From a travel fellowship to starting your own lab
Posted by Mirana Ramialison, on 4 February 2014
Until last year, I was a bioinformatician post-doc in the laboratory of Prof. Richard Harvey, at the Victor Chang Cardiac Research Institute in Sydney. My project consisted of deciphering the regulatory network that controls heart development and how this network might be perturbed in disease conditions, such as congenital heart disease (which in Australia affects one in a hundred newborn babies). For this, we used a mouse cell line (HL-1 cells) which was a great system to rapidly obtain all the genome-wide information that we needed for the project, but had its limitations since it is an adult cell line that is cultured in vitro. In the meantime, the laboratory of Dr. Eileen Furlong at EMBL Heidelberg in Germany published a new method, BiTS-ChIP-seq (Bonn et al., 2012), that allows to obtain genome-wide information from tissue specific cell-types in vivo. Using this method in the developing zebrafish, in a transgenic line that specifically labels heart nuclei (cf image), would be the perfect experiment to gain insight into the nature of this gene regulatory network in vivo.
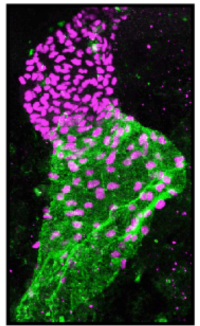
Zebrafrish embryonic heart nuclei (red) and atrium (green)
When I started to plan the experiment, I was daunted by the detailed published protocol due to its length and complexity, but mainly due to my own limitations- it included many techniques that I had never performed before. My time was limited, as my post-doc was ending at the end of the year. The quickest way to get these experiments going was to get “hands-on” experience on this protocol. These are the times when I miss Europe, where I could just hop on a train and visit a large number of diverse laboratories nearby. Yes, I am in Australia, and “popping-in” to visit any European laboratory involves a 24-hour flight (one way) and lots of $$. Fortunately, our grants management officer encouraged me to apply for a Development travelling fellowship from the Company of Biologists to support this visit. Thanks to this fellowship, I had the hands-on experience to learn the protocol and its “tricks” that I wanted, which probably saved me months of trials to get it right.
Far beyond the technical knowledge that I gained, receiving the fellowship also had great repercussions on my career path. At the time I was doing this work, I was also in the process of looking for independent positions and writing grants to fund it. I submitted a proposal to the Australian Research Council that included the BiTS-ChIP-seq experiments, in collaboration with Dr. Furlong’s laboratory. An important criterion for the grant was to provide evidence of this collaboration, ideally by showing co-authored publications. Since we had not published together at that time, the visits to Dr. Furlong’s laboratory definitely testified for the feasibility of these experiments in my hands, which was essential for the success of the grant.
In summary, receiving the Development travelling fellowship allowed me to save time on experiments, strengthen my collaborations around the world, and help to win my first grant to start my lab at the Australian Regenerative Medicine Institute in Melbourne.


 (7 votes)
(7 votes)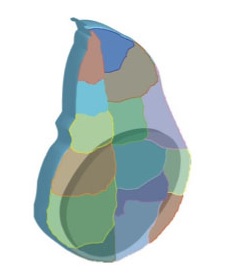

 (No Ratings Yet)
(No Ratings Yet)