An interview with Caroline Dean
Posted by the Node, on 18 August 2015
This interview first featured in Development.
Caroline Dean is a plant biologist based at the John Innes Centre in Norwich, UK. She helped to establish Arabidopsis as a model plant organism, and has worked for many years on the epigenetic mechanisms that regulate vernalisation, the process by which plants accelerate their flowering after periods of prolonged cold. We met Caroline at the recent Spring Meeting of the British Society for Developmental Biology. We asked her about her career, her thoughts on the plant field and being awarded this year’s FEBS EMBO Women in Science Award.
 We are here at the Spring Meeting of the British Society for Developmental Biology. How are you finding the meeting?
We are here at the Spring Meeting of the British Society for Developmental Biology. How are you finding the meeting?
This is not a meeting that I come to very often, but the questions asked here are always interesting because people think about my topic in a slightly different way. It’s always very interesting and illuminating to see what grabs people’s attention and what they get confused about.
This year you were awarded the FEBS EMBO Women in Science Award. What does receiving this prize mean to you, and what advice do you have for young female scientists?
I was thrilled to pieces! I heard on New Year’s Eve that I’d won it, so it was a great celebration that evening. It is a real honour because the women who have won this prize over the years are really fantastic scientists.
People have asked me how to encourage younger female scientists to break through the glass ceiling. There are still relatively few senior women scientists in Europe, although it’s not as bad as it used to be. My advice is to take the next obvious step at each stage of your career and not to worry too much about the long-term difficulties of a research career. When I started my career I didn’t have a clear goal, I didn’t aim to become a big leader in research. I just did research because I enjoyed it and found it an interesting topic. I really enjoyed being a graduate student and a postdoc, and now I really enjoy running a lab. So I would encourage young females to think “Yes, I can do it”, instead of worrying about “Well, can I achieve a work-life balance, can I compete in the current funding scenario, can I do this, can I do that?” I say to my postdocs “Jump, and then think!”
The nice thing about a science career is that it’s very flexible. As a PI you are your own boss, so if I needed to go off when the children were sick I did, and then caught up later. It’s not the type of job where you have to go in at set hours, so the flexibility helps enormously with the work-life balance. And children grow up very fast. There’s a lot of life left once they’ve left home!
Do you think role models, such as the women who have won this prize, are helpful?
I think role models do help. When I was a teenager my mother was the breadwinner, so for me the idea of a working mother was not unusual. At the John Innes Centre, I am involved in mentoring a number of project leaders and I also enjoy chatting to students and postdocs about research careers. I hope the younger scientists think “she managed to have a career and two kids who seem perfectly normal, and she is still quite into her science”, and that encourages them to continue.
How did you first become interested in biology? Was there someone who inspired you?
I loved the documentaries of the marine biologist Jacques Cousteau. I actually dragged my then boyfriend all the way down to Marseilles when I was 18 to see if I could meet him! Of course I didn’t, but I started off studying marine biology at university. I then discovered biochemistry, which I hadn’t really done at school. There was a particular plant biochemistry practical that I loved, where we isolated chloroplasts and did electron transport analysis. It really hooked me on lab work. I decided to be a technician so that I could continue doing lab work and after a year began a PhD – and it just rolled from there.
You did your undergraduate and PhD in the UK, but moved to California to do a postdoc in a biotech company. What was it like to work in industry?
The prospect of genetically modifying plants emerged when I finished my PhD, and venture capital funded a few start-up biotech companies. I did my postdoc in one of them, Advanced Genetic Sciences, in Oakland, California. The director, John Bedbrook (who came from academia), hired a bunch of academics. We had five years’ worth of money and our aim was to learn how to modify plants genetically and get foreign genes expressed in them. I learned all my molecular biology there. Very exciting times, because science was moving very quickly. So yes, it was a biotech company – but I could do fundamental academic research as part of the more biotech projects, e.g. generation of herbicide-tolerant plants. After five years, in 1988, I then moved back to the UK, to the John Innes Centre.
Your lab works on vernalisation. Why this scientific question?
When I was doing my postdoc in America, the Arabidopsis wave started – Elliot Meyerowitz and Chris Somerville initiated the use of a molecular genetic approach in Arabidopsis thaliana. I hadn’t done any such work while at the company, but I could see that this would open up analysis of really complex traits. For a trait like flowering time or developmental timing, we had no clue about which genes would be involved – you had to take a genetic approach. I chose to study vernalisation because of a conversation with a seller in a nursery garden in California. While I was buying some tulip bulbs he said to me “now put them in the fridge for six weeks before you plant them.” I was so intrigued that I looked it up: it turns out tulips need prolonged cold to flower in spring. I thought the whole ability of plants to monitor seasons was a really fascinating question. No one knew anything about molecular regulation of flowering time, let alone vernalisation, the acceleration of the flowering by cold that is very important for crop plants. Development of winter and spring varieties has significantly extended the geographical range of their production, but there was no molecular understanding of this trait. So I tackled that question using Arabidopsis genetics. I started this project in 1988 and we’re still doing it today!
We started off addressing different angles of the same question. Why do some plants need winter and others don’t? How does the plant actually remember that it has had winter? And how do plants cope with different lengths of winter? These research avenues all converged on a single regulator, a protein that blocks the transition to flowering: FLC. Whether you need winter or not depends on the expression level of that gene. Response to winter depends on epigenetically silencing that gene, and adaptation to different climates involves changes in that silencing mechanism.
The regulation of FLC involves conserved chromatin mechanisms, for example Polycomb silencing, and antisense RNAs. Subtle changes to the anti-sense transcripts are important to set the expression state, silence the gene or adapt to a different type of winter. It is a good system to understand the integration of many different layers of epigenetic regulation, which are often quite hard to dissect in other systems.
What are the next scientific questions that you would like to tackle?
I was lucky to be awarded a European Research Council grant to study how plants monitor and integrate fluctuating winter temperature. Plants integrate temperature changes over several weeks in order to monitor seasonal progression. I want to understand how they do that. What are the actual molecular thermosensors and how is their action channelled and integrated to regulate this one gene? We also want to understand how this process changes as plants adapt to different climates. These questions will be our focus over the next five years.
How has the use of Arabidopsis as a model system changed during your career?
Within the Arabidopsis community, there has been the development of very many extremely useful resources: genome sequences of 1000 natural accessions (there is a whole generation of students and postdocs that can’t remember what it was like doing science without a full genome sequence!); T-DNA collections knocking out all genes… Analysis of complex traits is now much faster in Arabidopsis. The scale of the international community also means you find things out by serendipity: you might think of a gene as flowering-specific and then find its mutation has been found in a completely different screen, which allows you to think of its function in a completely different way.
We need to fight the move for funders to think “let’s fund this reference plant for a bit, get the information we need and then concentrate on crop plants”. For example, flowering time is regulated by a variety of inputs: vernalisation, photoperiodicity, ambient temperature, metabolic signals. We are really quite a long way from understanding how all these fit together. We need to keep funding the basics to improve most rapidly our understanding of this complex adaptive trait – if we focus on flowering analysis in a few reference plants we will then be able to interpret experimental data on flowering in other species much faster. Gerry Fink, a yeast geneticist with an interest in Arabidopsis, made a very controversial statement in 1990: “If you want to understand wheat you should work on Arabidopsis, because four years of work on Arabidopsis will tell us more about wheat than working on wheat for four years.” We are still at the stage where it is very important to aim at a full understanding of how a whole plant works, responds to the environment and adapts. This is best done in reference plant systems at the same time as aiming to effectively translate this information into plant and crop biology as a whole.
At broad developmental biology meetings like this, there’s generally a very low representation of plant science. Would you like to see better integration between the plant and animal fields?
Plant science shouldn’t be seen as a poor relation. We should be very proud of what plant biology has contributed. If you look back at the history of biology, really fundamental discoveries – such as the concept of genetics, chromosomes, transposable elements, heterochromatin, small RNAs – have come from the plant field. Plant science should always be integrated into mainline biology. For example, our work has broad appeal because, although we are looking at a very plant-specific process, we have ended up studying epigenetic pathways that are widely conserved in eukaryotes. However, and this is true of many other communities, plant scientists tend to attend plant-specific meetings and publish in plant-specific journals, and this isolates them. I think it is good to reach beyond your own field. But fields operate using different languages, and this jargon (like the species-specific gene names) complicates comparisons. This is why it is quite good to have joint sessions as in this meeting.
What would people be surprised to find out about you?
I used to sail a lot before my children were born, but in the last few years my life has been a mosaic of family life and work. However, my children have now both gone to university, so I am thinking to myself that I need a hobby! Of course, when you don’t have a hobby what happens is that you work all the time. I am very privileged in that I really enjoy my job and I love being in touch with all that is going on in my lab!


 (4 votes)
(4 votes) The accurate control of gene expression is essential for cell differentiation during development but how do heterogeneous and fluctuating gene expression levels influence cell fate choices? Here (p.
The accurate control of gene expression is essential for cell differentiation during development but how do heterogeneous and fluctuating gene expression levels influence cell fate choices? Here (p. 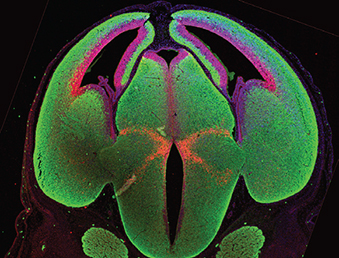


 (No Ratings Yet)
(No Ratings Yet)