Gone Fishing. Zebrafish Meeting 2014, Madison, WI.
Posted by Denise Zannino, on 15 October 2014
“In scientific thinking are always present elements of poetry. Science and music requires a thought homogeneous.” ~Albert Einstein.
This year the biannual Zebrafish meeting started out in the best possible fashion, combining music with science. The exceptionally talented organist Dr. Graham Lieschke performed three pieces composed by Bach, and Overture Hall never sounded better! Having a musical background myself, I was enthralled. Not to mention such a beautiful set of pipes deserves to be showcased. It was blissful.
I do hope this is repeated in the future! But getting back to the meeting.
Every two years the Zebrafish community descends upon Madison, WI to discuss the recent advances in the field, new technologies pertinent to our model system, and just catch up with old friends. This year was no different. Being my third Madison meeting, plus one international zebrafish meeting, I have a lot of friends to catch up with. It is especially great seeing grad student friends who are now postdocs, and postdoc friends who have moved on to start their own labs. Some friends I only see at this meeting, as they have moved to other countries.
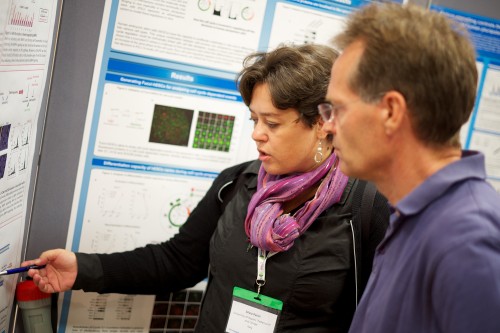
Here are some of the current and former Vanderbilt folks, catching up between sessions!
One thing that strikes me every time I attend this meeting is the vast array of biological systems that researchers are using zebrafish to study. Originally established as a model to study development, zebrafish are also well adapted for many other areas of study including cancer and growth control, neural degeneration, genetics and genomics, signaling and gene regulation, chemical biology, and behavior. There is never a problem finding something of interest. In fact, I often wanted to be in two places at the same time! Being in the field of neurodevelopment, I had lots of options, from sessions on neural degeneration, regeneration and disease; neural circuits, neurophysiology and behavior; and neural development, just to name a few. While the sessions tended to be organized by subject, the workshops were more technique based that could be utilized by many research areas. These techniques ranged from genome editing and lineage tracing to neural and vascular imaging to maternally controlled development and post embryonic development.
This year we had two Keynote Speakers, Dr. Sarah Tishkoff and Dr. John Postlethwait. Dr. Tishkoff spoke about her work studying human genetic variation in Africa using genome-scale genetic variation analyses conducted across diverse African populations. Also using genome analysis, Dr. Postlethwait uses Teleost fish as a model for understanding vertebrate development and disease by determining the origin of genes and gene families across Teleost species. In addition to these renowned researchers, there were also five plenary sessions running the gamut from imaging, genetics and genomics, to physiology and disease, to morphogenesis and organogenesis.
Each year I most look forward to learning about the latest and coolest technologies emerging in the field, and especially for me, the advances in imaging technologies. Zebrafish are so well suited for imaging studies and I can always count on a session/workshop or two at these meetings discussing the recent advances, complemented by beautiful timelapse movies. Of particular interest to me this year was the Workshop “Emerging Technologies in Neural Circuit Analysis” organized by Filippo del Bene and Marnie Halpren. Here several prominent researchers were invited to speak on new advances using transgenic tools to correlate neural activity to behavior and the some of the challenges that remain. Some of these new tools include optogenetic probes, genetically encoded voltage and calcium indicators, and new and interesting behavioral models. The use of zebrafish for behavioral studies is increasing and Erik Duboué discussed means to quantify the response of zebrafish larva to aversive stimuli. Additionally, Kurt Marsden uses a head-restrained method to map neural circuits activated during the startle response. Two esteemed researchers from Janelia Farm spoke on imaging neural activity; Douglas Kim discussed the optimization of fluorescent probes while Misha Ahrens described his research using calcium indicators to create functional maps of the entire fish brain. Other researchers also use adult zebrafish, including Hitoshi Okamoto who utilizes optogenetics, and has generated a plethora of valuable transgenic lines, and Koichi Kawakami who uses the Gal4-UAS system to genetically dissect the adult brain. Other researchers are studying neural connectivity, including Claire Wyart who studies the spinal cord using Channel Rhodospin and Alex Schier who discussed an immunohistochemical approach to study neural activity. Finally, other exciting techniques involve manipulating the activity of the neural circuit, as discussed by Harold Burgess who uses transgenic tools, and David Prober who uses TRP channels to ablate or activate specific neurons. One element I especially appreciated from this workshop was the extended question and answer time following the presentations.
Another really neat element of this year’s meeting was seeing the use of new technologies leading to great research! Two years ago the relatively new technology of genome editing via TALENs was the “it” topic of the fish meeting. Everyone was either already trying it, or going to. Every poster presenter who used the traditional morpholino approach was asked if they had considered TALENs, and entire workshops and sessions were devoted to discussing the new technology, alternative approaches, and troubleshooting some of the difficulties. This year it was great to see new data based on TALEN induced mutants! Just a wonderful example of the zebrafish community at large embracing and sharing a new technique which leads to the advancement of research across biological disciplines.
As it’s become the tradition, the last night of the meeting involves dinner on the waterfront Memorial Union Terrace with live music from Terrace After Dark, the past two meetings this has been Natty Nation. The waterfront area is perfectly setup to not only mingle with other conference attendees, but also get a local flavor with plenty of outdoor seating and an awesome view of the sunset over the water.
These festivities were followed by DJ Mike Carlson for a conference party, where everyone from PIs to graduate students can get their groove on. Overall, it’s just a great time to hang out with fellow researchers, enjoy the view, and just have some fun, as it has been for many many years. Although the next US zebrafish meeting is slated to occur in Orlando with a joint GSA model system meeting and I do not know the locational fate after that, I do know that Madison will always have a warm spot in the zebrafish community’s heart.




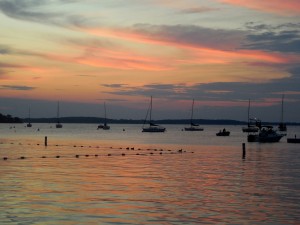
 (2 votes)
(2 votes) (No Ratings Yet)
(No Ratings Yet)
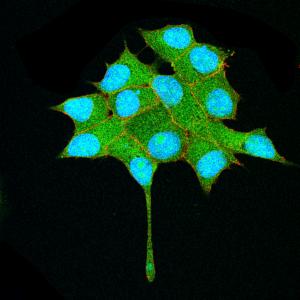





 The repair of cartilage and bone following damage remains a clinical challenge. Current cell-based therapies rely mostly on adult mesenchymal stromal cells, but the expansion of these into correctly differentiated and functionally competent chondrocytes, which give rise to cartilage and then bone, remains problematic. Here, Naoki Nakayama and colleagues develop a small molecule-based approach that mimics the embryonic somitic chondrogenesis programme and can be used to differentiate mouse embryonic stem cells (ESCs) into chondrocytes in vitro (p.
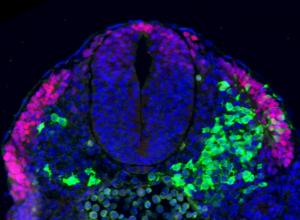
The repair of cartilage and bone following damage remains a clinical challenge. Current cell-based therapies rely mostly on adult mesenchymal stromal cells, but the expansion of these into correctly differentiated and functionally competent chondrocytes, which give rise to cartilage and then bone, remains problematic. Here, Naoki Nakayama and colleagues develop a small molecule-based approach that mimics the embryonic somitic chondrogenesis programme and can be used to differentiate mouse embryonic stem cells (ESCs) into chondrocytes in vitro (p.  The gene orthodenticle homologue 2 (Otx2) encodes a paired-type homeodomain transcription factor that is known to play a role in head morphogenesis. In the mouse, Otx2 is expressed in the anterior neurectoderm, where it is required for the differentiation of anterior neural tissues. Otx2 is also expressed in the anterior mesendoderm (AME) but its role here is unknown. On p.
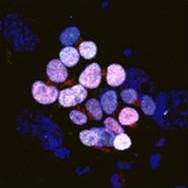
The gene orthodenticle homologue 2 (Otx2) encodes a paired-type homeodomain transcription factor that is known to play a role in head morphogenesis. In the mouse, Otx2 is expressed in the anterior neurectoderm, where it is required for the differentiation of anterior neural tissues. Otx2 is also expressed in the anterior mesendoderm (AME) but its role here is unknown. On p.  Numerous transcription factors (TFs), including PU.1 and Scl, are known to play important roles during haematopoiesis, but how these act within wider TF networks is unclear. Now, Berthold Göttgens and colleagues use transcription activator-like effectors (TALEs) to manipulate the expression of PU.1 and Scl and determine how these TFs function during developmental haematopoiesis (p.
Numerous transcription factors (TFs), including PU.1 and Scl, are known to play important roles during haematopoiesis, but how these act within wider TF networks is unclear. Now, Berthold Göttgens and colleagues use transcription activator-like effectors (TALEs) to manipulate the expression of PU.1 and Scl and determine how these TFs function during developmental haematopoiesis (p.  Pattern formation during development often depends on the differential regulation of gene expression in response to a morphogen gradient, but how such gradients govern gene expression is unclear. A simplified view suggests that the morphogen activates a transcriptional activator, and that differential gene expression is dependent on the affinity or number of binding sites for this activator within target genes. However, this model does not account for bifunctional transcriptional effectors – those that function as activators and repressors – and has also been questioned by recent experimental results. Here, James Briscoe and colleagues describe a unifying mathematical model of morphogen-dependent gene expression that can explain recent counterintuitive findings (p.
Pattern formation during development often depends on the differential regulation of gene expression in response to a morphogen gradient, but how such gradients govern gene expression is unclear. A simplified view suggests that the morphogen activates a transcriptional activator, and that differential gene expression is dependent on the affinity or number of binding sites for this activator within target genes. However, this model does not account for bifunctional transcriptional effectors – those that function as activators and repressors – and has also been questioned by recent experimental results. Here, James Briscoe and colleagues describe a unifying mathematical model of morphogen-dependent gene expression that can explain recent counterintuitive findings (p. 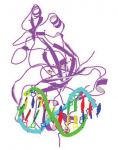 The T-box family of transcription factors exhibits widespread involvement throughout development in all metazoans. Here, Virginia Papaioannou provides an overview of the key features of T-box transcription factors and highlights their roles and mechanisms of action during various stages of development and in stem/progenitor cell populations. See the Primer on p.
The T-box family of transcription factors exhibits widespread involvement throughout development in all metazoans. Here, Virginia Papaioannou provides an overview of the key features of T-box transcription factors and highlights their roles and mechanisms of action during various stages of development and in stem/progenitor cell populations. See the Primer on p. 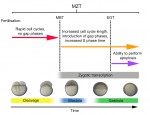 The initial phases of embryonic development occur in the absence of de novo transcription and are instead controlled by maternally inherited mRNAs and proteins. Following this period of transcriptional silence, zygotic transcription begins, the maternal influence on development starts to decrease, and dramatic changes to the cell cycle take place. Here, Steven Harvey and colleagues discuss recent work that is shedding light on the maternal to zygotic transition. See the Review on p.
The initial phases of embryonic development occur in the absence of de novo transcription and are instead controlled by maternally inherited mRNAs and proteins. Following this period of transcriptional silence, zygotic transcription begins, the maternal influence on development starts to decrease, and dramatic changes to the cell cycle take place. Here, Steven Harvey and colleagues discuss recent work that is shedding light on the maternal to zygotic transition. See the Review on p.