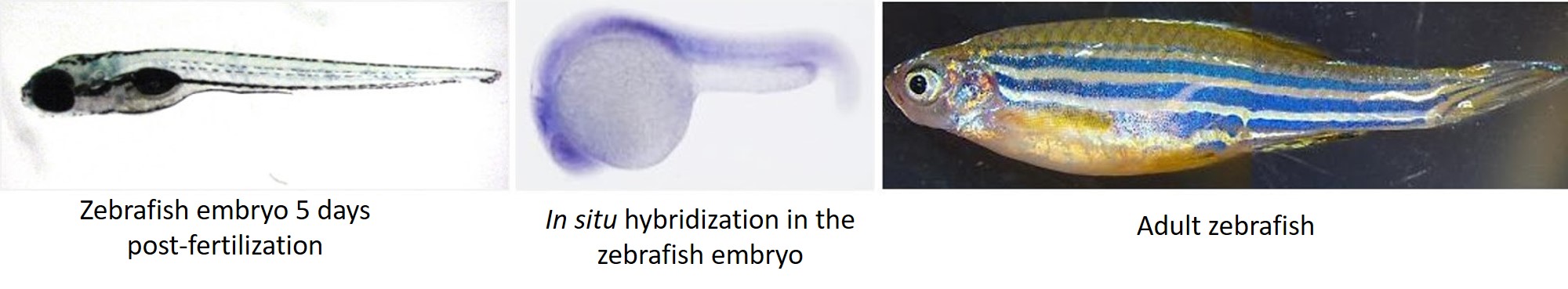
After my initial love affair with the sea urchin, I transitioned to zebrafish during my postdoc where I made zebrafish models for rare human diseases. Unlike the average sea urchin lab where adult animals are acquired from an external supplier, zebrafish labs often have several stable lines in-house, maintained by breeding adult fish and raising their offspring. While all this aquaculture can be time-consuming for a graduate student or postdoc, many institutions have fish core facilities and/or staff hired specifically for fish husbandry. Male and female zebrafish are usually kept separately, and to get them to breed, you only need to put them together in the same tank and they take over from there.
The zebrafish has many of the same advantages as the sea urchin, as it is easy to get large numbers of relatively transparent embryos. Adult animal and embryo care is similar to the sea urchin, as are methods of microinjection and analysis of results. The distinct advantage of the zebrafish (and indeed the main reason I found it attractive as a model system) is the immense power of transgenics in this system. Stable transgenic lines and knockouts are standard in the zebrafish field, and this opens up a world of possibilities for a researcher.
Another advantage of this system is the ability to culture embryos into adulthood. This lets you study a phenotype throughout the lifespan of the fish. Moreover, since the zebrafish is a closer relative to humans, antibodies raised against human proteins work better in the fish than they do in sea urchins. Aside from learning about zebrafish morphology and how to make the best use of the transgenic toolkit, my transition from sea urchin to zebrafish was seamless.
Chicken, anyone?
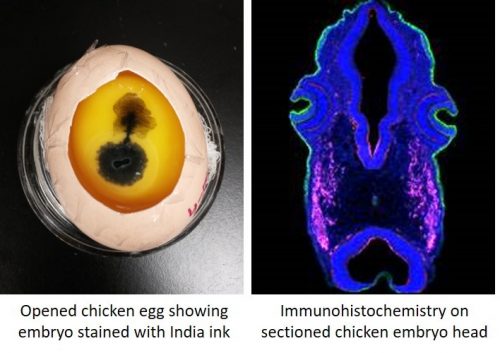
My current postdoctoral work is on neural crest and placode cells in the chicken embryo, which is distinct from the first two systems described. We get our embryos by incubating fertilized chicken eggs shipped in from a supplier, and as orders for eggs must be placed relatively far in advance, a researcher’s plan for experiments can fizzle out very quickly if egg quality is poor for a specific shipment. Also, a lot of time is spent on egg prep work, and you start out with fewer eggs, which means fewer embryos for your experiments.
Unlike aquatic embryos where you can just peek into a petri dish to tell what stage of development your embryos are, chicken eggs have to be opened and sometimes stained to check embryo development. Very early embryos especially do not take too kindly to this abuse, and may stop growing. My favorite thing about chick work is that you can stop and restart development multiple times simply by changing incubation temperature. This means that you can set a timer on an incubator and have your embryos ready at the stage at which you want them, when you want them. This cuts down on those infamous late nights in the lab.
The toolkit available to a chicken embryologist is relatively large and effective, and of the three systems described, antibodies generally work best in the chick. It is possible (but not at all common) to create stable transgenic lines of chicken, and other more common gene manipulation techniques such as morpholinos also work well. Since chicken embryos are relatively large, they often must be sectioned to observe the results of experiments.
The one thing I couldn’t escape with all three systems was the fact that embryo quality changed with the weather and the seasons.
The verdict?
Through all these transitions, I have found that studying development is a state of mind that transcends any specific model system, and the expertise you gain from one system is very easily transferable to another. However, if forced to pick a favorite, I would confess that my heart still belongs to the sea urchin.

 (13 votes)
(13 votes)
 Loading...
Loading...


 (No Ratings Yet)
(No Ratings Yet)
 recommendation being the elimination of journal-based metrics by funding agencies, institutions and publishers when judging research and researchers.
recommendation being the elimination of journal-based metrics by funding agencies, institutions and publishers when judging research and researchers.
 (2 votes)
(2 votes)



 (13 votes)
(13 votes)