Here are the highlights from the current issue of Development:
Human embryos: a mixed bag

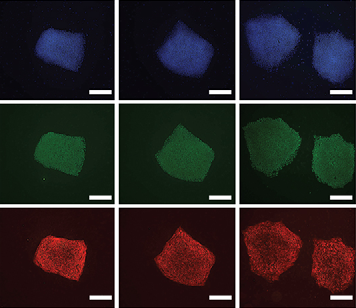
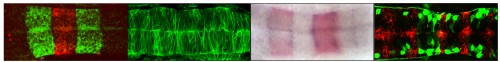
Human embryonic stem cells (hESCs) are conventionally derived from embryos that are at the blastocyst stage of development. Now, to gain insight into lineage allocation in early human development, Susan Fisher and colleagues report the derivation of various hESC lines using single blastomeres from 8- and 12-cell human embryos (p. 4010). The characterisation of these lines reveals that, at the transcriptome level, they differ from blastocyst-derived lines. Notably, blastomere-derived hESCs are enriched for genes expressed by trophoblasts and the ectoplacental cone. The researchers further show that blastomere-derived lines are hypomethylated in genomic regions that control trophoblast differentiation and early developmental processes, indicative of trophoblast competence. Variations in gene expression profiles are also observed among the various blastomere-derived lines (all of which are derived from embryos from a single couple), highlighting the existence of blastomere heterogeneity. In line with this, the authors show that levels of EOMES, T, GDF15 and active b-catenin differ among the blastomeres of 8- to 10-cell human embryos. Finally, the researchers report the derivation of the first human trophoblast stem cell line, which could be used in the future to model placental development and disorders.
Cilia: at the heart of chamber maturation

Increasing evidence suggests that blood flow and biomechanical forces generated by the developing heart feedback to regulate cardiac chamber formation and maturation. How such forces are sensed and translated, however, remains unclear. Here, Jiandong Liu and co-workers show that, in zebrafish, cardiac contraction activates Notch signalling in the endocardium to control chamber maturation (p.4080). By analysing embryos deficient for troponin T type 2a (tnnt2a), which encodes an essential component of the cardiac contraction apparatus, the researchers first show that cardiac contraction is required for the formation of cardiac trabeculae – the luminal protrusions that are indicative of cardiac chamber maturation. They further show that cardiac contraction controls Notch signalling in the endocardium; notch1b expression is reduced in tnnt2a–/– embryos. Notch activation, they report, induces the expression of ephrin b2a and neuregulin 1 to promote trabeculation. Finally, the authors demonstrate that shear stress controls notch1b expression in a primary cilia-dependent manner, suggesting that primary cilia in this context are responsible for detecting fluid flow. In summary, these findings highlight a molecular mechanism that links flow sensing to the transcriptional changes that regulate cardiac development.
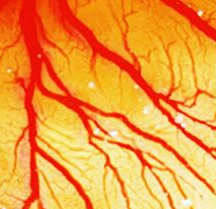
Vascular development in full flow

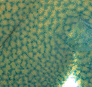
Vascular development and angiogenesis are known to be regulated by various signals, but the roles played by blood flow and biomechanical signals are unclear. This is mostly because the ability to image and measure changes in blood flow has been limited. Now, Elizabeth Jones and colleagues develop a method to simultaneously image blood flow dynamics and vascular remodelling (p. 4158), and use this technique to show that flow dynamics control sprout location and elongation in quail embryos (p.4151). Their imaging approach uses micro-particle image velocimetry: embryos are injected with a fluorescent dye that labels endothelial cells and with fluorescent microspheres that act as tracers of fluid motion. Subsequent imaging via a high-speed camera allows changes in blood flow dynamics and vessel geometry to be quantified. Using this method, the researchers demonstrate that sprout location can be predicted based on flow dynamics; sprouts form from vessels that are at a lower pressure towards vessels at a higher pressure and localise to points where shear stress, a force created by flow, is at a minimum. In addition, the rate of sprout elongation is proportional to the pressure difference between the two vessels. These studies provide insights into the hemodynamic forces at play during vascular development and open the door to further studies of the biomechanical control of vascular remodelling.
Con-Nek-ting cilia biogenesis and resorption

The left-right organiser (LRO) is a transient ciliated structure that plays a key role in establishing left-right (LR) asymmetry in the vertebrate embryo. However, the mechanisms that control the formation and resorption of cilia on this structure are unclear. In this issue (p. 4068), Martina Brueckner and colleagues reveal that Nek2 regulates cilia biogenesis and resorption at the Xenopus LRO. They show that both the knockdown and the overexpression of nek2, which encodes a NIMA-like kinase, result in reduced cilia numbers and motility at the LRO and hence abnormal LR development. Nek2 is known to play a role in centriole separation and, in line with this, the authors reveal that the knockdown of nek2results in centriole defects in the LRO. They further show that Nek2 acts upstream of the tubulin deacetylase Hdac6, and that it interacts with the nucleoporin Nup98, to control cilia resorption. Together, these findings demonstrate that Nek2 is involved in multiple stages of the cilia life cycle. Given that NEK2 has previously been implicated in abnormal laterality in humans, these findings also provide further evidence that links Nek family kinases to human ciliopathies.
An appendage to Hox gene function

Hox genes are best known for their role in axial patterning during embryogenesis, but they have also been implicated in the development and patterning of cutaneous accessory organs, such as hair follicles and mammary glands. How do they function in this context? Here, by showing that Hoxc8 can initiate an ectopic mammary gland programme in mice, Lara Carroll and Mario Capecchi propose that Hox genes regulate the distribution of cutaneous appendages (p. 4056). They first show that Hoxc8 is transiently expressed in the early surface ectoderm prior to mammary line formation. The researchers then show that the conditional overexpression of Hoxc8 – to express it in regions where it is not normally expressed – results in the formation of ectopic mammary placodes. These ectopic rudiments express known mammary placode markers, such as Tbx3 and Wnt10b. The authors further report that the ablation of ectodermal Tbx3 prevents the formation of both normal and ectopic mammary rudiments, suggesting that Tbx3 is directly regulated by Hoxc8 during mammary development. Together, these and other findings highlight a role for Hoxc8 in the initial stages of mammary development and suggest that Hoxc8 and other Hox genes play roles during the regional specification of cutaneous appendages.
PLUS…
How to make an oligodendrocyte
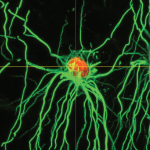 The loss of oligodendrocytes – the cells that produce the myelin sheath of axons – can result in a broad array of diseases including cerebral palsy and multiple sclerosis. Accordingly, replacing lost oligodendrocytes holds great promise as a therapeutic strategy. Here, Goldman and Kuypers describe the molecular events regulating oligodendrocyte development in vivo and discuss how our understanding of this process has led to the establishment of methods for producing oligodendrocytes in vitro. See the Primer on p. 3983
The loss of oligodendrocytes – the cells that produce the myelin sheath of axons – can result in a broad array of diseases including cerebral palsy and multiple sclerosis. Accordingly, replacing lost oligodendrocytes holds great promise as a therapeutic strategy. Here, Goldman and Kuypers describe the molecular events regulating oligodendrocyte development in vivo and discuss how our understanding of this process has led to the establishment of methods for producing oligodendrocytes in vitro. See the Primer on p. 3983
*Also see the other articles in the “How to make…” series here
Morphogen rules: design principles of gradient-mediated embryo patterning
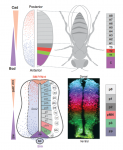 The Drosophila blastoderm and the vertebrate neural tube are archetypal examples of morphogen-patterned tissues that create precise spatial patterns of different cell types. Here, James Briscoe and Stephen Small compare these systems in the context of gene regulatory networks and dynamical systems theory. This comparison reveals several shared features that suggest that a set of common design principles underpins the patterning of both tissues. See the Review on p. 3996
The Drosophila blastoderm and the vertebrate neural tube are archetypal examples of morphogen-patterned tissues that create precise spatial patterns of different cell types. Here, James Briscoe and Stephen Small compare these systems in the context of gene regulatory networks and dynamical systems theory. This comparison reveals several shared features that suggest that a set of common design principles underpins the patterning of both tissues. See the Review on p. 3996
 (No Ratings Yet)
(No Ratings Yet)
 Loading...
Loading...



 (No Ratings Yet)
(No Ratings Yet)
 (7 votes)
(7 votes)
 – Marcela
– Marcela  (1 votes)
(1 votes)




 The loss of oligodendrocytes – the cells that produce the myelin sheath of axons – can result in a broad array of diseases including cerebral palsy and multiple sclerosis. Accordingly, replacing lost oligodendrocytes holds great promise as a therapeutic strategy. Here,
The loss of oligodendrocytes – the cells that produce the myelin sheath of axons – can result in a broad array of diseases including cerebral palsy and multiple sclerosis. Accordingly, replacing lost oligodendrocytes holds great promise as a therapeutic strategy. Here,  The Drosophila blastoderm and the vertebrate neural tube are archetypal examples of morphogen-patterned tissues that create precise spatial patterns of different cell types. Here, James Briscoe and Stephen Small compare these systems in the context of gene regulatory networks and dynamical systems theory. This comparison reveals several shared features that suggest that a set of common design principles underpins the patterning of both tissues. See the Review on p.
The Drosophila blastoderm and the vertebrate neural tube are archetypal examples of morphogen-patterned tissues that create precise spatial patterns of different cell types. Here, James Briscoe and Stephen Small compare these systems in the context of gene regulatory networks and dynamical systems theory. This comparison reveals several shared features that suggest that a set of common design principles underpins the patterning of both tissues. See the Review on p.