Gene regulation and developmental biology at the Royal Society
Posted by Rachael Inglis, on 5 November 2012
Last week, the Royal Society hosted a meeting entitled “Regulation from a distance: Long-range control of gene expression in development and disease”. The impressive London offices of the Society (complete with double helix-inspired door handles) added a sense of occasion to what was bound to be a fascinating meeting, based on the list of excellent speakers from Europe and the US. It brought together a diverse range of scientists from different backgrounds, united by their common interest in the vast stretches of non-coding DNA in the genome, and the role that it plays in gene regulation. I’ll try to give a general overview of the talks here, but with greater focus on the developmental biology topics.
The meeting was opened by Robert Hill (University of Edinburgh) who described an extreme example of a long-range enhancer: the ZRS enhancer that drives Shh expression in the developing limb bud from a distance of 1 Mb. His group have studied this enhancer’s role in preaxial polydactyly (extra digits). The talks that followed showcased a range of useful methods for identifying such distant enhancers.
Francois Spitz (EMBL) described the use of the GROMIT method (Genome Regulatory Organisation Mapping with Integrated Transposons) to mobilise a transposon containing a “regulatory sensor” around the genome, giving reporter gene expression as a read-out of the tissue-specific enhancer signals at different genomic locations. Ivan Ovcharenko (NIH) presented a computational approach for identifying such tissue-specific enhancers, showing that the lessons learned from analysing the tissue-specific transcription factors bound at promoters are also useful for locating enhancers of those tissue-specific genes. Joanna Wysocka (Stanford) presented a combination of different approaches for enhancer identification including conservation analysis, identification of DNaseI hypersensitive sites and ChIP-seq for histone modifications and p300. Her group has used these methods to identify neural crest-specific enhancers that drive novel regulators of craniofacial development.
Several speakers discussed the chromosome conformation capture or ‘C’ techniques, which are used to visualise physical interactions between distant regions of the genome, such as those between gene promoters and enhancers. Denis Duboule (University of Geneva/EPFL) described the use of 4C to investigate the regulation of the Hoxd gene cluster during limb development, and Wouter de Laat (Hubrecht Institute) spoke about recent advances that have increased the resolution of 4C to allow identification of interactions as close as 10 kb. His group have used this technology to identify a novel enhancer of Oct4 in embryonic stem cells. Wendy Bickmore (University of Edinburgh) emphasised the importance of corroborating findings from the ‘C’ techniques using other methods, such as FISH (fluorescent in situ hybridisation).
Although many methods can be used to identify potential enhancers, it is also important to validate their activity. Len Pennacchio (LBNL) described recent efforts in compiling the VISTA enhancer browser: a publicly available database of enhancers that drive tissue-specific gene expression in vivo in transgenic mouse embryos.
In addition to the developmental biology talks, several speakers focused on the role of long-range gene regulation in human disease, which is an important consideration given that many mutations associated with disease are in non-coding regions of the genome. We heard from Doug Higgs (University of Oxford) on the alpha-globin genes and thallasaemias, Matthew Freedman (Harvard) on the Cancer Genome Atlas, and Gioacchino Natoli (European Institute of Oncology) on enhancers that are activated during the inflammatory response.
Enhancer mutations are also an important driving force in evolution, as was explained by David Kingsley (HHMI/Stanford) who studies the rapid natural evolution of three-spined stickleback populations as they colonise new lakes. Major phenotypic changes in these populations can be explained by mutations in a few key developmental genes, but while mutations that inactivate these genes are lethal, mutations in enhancers can alter their expression patterns in a way that produces an evolutionary advantage. Also on the population genetics theme, Manolis Dermitzakis (University of Geneva) presented human data that he has used to investigate the genetic and epigenetic contributions to gene expression.
The majority of speakers focused on long-range gene regulation by enhancers, but it is also important to consider the role of insulators. This was addressed by Bing Ren (UCSD) in the final talk of the meeting. His group have used the HiC technique to create a genome-wide map of chromosomal interactions, revealing that the three-dimensional structure of the genome is divided in to topological domains. Many long-distance interactions occur within these domains, but very few occur between different domains, suggesting that the domain boundaries represent insulators. However, the role of CTCF as an insulator factor was debated at length in the subsequent discussion session.
The Royal Society meetings have an unusual format, with sets of two half-hour talks followed by half an hour of discussion. These long discussion sessions allowed for much deeper probing of the subject, and encouraged debate that became quite lively at times, showing just how exciting this area of research is! Some topics were revisited throughout the meeting, establishing links between different areas, which is important in driving the field forward. The variety of experimental approaches presented showed how insights can be gained by investigating regulation of gene expression at multiple levels. In his talk, John Stamatoyannopoulos (University of Washington) mentioned that the genome contains over 4 million distinct elements that are associated with gene regulation, so there is clearly a lot of work to be done in this field in the future!
The Royal Society have made the audio of all talks from this meeting available online, so check their website for more details on any of the presentations.


 (7 votes)
(7 votes)

 (No Ratings Yet)
(No Ratings Yet)





 What made you decide to study the biology of ageing?
What made you decide to study the biology of ageing?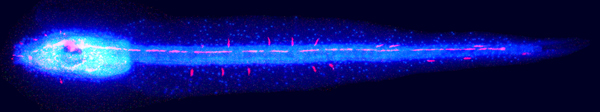
 To mark
To mark  (20 votes)
(20 votes)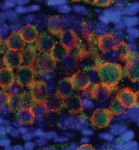 During mammalian gonadal development, somatic cues regulate the sex-specific development of foetal germ cells and control the transition between proliferation and cell-fate commitment. This transition is particularly important for male germ cells: too little proliferation reduces sperm numbers and fertility, whereas escape from commitment and prolonged pluripotency can cause testicular germ cell tumours. Now, Josephine Bowles, Peter Koopman and colleagues (p.
During mammalian gonadal development, somatic cues regulate the sex-specific development of foetal germ cells and control the transition between proliferation and cell-fate commitment. This transition is particularly important for male germ cells: too little proliferation reduces sperm numbers and fertility, whereas escape from commitment and prolonged pluripotency can cause testicular germ cell tumours. Now, Josephine Bowles, Peter Koopman and colleagues (p. 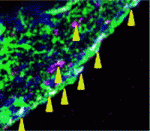 Unlike adult mammals, adult zebrafish can regenerate injured heart tissue. Heart regeneration in zebrafish is known to involve partial de-differentiation and proliferation of cardiomyocytes, but are cardiomyocytes involved in any other processes during heart repair? Here (p.
Unlike adult mammals, adult zebrafish can regenerate injured heart tissue. Heart regeneration in zebrafish is known to involve partial de-differentiation and proliferation of cardiomyocytes, but are cardiomyocytes involved in any other processes during heart repair? Here (p. 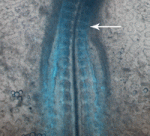 In amniote embryos, three kidneys – the pronephros (a transient embryonic structure), the mesonephros (the embryonic kidney) and the metanephros (the adult kidney) – form sequentially along the anterior-posterior (AP) axis of the intermediate mesoderm (IM). Here (p.
In amniote embryos, three kidneys – the pronephros (a transient embryonic structure), the mesonephros (the embryonic kidney) and the metanephros (the adult kidney) – form sequentially along the anterior-posterior (AP) axis of the intermediate mesoderm (IM). Here (p. 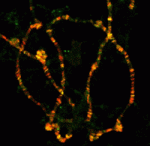 Polycomb group (PcG) genes encode transcriptional repressors that regulate gene expression during development. Most PcG genes encode subunits of chromatin-modifying complexes, but exactly how PcG proteins repress transcription is unclear. Now, Judith Kassis and colleagues report that Wapl, a cohesin-associated protein involved in cohesin removal from chromosomes, promotes PcG silencing in Drosophila (p.
Polycomb group (PcG) genes encode transcriptional repressors that regulate gene expression during development. Most PcG genes encode subunits of chromatin-modifying complexes, but exactly how PcG proteins repress transcription is unclear. Now, Judith Kassis and colleagues report that Wapl, a cohesin-associated protein involved in cohesin removal from chromosomes, promotes PcG silencing in Drosophila (p. 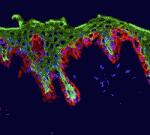 The skin protects the body from microbial pathogens by employing Toll-like receptors (TLRs) and other molecules that recognise pathogen-associated molecular patterns to initiate innate immune responses and to direct subsequent adaptive immunity. But when does the innate immune system in human skin become immunologically competent? On p.
The skin protects the body from microbial pathogens by employing Toll-like receptors (TLRs) and other molecules that recognise pathogen-associated molecular patterns to initiate innate immune responses and to direct subsequent adaptive immunity. But when does the innate immune system in human skin become immunologically competent? On p. 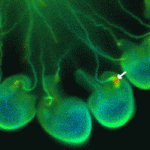 During sexual reproduction in flowering plants, cellular interactions guide the growth of the pollen tube from the stigma to the embryo sac where fertilisation occurs. The cytoplasmic Ca2+ concentration ([Ca2+]cyt) regulates pollen tube growth, but does it also regulate pollen tube guidance and reception? On p.
During sexual reproduction in flowering plants, cellular interactions guide the growth of the pollen tube from the stigma to the embryo sac where fertilisation occurs. The cytoplasmic Ca2+ concentration ([Ca2+]cyt) regulates pollen tube growth, but does it also regulate pollen tube guidance and reception? On p.  In 1991, John Bowman, David Smyth and Elliot Meyerowitz published a paper in Development that proposed the ABC model of flower development. Now, the authors look back on their paper and discuss several aspects of this story.
In 1991, John Bowman, David Smyth and Elliot Meyerowitz published a paper in Development that proposed the ABC model of flower development. Now, the authors look back on their paper and discuss several aspects of this story. In December 1996, in a special issue of Development, 37 papers reported the results of two large screens for zebrafish mutants performed in Tübingen and Boston. Now, Christiane Nüsslein-Volhard gives a personal account of the history of this unique endeavor.
In December 1996, in a special issue of Development, 37 papers reported the results of two large screens for zebrafish mutants performed in Tübingen and Boston. Now, Christiane Nüsslein-Volhard gives a personal account of the history of this unique endeavor.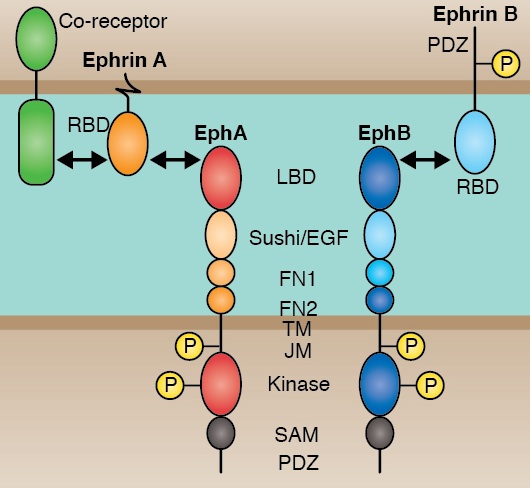 Eph receptors and their membrane-tethered ephrin ligands have important functions in development. Rudiger Klein provides an overview of the general structures and signalling mechanisms underlying Eph/ephrin signalling in development.
Eph receptors and their membrane-tethered ephrin ligands have important functions in development. Rudiger Klein provides an overview of the general structures and signalling mechanisms underlying Eph/ephrin signalling in development.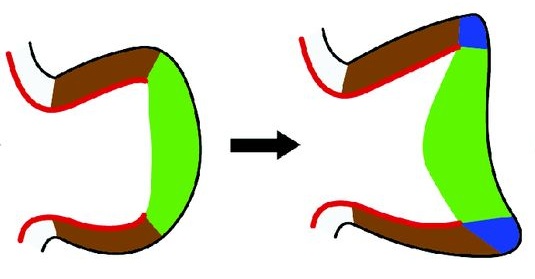 Organ formation during embryogenesis is a complex process that involves various local cell-cell interactions at the molecular and mechanical levels. Yoshiki Sasai and colleagues discuss how, despite this complexity, organogenesis can be modelled in vitro using stem cells.
Organ formation during embryogenesis is a complex process that involves various local cell-cell interactions at the molecular and mechanical levels. Yoshiki Sasai and colleagues discuss how, despite this complexity, organogenesis can be modelled in vitro using stem cells.