Cricket Leg Regeneration: Histone Modification Matters
Posted by Hideyo Ohuchi, on 22 October 2015
In autumn, crickets generally exhibit chirping songs in the temperate East Asian country of Japan. While the African field cricket Gryllus bimaculatus originates from tropical countries, it is an emerging model animal globally because of its ability to regenerate amputated legs during nymph and its developmental mode (short germ band) (Mito and Noji, 2008).
Many living organisms in the animal kingdom are able to regrow their body parts following injury. Examples of body parts that may be regrown include the lens and tail of amphibians, the head of planarians, and the heart of fish. In contrast, it has long been assumed that humans cannot restore lost body parts, except for particular tissues, including the epidermis, the liver, and the ovarian surface after ovulation. Therefore, it is important to elucidate the molecular mechanisms involved in regeneration processes using animal models that are able to regenerate body parts for subsequent application in non-regenerative human organs and tissues.
Within the last 2 years, comparative genomic studies of two planarian species with different regenerative abilities led to the successful regeneration of heads by reducing beta-catenin activity from otherwise non-regenerative tail fragments (Umesono et al., 2013). Studies of vertebrates with the ability to restore limbs, including newts, frogs, and salamanders, have demonstrated that limb regeneration occurs in a stepwise manner. The limb regeneration process is divided into at least three phases: wound healing, dedifferentiation, and redevelopment, with the redevelopment phase mimicking embryonic development (Endo et al., 2004).
The cricket leg is composed of six segments that are arranged along the proximo-distal (PD) axis: coxa, trochanter, femur, tibia, tarsus, and claw (Figure 1). 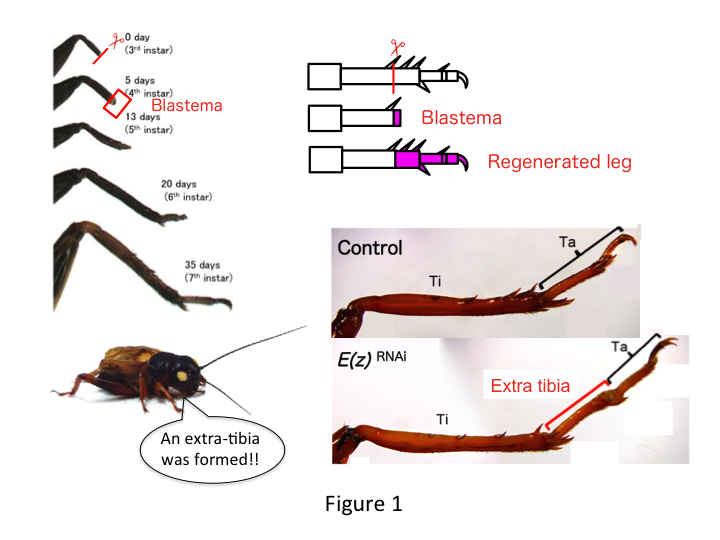 The tarsus is further subdivided into three tarsomeres. When the tibia of the third-instar nymph is amputated, the leg regenerates and recovers its allometric size and proper shape by the sixth instar (i.e., within 20 days of amputation), being restored to almost normal adult size and shape. Soon after healing, the blastema (a pool of cells that proliferate) develops in the distal region of the amputated leg. Blastema cells proliferate and form the missing structures by intercalary processes between the most distal region and the remaining part of the leg (French et al., 1976).
The tarsus is further subdivided into three tarsomeres. When the tibia of the third-instar nymph is amputated, the leg regenerates and recovers its allometric size and proper shape by the sixth instar (i.e., within 20 days of amputation), being restored to almost normal adult size and shape. Soon after healing, the blastema (a pool of cells that proliferate) develops in the distal region of the amputated leg. Blastema cells proliferate and form the missing structures by intercalary processes between the most distal region and the remaining part of the leg (French et al., 1976).
Previously, we performed comparative transcriptome analysis of regenerating and normal amputated legs of crickets to profile mRNA expression associated with leg regeneration (Bando et al., 2013). We first focused on the upregulation of Jak/Stat pathway genes, which are linked to the immune system. RNA interference (RNAi) of genes in this pathway thoroughly disturbed leg regeneration. In contrast, RNAi against Socs, a suppressor of cytokine signaling, caused leg elongation. Additional experiments showed that the Jak/Stat pathway promotes cell proliferation downstream of the Ds/Fat pathway.
Subsequently, we investigated epigenetic regulation during cricket leg regeneration. Tetsuya Bando, a senior investigator in our group, identified one gene for histone H3 lysine 27 (H3K27) methyltransferase, E(z), and one gene for histone H3K27 demethylase, Utx, in G. bimaculatus. Cloning Gryllus genes is now a straightforward process due to information being available about the cricket genome (Mito and Noji, personal communication). Methylation of histone H3K27 by E(z) represses the expression of target genes by recruiting Polycomb group proteins. Conversely, demethylation of the trimethylated histone H3K27 by Utx promotes gene expression. Tetsuya found that the transcription of both E(z) and Utx genes is upregulated in the blastema cells of amputated legs (Bando et al., 2013). In situ hybridization verified that both genes are ubiquitously transcribed in the regenerating legs of crickets, and that both genes are expressed in developing embryos (Hamada et al., 2015). Immunostaining on the amputated tiny legs after RNAi by Yoshimasa Hamada (a PhD student) confirmed that E(z) and Utx contribute to the methylation and demethylation at histone H3K27me3, respectively, during leg regeneration.
However, Yoshimasa unexpectedly found that the extra leg segment is formed after RNAi against E(z) (Figure 1). Initially, we were not able to determine the identity of the leg segment. Morphologically, the leg segment appeared to be a tibia, because it had spines and spurs that were characteristic to an authentic tibia. Our opening hypothesis was that the phenotypes after RNAi might depend on the amputation site in the tibia. However, even when a leg is amputated in the distal part of the femur, the extra tibia-like segment emerges. Pattern formation along the antero-posterior and dorso-ventral axes remained unchanged, except along the PD axis. We then examined whether the amputation site along the PD axis in the tibia influenced phenotypic severity. The extra-tibia that formed became longer the more proximal the amputation sites on the tibia (Figure 1). Conversely, RNAi against Utx resulted in the loss of joint formation between tarsomere 1 (Ta1) and Ta2 (Figure 2). 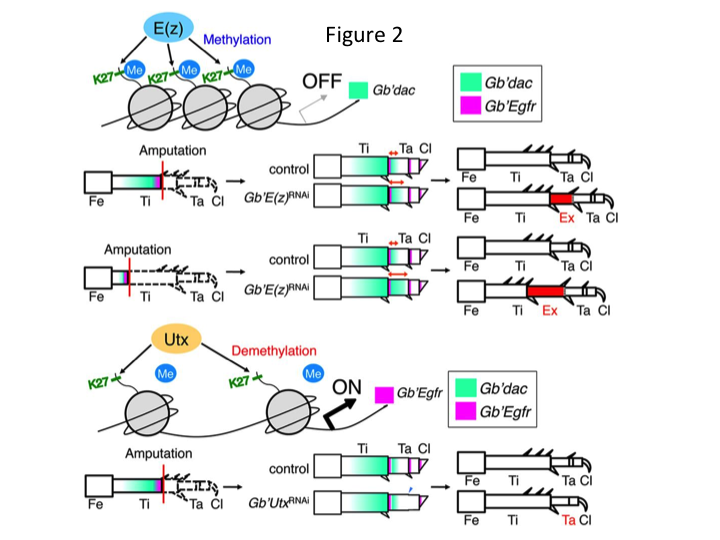 In situ hybridization showed that the expression of leg patterning genes altered along the PD axis. Specifically, the domain of dachshund (dac) expression expanded in E(z)RNAi regenerating legs, whereas Egfr expression diminished in UtxRNAi legs. Therefore, E(z) may repress dac expression during normal leg regeneration, whereas Utx induces Egfr expression.
In situ hybridization showed that the expression of leg patterning genes altered along the PD axis. Specifically, the domain of dachshund (dac) expression expanded in E(z)RNAi regenerating legs, whereas Egfr expression diminished in UtxRNAi legs. Therefore, E(z) may repress dac expression during normal leg regeneration, whereas Utx induces Egfr expression.
dac encodes a transcriptional co-repressor that is categorized in leg gap genes. dac produces crude positional values along the PD axis of the leg and mediates the formation of the distal tibia and Ta1 (the proximal tarsomere) during cricket leg regeneration (dac expression domain is shown in green in Figure 2) (Ishimaru et al., 2015). Specifically, dac promotes tibial cell proliferation. Therefore, because RNAi against E(z) upregulates dac, E(z) expression in the blastema cells may suppress the blastemal overproliferation by repressing extra dac expression.
This information raises the question of how E(z) specifically regulates dac expression. Furthermore, what is the mechanism that determines the target genes of E(z)? E(z) belongs to the Polycomb repressive complex 2 (PRC2), which is one of three Polycomb group (PcG) complexes (Schuettengruber et al., 2007). During cricket embryogenesis, E(z) represses the anterior expansion of Hox gene expression and provides proper identity in embryos (Matsuoka et al., 2015). This information indicates that the target genes of E(z) differ depending on the cellular context. A DNA binding protein, Pleiohomeotic (Pho), along with other factors, binds to the Polycomb response elements (PRE) of target genes, after which E(z) trimethylates histone H3K27. Although PREs have only been identified in Drosophila, the meta-analysis of putative target genes for PcG proteins has shown that many of the target genes are common to the fly, mouse, and humans. dac and Egfr are included among these genes (Schuettengruber et al., 2007). Thus, the regulatory region of the cricket dac gene probably contains PREs, through which E(z) epigenetically regulates the expression of dac during cricket leg regeneration (Figure 2). Ongoing research is focused on characterizing the functions of the Pho gene and other PcG complex genes and epigenetic modifiers during Gryllus leg regeneration.
Finally, why does E(z) RNAi cause extra-tibia formation? One hypothetical scenario is that when the tibia is amputated at the proximal position where dac expression is low, Utx expression (which dominates E(z) expression) permits dac expression (Figure 3a) to restore the tibia. 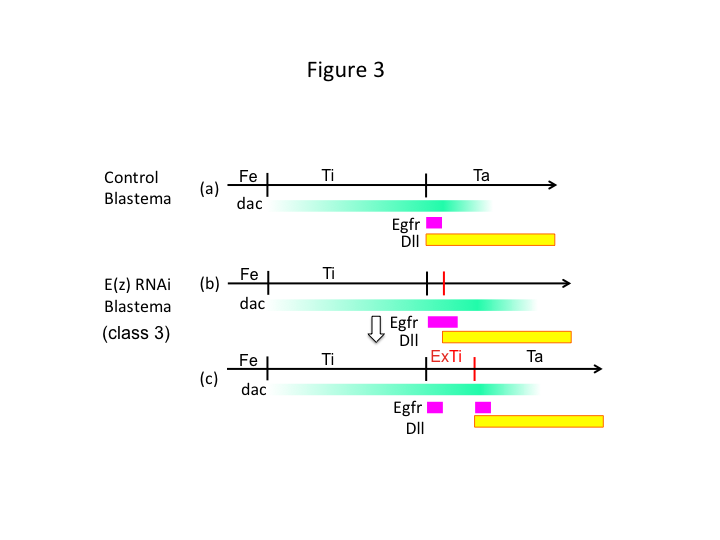 Thus, these histone modifiers sense the positional values along the PD axis of the amputation site, and fine-tune the expression level of leg patterning genes, like dac. In the case of E(z) RNAi just before proximal amputation, intense dac expression is induced and expands in the regenerating leg (Figure 3b). Distal-less (Dll) expression, which is another leg gap gene that specifies the distal domain of the leg (Angelini and Kaufman, 2005), may shift more distally depending on expanded dac expression (Figure 3b). Thus, the Egfr-expressing domain may be separated into two parts where (1) Dll expression is low and (2) Dll is high. The extra-tibia probably forms between the two different Egfr-expressing domains by intercalating cell proliferation and patterning (Figure 3c).
Thus, these histone modifiers sense the positional values along the PD axis of the amputation site, and fine-tune the expression level of leg patterning genes, like dac. In the case of E(z) RNAi just before proximal amputation, intense dac expression is induced and expands in the regenerating leg (Figure 3b). Distal-less (Dll) expression, which is another leg gap gene that specifies the distal domain of the leg (Angelini and Kaufman, 2005), may shift more distally depending on expanded dac expression (Figure 3b). Thus, the Egfr-expressing domain may be separated into two parts where (1) Dll expression is low and (2) Dll is high. The extra-tibia probably forms between the two different Egfr-expressing domains by intercalating cell proliferation and patterning (Figure 3c).
Our goal is to elucidate blueprints for “making a regenerated leg” by using this attractive hemimetabolous insect model. The blueprints are expected to clarify how the number of leg segments is determined. Our striking observations on RNAi against E(z) leading to “extra tibia formation” represent an important step towards elucidating this process.
References
- Mito, T. and Noji, S. (2008). The Two-Spotted Cricket Gryllus bimaculatus: An emerging Model for Developmental and Regeneration Studies. Cold Spring Harb Protoc, 331-346.
- Umesono, Y., Tasaki, J., Nishimura, Y., Hrouda, M., Kawaguchi, E., Yazawa, S., Nishimura, O., Hosoda, K., Inoue, T. and Agata, K. (2013). The molecular logic for planarian regeneration along the anterior–posterior axis. Nature 500, 73–76.
- Endo, T., Bryant, S. V. and Gardiner, D. M. (2004). A stepwise model system for limb regeneration. Dev Biol 270, 135–145.
- French, V., Bryant, P. J. and Bryant, S. V. (1976). Pattern regulation in epimorphic fields. Science 193, 969-981.
- Bando, T., Ishimaru, Y., Kida, T., Hamada, Y., Matsuoka, Y., Nakamura, T., Ohuchi, H., Noji, S. and Mito, T. (2013). Analysis of RNA-Seq data reveals involvement of JAK/STAT signalling during leg regeneration in the cricket Gryllus bimaculatus. Development 140, 959-964.
- Hamada, Y., Bando, T., Nakamura, T., Ishimaru, Y., Mito, T., Noji, S., Tomioka, K. and Ohuchi, H. (2015). Leg regeneration is epigenetically regulated by histone H3K27 methylation in the cricket Gryllus bimaculatus. Development 142, 2916-2927.
- Ishimaru, Y., Nakamura, T., Bando, T., Matsuoka, Y., Ohuchi, H., Noji, S. and Mito, T. (2015). Involvement of dachshund and Distal-less in distal pattern formation of the cricket leg during regeneration. Sci Rep 5, 8387.
- Schuettengruber, B., Chourrout, D., Vervoort, M., Leblanc, B. and Cavalli, G. (2007). Genome regulation by polycomb and trithorax proteins. Cell 128, 735-745.
- Matsuoka, Y., Bando, T., Watanabe, T., Ishimaru, Y., Noji, S., Popadić, A. and Mito, T. (2015). Short germ insects utilize both the ancestral and derived mode of Polycomb group-mediated epigenetic silencing of Hox genes. Biol Open 4, 702-709.
- Angelini,R. and Kaufman, T. C. (2005). Insect appendages and comparative ontogenetics. Dev Biol 286, 57-77.


 (2 votes)
(2 votes)



 MyoD and Twist-1 are transcription factors known to promote and inhibit muscle cell differentiation respectively. Phylactou and co-workers identify a mechanism of myogenesis in which MyoD and miR-206 downregulate the expression of Twist-1. This pathway might also play an important role in muscle disease. Read the paper
MyoD and Twist-1 are transcription factors known to promote and inhibit muscle cell differentiation respectively. Phylactou and co-workers identify a mechanism of myogenesis in which MyoD and miR-206 downregulate the expression of Twist-1. This pathway might also play an important role in muscle disease. Read the paper 

 (No Ratings Yet)
(No Ratings Yet)


 Atoh1 is a key regulator of the differentiation of hair cells, the sensory cells that support audition: it is upregulated during their differentiation and downregulated at postnatal stages. But what are the mechanisms underlying Atoh1 transcriptional regulation during inner ear development? To address this question, Neil Segil and co-workers (p.
Atoh1 is a key regulator of the differentiation of hair cells, the sensory cells that support audition: it is upregulated during their differentiation and downregulated at postnatal stages. But what are the mechanisms underlying Atoh1 transcriptional regulation during inner ear development? To address this question, Neil Segil and co-workers (p. 


 Mike Levine, director of the Lewis-Sigler Institute for Integrative Genomics at Princeton University, is a developmental biologist who has dedicated his career to understanding how gene expression is regulated during development. We had a lively chat with Mike at this year’s Society for Developmental Biology meeting, where he was awarded the Edwin Grant Conklin Medal. See the Spotlight article on p.
Mike Levine, director of the Lewis-Sigler Institute for Integrative Genomics at Princeton University, is a developmental biologist who has dedicated his career to understanding how gene expression is regulated during development. We had a lively chat with Mike at this year’s Society for Developmental Biology meeting, where he was awarded the Edwin Grant Conklin Medal. See the Spotlight article on p. 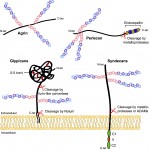 Heparan sulfate proteoglycans (HSPGs) have long been implicated in a wide range of cell-cell signaling and cell-matrix interactions, both in vitro and in vivo in invertebrate models. Here,
Heparan sulfate proteoglycans (HSPGs) have long been implicated in a wide range of cell-cell signaling and cell-matrix interactions, both in vitro and in vivo in invertebrate models. Here,