deadline tomorrow – EMBO practical course on 3D Developmental Imaging (July 2018 – Portugal)
Posted by gaby, on 10 April 2018
Posted by gaby, on 10 April 2018
Posted by Burkhardt Group, on 10 April 2018
Closing Date: 15 March 2021
There is a vacancy for a postdoctoral research fellow position at the Sars International Centre for Marine Molecular Biology (www.sars.no) in the research group headed by Dr. Pawel Burkhardt. The position is for a period of 4 years and is funded on the Sars Centre core budget. The Sars Centre belongs to the University of Bergen and is partner of the European Molecular Biology Laboratory (EMBL) (www.embl.de). The place of work will be at the Sars Centre. The starting date is negotiable but preferably no later than 01 September 2018.
About the project/work tasks:
The Burkhardt group combines comparative biological systems in the laboratory to understand when and how the first synapses and neurons evolved. The group is particularly interested in studying the origin and evolution of synaptic proteins (Burkhardt et al, 2011 PNAS; Burkhardt et al, 2014 MBE; Bhattacharyya et al, 2016 eLife). We are looking for a highly self-motivated and enthusiastic Postdoctoral Research Fellow with interests in evolutionary biology, neurobiology and cell biology. The project will focus on the characterization of synaptic protein homologs in sponges and ctenophores to better understand the evolution of first neuron-like cell types in animals. The successful candidate will undertake research with the possibility to use a variety of techniques, ranging from super resolution immunofluorescence and electron microscopy, biochemical methods (protein purification, co-IPs, and analytical ultracentrifugation), mass spectrometry to X-ray crystallography to study synaptic protein homologs in sponges and ctenophores. The successful candidate will work in close association with the group leader and other lab members with the aim to eventually contribute to the further development of the project in line with her/his interests.
Qualifications and personal qualities:
About the position of postdoctoral research fellow:
The position of postdoctoral research fellow is a fixed-term appointment with the primary objective of qualifying the appointee for work in top academic positions. The fixed-term period for this position is 4 years. Individuals may not be hired for more than one fixed-term period as a postdoctoral research fellow at the same institution. Upon appointment, applicants must submit a project proposal for the qualifying work including a work schedule. It is a requirement that the project is completed in the course of the period of employment.
We can offer:
Your application in English must include:
The application and appendices with certified translations into English or a Scandinavian language must be uploaded at JobbNorge (https://www.jobbnorge.no/en/available-jobs/job/150623/postdoctoral-research-fellow-positon-evolutionary-origin-of-synapses-and-neurons).
Application Deadline: 02 May 2018.
General information:
Detailed information about the position can be obtained by contacting: Group Leader Pawel Burkhardt, tlf.: +47 55 58 43 57, email: Pawel.Burkhardt@uib.no.
The state labour force shall reflect the diversity of Norwegian society to the greatest extent possible. Age and gender balance among employees is therefore a goal. People with immigrant backgrounds and people with disabilities are encouraged to apply for the position.
We encourage women to apply. If multiple applicants have approximately equivalent qualifications, the rules pertaining to moderate gender quotas shall apply.
The University of Bergen applies the principle of public access to information when recruiting staff for academic positions.
Information about applicants may be made public even if the applicant has asked not to be named on the list of persons who have applied. The applicant must be notified if the request to be omitted is not met.
Further information about our employment process can be found here.
Posted by Burkhardt Group, on 10 April 2018
Closing Date: 15 March 2021
There is a vacancy for a PhD position at the Sars International Centre for Marine Molecular Biology (www.sars.no) in the research group headed by Dr. Pawel Burkhardt. The position is for a fixed-term period of 4 years and is subject to funding on the Sars Centre core budget. The Sars Centre belongs to the University of Bergen and is partner of the European Molecular Biology Laboratory (EMBL) (www.embl.de). The place of work will be at the Sars Centre. The starting date is negotiable but preferably no later than 01 September 2018.
About the project/work tasks:
The goal of the Burkhardt group is to reconstruct the evolutionary origin of synapses and neurons. The group is particularly interested in studying synaptic protein homologs in choanoflagellates, sponges and ctenophores. We are looking for a highly self-motivated and enthusiastic PhD student with interests in evolutionary biology, neurobiology and cell biology. The project will focus on when the protein signalling complexes required for synaptic activity first evolved and how they functioned at a molecular level (Burkhardt et al, 2011 PNAS; Burkhardt et al, 2014 MBE; Bhattacharyya et al, 2016 eLife). The successful candidate will undertake research with the possibility to use a variety of techniques, ranging from super resolution immunofluorescence and electron microscopy, various biochemical methods to X-ray crystallography to study synaptic protein homologs in choanoflagellates. The successful candidate will work in close association with the group leader and other lab members with the aim to eventually contribute to the further development of the project in line with her/his interests.

Qualifications and personal qualities:
About the PhD:
The duration of the PhD position is 4 years, of which 25 per cent of the time each year comprises required duties associated with research, teaching and dissemination of results. The employment period may be reduced if you have previously been employed in a recruitment position.
About the research training:
As a PhD Candidate, you must participate in an approved educational programme for a PhD degree within a period of 4 years. A final plan for the implementation of the research training must be approved by the faculty within three months after you have commenced in the position. It is a condition that you satisfy the enrolment requirements for the PhD programme at the University of Bergen.
We can offer:
Your application in English must include:
The application and appendices with certified translations into English or a Scandinavian language must be uploaded at Jobbnorge (https://www.jobbnorge.no/en/available-jobs/job/150622/phd-position-origin-and-evolution-of-synaptic-proteins)
Application Deadline: 02 May 2018.
General information:
Detailed information about the position can be obtained by contacting: Group Leader Pawel Burkhardt, tlf.: +47 55 58 43 57, email: Pawel.Burkhardt@uib.no
The state labour force shall reflect the diversity of Norwegian society to the greatest extent possible. Age and gender balance among employees is therefore a goal. It is also a goal to recruit people with immigrant backgrounds. People with immigrant backgrounds and people with disabilities are encouraged to apply for the position.
We encourage women to apply. If multiple applicants have approximately equivalent qualifications, the rules pertaining to moderate gender quotas shall apply.
The University of Bergen applies the principle of public access to information when recruiting staff for academic positions.
Information about applicants may be made public even if the applicant has asked not to be named on the list of persons who have applied. The applicant must be notified if the request to be omitted is not met.
The successful applicant must comply with the guidelines that apply to the position at all times.
Posted by the Node Interviews, on 9 April 2018
Cell fate commitment relies on both activation of appropriate genes and suppression of inappropriate ones. Polycomb group proteins are known to be crucial epigenetic silencers of developmental genes, but the manner by which they control fate in vivo, and the relative roles of different Polycomb proteins in silencing, have remained unclear. A new paper in Development tackles this problem using the Drosophila eye a developmental model – we caught up with authors Jinjin Zhu and Justin Kumar, Professor of Biology at Indiana University in Bloomington, to find out more.

JK I started my career in Drosophila eye development while I was an undergraduate in the laboratory of Karl Fryxell at the University of California, Riverside. He was a wonderful mentor and it was my time in his lab that convinced me that I wanted to be a professor one day. While I was in Karl’s lab, I read Don Ready’s seminal paper on the morphogenetic furrow (Ready et al., 1976) and was totally amazed by the cellular mechanism of pattern formation. From the images in the paper, I could see a field of undifferentiated cells being transformed into the periodic units of photoreceptor clusters right before my eyes. I knew then that I wanted to join his lab for my PhD studies. Being in Don’s lab at Purdue University as a graduate student was a privilege. He taught me how to love the fly eye for its own sake and to appreciate its intrinsic beauty.
I then went on to do my post-doctoral fellowship with Kevin Moses initially at the University of Southern California and then at Emory University. It was in Kevin’s lab that I finally settled on the research questions that still drive me today. I stumbled on what I thought to be an astonishing phenotype. Quite by accident I discovered that manipulations of the Notch and EGF Receptor signalling pathways led to the homeotic transformation of the eye into an antenna. While Hox mutants change entire body segments, I was able to observe fate transformations occurring within a single imaginal disc. When I joined the faculty of Indiana University I set out to understand how the fly eye-antennal disc is first set apart from the other discs (i.e. leg, wing, haltere, genital) and then how it is later subdivided into distinct territories such as the compound eye, ocelli, antenna, maxillary palps, and head epidermis. Over the years my research group has discovered that while gene regulatory networks promote fate specification, growth, and patterning, they also influence development by repressing alternate and inappropriate tissue fates. My current interests are to understand how the retinal determination gene regulatory network cooperates with signalling pathways and epigenetic complexes to prevent the eye from adopting distant fates such as the wing and more local fates such as the head epidermis.
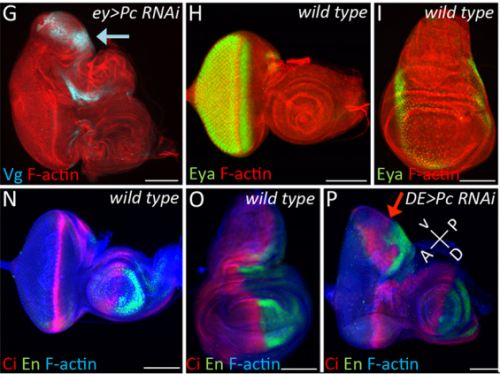
JZ I was really interested in the fate transformation caused by mis-expression of Hox genes when I was in college. When I joined the Justin’s lab, I did a genetic screen to find upstream regulators of eyeless in the developing eye disc. I knocked down eyeless and Sfmbt together and found this amazing eye-to-wing transformation phenotype. Meanwhile, Ali Ordway (second author) joined the lab and decided to screen other PcG proteins. She knocked down Pc and saw similar phenotypes. We were both fascinated by the images of these chimera tissue, in which the dorsal part of the eye disc turned into a wing disc while the ventral part remained as an eye disc, so we decided to figure out what happened to these mutant discs.
JK I got interested in transdetermination purely by accident. When I joined Don’s lab in 1991, I made sure that I read every paper that he had published. In one of his papers, he and Ricky Lebovitz had transplanted eye-antennal disc fragments into host larvae and then recovered the tissue after the host had emerged as an adult (Lebovitz and Ready, 1986). These experiments were aimed at understanding if the morphogenetic furrow was pushed or pulled across the eye field. I was really intrigued by the disc transplantation method used in his paper so I started reading about the history of this method, which I learned was pioneered by Beadle and Ephrussi in the 1930s. In the course of these readings I came across the work of Ernst Hadorn. It was from his papers that I learned about the concepts of determination and transdetermination. When Jinjin and I saw the eye to wing transformation, it reminded me of the fact that Hadorn remarked that the eye could only transdetermine into a wing. And it was at that point that I realized that the loss of PcG and Pax6 that we noticed gave the eye to wing switch might be the molecular explanation for the eye to wing transdetermination event that Hadorn observed many decades ago.
JK In this paper we demonstrate that the Pax6 transcription factor and the Polycomb group (PcG) of epigenetic silencers prevent the eye from adopting a wing fate. This decision is made early in development during the last stages of embryogenesis and during the first larval instar. The eye transforms into a wing because the chromatin around the Antp locus fails to be compacted thereby allowing for its activation by the zinc finger transcription factor Teashirt (Tsh), which is normally expressed in and required for the specification of the eye. Our findings suggest that in addition to promoting the primary fate of a tissue or organ, gene regulatory networks must play defense and suppress the activation of inappropriate selector genes and/or entire gene regulatory networks. This repressive activity appears to require cooperation from epigenetic silencing complexes such as PcG.
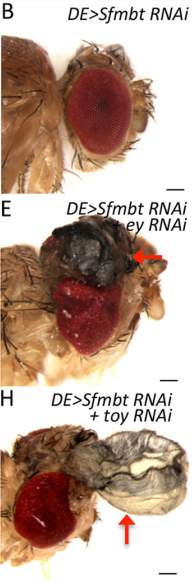
JZ This is a very interesting question and I think the answer lies down to the gene regulatory network controlling dorsal eye disc. 1) Wingless (wg), which is normaly required for wing development, is expressed in a higher level in the dorsal eye disc than in its ventral compartment. 2) engrailed (en) and cubitus interruptus (ci), which regulates A/P compartment of the wing disc also meets in the dorsal portion of the eye disc. Thus, the expression pattern of the endogenous genes in the dorsal eye disc contributes a lot to the eye-to-wing fate transformation. In addition, the dorsal eye disc is more susceptible to adopt wing fate because the level of Pax6 is significantly lower in the dorsal eye disc, especially in the region which will become the future dorsal head capsule. We have demonstrated that eye disc is more resistant to loss of Sfmbt function than the antennal disc. This is very likely due to the presence of Pax6 in the eye progenitor cells, because simultaneously knocking down Pax6 and Sfmbt induces the fate transformation.
JK Yes, it was indeed satisfying to provide a modern perspective to an old problem/observation. In general I really enjoy reading the literature from the pre-molecular age – that is how I got interested in transdetermination in the first place. I also like scour the older literature for problems in eye development that were once studied but later abandoned due to the lack of the right genetic or molecular tools. If you look carefully enough, there is a wealth of such studies out there that are ripe for a modern perspective. For example, one of the exciting new areas of research in my laboratory today is the role that the peripodial epithelium, a tissue that overlies the eye-antennal disc, plays in development. There were several papers in the 1970s that suggested that its role was limited to the fusion of the two eye-antennal discs during pupal development. However, between 2000-2002 several laboratories provided evidence that signalling existed between these two tissues. But very little has been done since on this topic. Now several researchers in my lab are looking at the role that transcriptional networks in the peripodial epithelium play in promoting the fates of the eye-antennal disc.
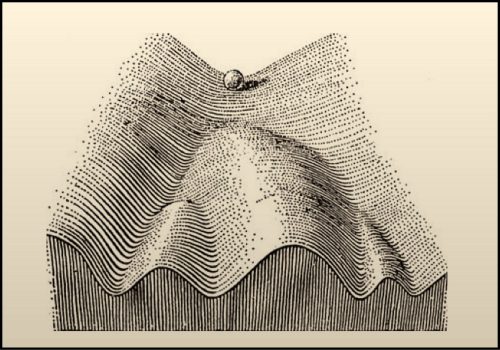
I also like how researchers of times past thought about development. The Epigenetic Landscape Model by C.H. Waddington is one of my favourite examples. To represent the process by which individual cells make fate decisions, he proposed that a cell can take different paths during development with each path representing a journey towards a unique fate. He drew a diagram to represent this idea – in his drawing a ball rolling down a mountainside presented a cell on its way to adopting a final fate. This drawing and the concepts that it represents is very inspiring to me. One can think about how to push cells developmentally down different trails or how to push the cells uphill (de-differentiation) and then down another trail (fate reassignment). For my own work, I try to think of the ball not as a single cell. Instead, to me it represents the entire eye-antennal disc. My lifetime goal is to figure out how the eye-antennal disc is guided down the mountain and how it ultimately gives rise to five distinct tissues and organs.
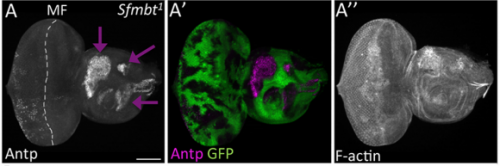
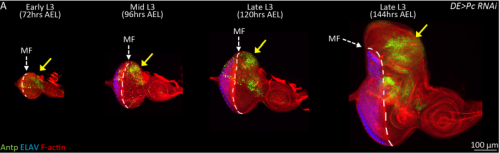
JZ Yes. For a long time, I couldn’t figure out why the eye disc adopts a wing fate but not any other tissue fates. I know it is likely due to the de-repression of Antp in the developing eye when PcG activity is impaired. However, why is Antp being activated when the epigenetic silencers are removed? This question was solved when a piece of data came back from another ongoing project, in which I found that over-expression of teashirt (tsh) was able to rescue the headless phenotype of Pax6 double mutant (Zhu et al, 2017). In those flies, little wings or thoracic bristles were found in the rescued head cuticle, so I wonder the endogenous expression of tsh might be the transcriptional activator of Antp when Pc is removed from the eye disc. I did an experiment to knockdown Pc and Tsh (Figure 3) at the same time and it turn out to be true. Tsh is required to activate Antp during the fate transformation. Although we do not know whether Tsh directly turns on Antp transcription or not, but at least we found some underlying mechanisms of the homology between the eye and the wing. I think the lesson here is to have multiple projects going on at the same time.
JZ For me, the frustration in this project is that isolating eye discs at early stages, such as in 1st and early 2nd larval instar, is almost impossible. Thus, we couldn’t verify our final model of PcG proteins being recruited to the genome at these specific stages using ChIP-Seq. I hope the techniques will advance and allow us to reveal how exactly these epigenetic regulators function in vivo.
JZ Yes, for my postdoctoral training, I will work with Dr. Robert Kingston, focusing on how PcG proteins control gene expression in mammalian system. The PcG proteins in mammals are much more complicated than in Drosophila. Different PcG complexes have multiple variants and each protein have multiple homologs. Thus, a diversity of possible mechanisms that might be used to generated a repressive state of gene expression, such as histone modification, chromatin compaction and higher-order genome organization. I think the eye-to-wing transformation project has brought me here, but I want to learn more about the underlying mechanisms of epigenetic regulators on the molecular level for my future research.
JK My research group has started studying how the interplay between transcriptional networks and epigenetic complexes such as Polycomb, Trithorax, and SAGA controls fate specification within the eye-antennal. These studies build upon the findings of our paper described here in Development in which Pax6 and PcG proteins cooperate to repress wing fates from being adopted in the developing eye. Currently, we have evidence that Pax6 cooperates with Trithorax and SAGA complexes to control the number of antennae that are produced. We also have evidence that these same complexes work together to prevent the duplication of the entire eye-antennal disc. I am very excited about these preliminary findings and I think my lab, for the near future, will be focused on using the tissues within the eye-antennal disc to revisit several very basic questions – how does an organism control the fate, number, and placement of all of the organs that it needs for survival.
JZ I travel with my husband during holidays and we have been to many national parks in US. We both like photographing wild animals. At home, I usually play piano and computer games if I don’t need to collect fly embryos.
JK When I am not in the lab or my office I enjoy playing tennis. It is a terrific sport and a good outlet to release stress. I try to mix tennis and work as well whenever possible. When I am at home working on a paper or grant, I have the Tennis Channel on in the background. If there is an important match, I will stream it on my work computer as well. I also take my camera wherever I go and enjoy photographing wild life and outdoor scenery.
Polycomb group (PcG) proteins and Pax6 cooperate to inhibit in vivo reprogramming of the developing Drosophila eye. Jinjin Zhu, Alison J. Ordway, Lena Weber, Kasun Buddika, Justin P. Kumar.
This is #39 in our interview series. Browse the archive here.
Posted by Eric Lai, on 7 April 2018
Closing Date: 15 March 2021
Eric Lai Lab, Sloan-Kettering, NYC
We have an opening for a postdoctoral position that will address fundamental questions in small RNA biology, genomic conflict, and speciation.
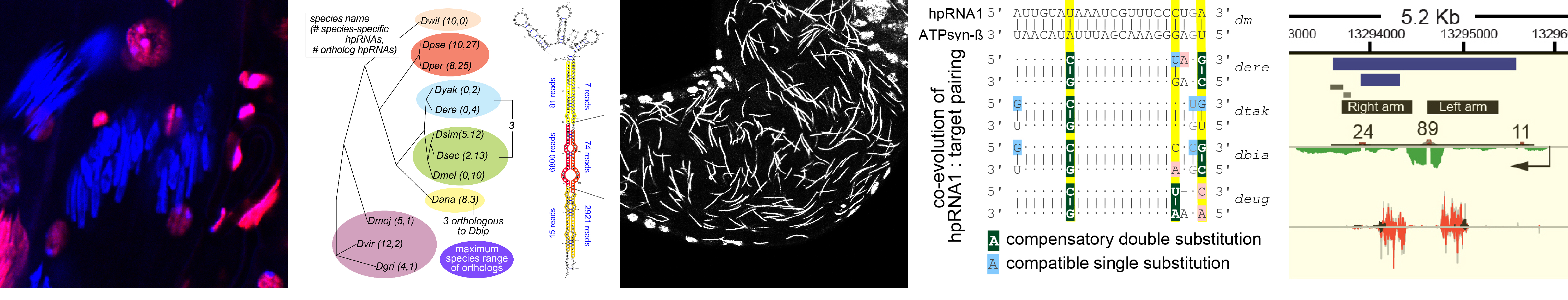
What is endogenous RNAi utilized for? We earlier described a mysterious class of endo-RNAi substrates termed hpRNAs (Okamura Nature 2008), and recently recognized that these mediate adaptive gene regulation in testis (Wen Molecular Cell 2015). These data open a window on a vital biology of RNAi, and now lead us to explore the evolution and function of RNAi systems across the Drosophilid phylogeny.
We discovered networks of rapidly evolving RNAi substrates we hypothesize resolve intragenomic conflicts, and successfully started to use CRISPR/Cas9 in non-model fly species to test some of these ideas (Lin 2018, in submission). Amazingly, while endogenous RNAi was not historically thought to have much phenotypic impact, we find that rapidly evolving genomic battles are being waged and critically depend upon the powerful weapon of RNAi silencing to propagate the species. Reciprocally, we are fascinated to understand by what novel molecular mechanisms de novo selfish meiotic factors can paradoxically drive population extinction.
We seek a motivated postdoctoral fellow with strong Drosophila molecular genetics experience and interest in integrating genome engineering, biochemistry, and bioinformatics to analyze the evolution and function of RNAi silencing systems in resolving deleterious intragenomic conflicts in testis. Although our entry point is RNAi biology, we anticipate that selfish factors yet to be discovered may mediate their effects through chromatin mechanisms. The successful candidate will integrate into a team that is actively engaged in diverse topics in gene regulation in Drosophila and mammalian models, and the Sloan-Kettering Institute provides a vibrant research community.
Funded position with housing and medical benefits are available immediately. Please provide CV, motivation letter and references to Eric Lai, laie@mskcc.org.
Relevant recent papers on hpRNAs and testis post-transcriptional regulation.
see also https://www.mskcc.org/research-areas/labs/eric-lai
Lin, C.-J., F. Hu, R. Dubruille, J. Wen, J. Vedanayagam, P. Smibert, B. Loppin and E. C. Lai (2018). The hpRNA/RNAi pathway is essential to resolve intragenomic conflict to preserve balanced sex ratio. Submitted.
Mohammed, J., A. Flynt, A. Panzarino, M. Mondal, M. DeCruz, A. Siepel and E. C. Lai (2018). Deep experimental profiling of miRNA diversity, deployment, and evolution across the Drosophila genus. Genome Research 28: 52-65.
Kondo S., J. Vedanayagam, J. Mohammed, S. Eizadshenass, L. Kan, N. Pang, R. Aradhya, A. Siepel, J. Steinhauer and E. C. Lai (2017). New genes often acquire male-specific functions but rarely become essential in Drosophila. Genes and Development 31: 1841–1846. (Highlighted in Genes and Dev 31: 1825-1826.)
Lin, C.-J., J. Wen, F. Bejarano, F. Hu, D. Bortolamiol-Becet, L. Kan, P. Sanfilippo, S. Kondo and E. C. Lai (2017). Characterization of a TUTase/RNase complex required for Drosophila gametogenesis. RNA 23: 284-296.
Wen, J., H. Duan, F. Bejarano, K. Okamura, L. Fabian, J. A. Brill, D. Bortolamiol-Becet, R. Martin, J. G. Ruby and E. C. Lai (2015). Adaptive regulation of testis gene expression and control of male fertility by the Drosophila hairpin RNA pathway. Molecular Cell 57: 165-78.
Posted by Pedro P Rocha, on 6 April 2018
Closing Date: 15 March 2021
We are at the National Institute of Child Health and Human Development (NICHD) at NIH, Bethesda, Maryland USA. Our lab is interested in understanding cell lineage differentiation, gene regulation and how non-coding DNA elements and the 3D architecture of chromosomes contribute to these processes during early mouse development.
Learn more at pedrorochalab.org
Who you are:
What we offer:
Apply: Send the following to gsrunit@gmail.com
The NIH is dedicated to building a diverse community in its training and employment programs.
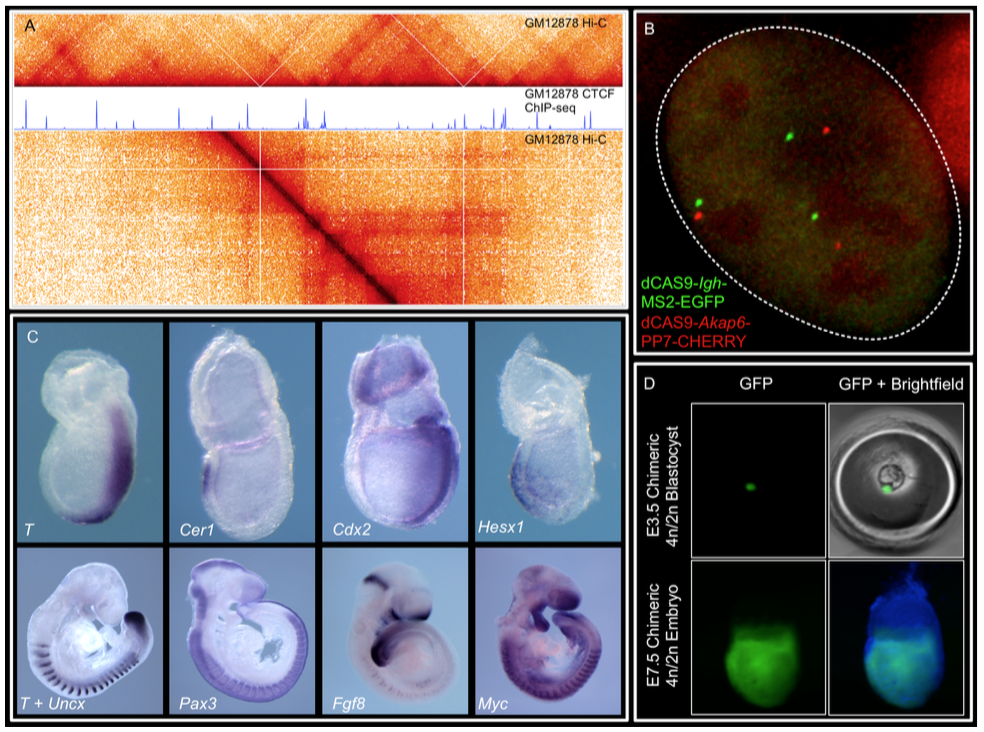
(A) Hi-C and CTCF ChIP- seq of GM1278 cells
(B) dCAS9 MCP-EGFP and PCP-CHERRY live imaging of the Igh and Akap6 loci.
The mouse embryo is an unparalleled system in mammalian biology for understanding how tissue- specific gene expression is achieved.
(C) Whole mount in-situ hybridization for patterning markers in mid and late gastrulating embryos.
(D) Tetraploid aggregation with GFP ES cells allows generation of fully ES-cell derived embryos.
Posted by Joachim Goedhart, on 5 April 2018
In a previous blog, I have disgraced parrots by associating them with P-values and discrediting them for their mechanic repetition. Nevertheless, I admire the vivid colours of these multifaceted birds. Here, I want to make it up by dedicating a pseudo-colour look-up table (LUT) to parrots.
The images produced by fluorescence microscopy are best displayed in grayscale for optimal contrast, using a linear relation between gray-level and pixel intensity (footnote 1). However, the number of gray levels that can be distinguished by humans is orders of magnitude smaller than the number of colours that can be discerned. Therefore, pseudo-colours (also often named false colours) can be used to achieve better contrast and convey a finer level of detail (Sheppard et al., 1968), at the cost of loosing the relation between intensities in the image and the actual pixel value. Thus, the choice between grayscale representation and pseudo-colour will depend on the information that is visualised.
A monochrome digital image can be converted into a pseudo-colour image by replacing each pixel value with a colour (Fink, 1976). The conversion is defined by a so-called look-up table (LUT). The LUT contains 256 RGB codes that define a colour for each of the pixel values between 0 and 255. The combinations are basically endless. A collection of LUTs for cartography, technical illustration and design is available here and the popular image processing software ImageJ/FIJI comes with a set of LUTs.
The LUT that is applied to an image largely depends on the information that one wants to visualise. Since the LUTs are critical as to the level of detail that can be discerned, careful selection is important. Here, I highlight different LUTs in the context of fluorescence images. Besides fluorescence intensity, several other parameters can be encoded by colour. Examples are (i) temporal encoding (figure 1 left), (ii) depth encoding (figure 1 right, and see several other examples here), and (iii) encoding of a spectroscopic parameter, e.g. fluorescence lifetime (Joosen et al., 2014).
Figure 1: Pseudo-colours are used (in the left panel) to encode time to show the dynamics of microtubule plus ends or (in the right panel) to encode depth in a 3D STORM image of microtubules. Image credits: Timelapse movie of EB3-mNeonGreen by Anna Chertkova, 3D STORM image by Christophe Leterrier.
Here, I will give examples of pseudo-coloured images that were obtained by FRET ratio-imaging. In FRET ratio-imaging the response of a biosensor is monitored by emission ratio-imaging. Both the spatial differences and the changes over time can be of interest. The spatial differences are usually presented with pseudo coloured maps of the FRET ratio. Movies of these FRET maps are generated to depict the ratio changes over time.
The ‘Fire’ LUT that comes with ImageJ works well for temporal colour coding (figure 1, left panel). A variant of the Fire LUT was generated and named ‘MorgenStemning’. This LUT is colourblind friendly and has a linear increase in luminance (Geissbuehler and Lasser, 2013). Similarly, several “perceptually uniform” colour palettes were generated that are also colourblind friendly. These LUTs, ‘Inferno’, ‘Viridis’, ‘Magma’ & ‘Plasma’, are described here. The Inferno LUT is a good choice, since it starts with black at zero intensity. This LUT does, however, not use white for maximal intensity, thereby not providing maximal contrast.
Some of the LUTs that we previously used are the ‘royal’ LUT of ImageJ (Unen et al., 2015) and a pseudocolor LUT that is part of MetaMorph software (Reinhard, 2016). Since I was not entirely satisfied with the existing LUTs, I designed a new LUT (footnote 2). The basic features are that it starts with black and ends with white to achieve maximal contrast between the minimal and maximal pixel values. The false colours that correspond with lower half of the pixel values does not have a red component to get a cyan colour which blends into bright green. For higher pixels values the false colours turn from green via orange and red to white. As the colours in the LUTs reminded me of parrots, I call this LUT ‘parrot’. Figure 2 shows how the parrot LUT compares to other LUTs. The parrot LUT is colour-blind-friendly (footnote 3), but it is not perceptually uniform and is suboptimal in this respect.
Figure 2: An overview of different LUTs applied to the same grayscale image that displays Rho GTPase activity in endothelial cells (Reinhard, 2017). The image and LUTs are available here.
Final words
The choice of a LUT is often based on a qualitative assessment of how it visualises the information. By trying a number of LUTS, the LUT that best conveys the information can be selected. Therefore, it useful to have a broad panel of LUTs to choose from and the parrot LUT is just another option. I encourage designing and sharing of LUTs (either as supplemental data to go with a paper or on data sharing platforms – footnote 4) to increase the number of options and the chances of finding the right LUT.
Acknowledgments: Thanks to Jakobus van Unen for tracing back the origin of the ‘pseudo-MM’ LUT to metamorph and to Marten Postma for making the LUT editor available.
Footnotes
Footnote 1: Achieving a true linear relationship between pixel value and displayed grayscale value may be impossible and irrelevant.
Footnote 2: The parrot LUT was created with a MatLab based LUT editor that was generated by Marten Postma
Footnote 3: I have a colour vision deficiency that prevents me from distinguishing blue from purple and bright green from yellow. This is the main reason to remove most of the red component and only have a limited band with a green/yellow component. Since I designed it, it is intrinsically colour-blind friendly. However, as there are several types of colour vision deficiency it may not work for everyone.
Footnote 4: The LUTs and related material described in this blog are available at Zenodo (Doi: 10.5281/zenodo.1211690).
.
Posted by the Node Interviews, on 5 April 2018
Development and homeostasis depend crucially on the maintenance of cell identity, and in gamete-producing tissues the somatic/germline distinction is paramount. A recent paper in Development explores how cell identity is secured in the Drosophila ovary by studying the function of the conserved tumour suppressor L(3)mbt. To find out more about the story, we caught up with first author Rémi-Xavier Coux and his supervisor Ruth Lehmann of the Skirball Institute at New York University School of Medicine.

RL I grew up in Germany and received my PhD in the lab of Christiane Nuesslein-Volhard. I was in her lab at the very beginning when the genes required for establishing embryonic polarity and pattern were identified. My project involved the genetic analysis of gap genes and a group of maternal effect genes, we termed the posterior group of genes, as mutation in these genes affect the development of the embryonic abdomen and my cytoplasmic transplantation experiments suggested a gradient of patterning activity emanating from the posterior pole. I became intrigued by this posterior pole plasm as it is also the site of germ cell formation in Drosophila. So, I made the more unusual and often discouraged decision to stay with my graduate project, yet decided to delve into molecular analysis of these genes. By the time I arrived for my first independent position at the Whitehead Institute at MIT in 1988, I had molecular entry points for the identification of the nanos, oskar and pumilio genes in my suitcase.
Analysis of these genes was really exciting as it showed that the mRNAs encoding Oskar and Nanos were localized to the posterior pole and that the translation of these RNAs was specifically regulated such that only the localized RNA (and now we know that is only a very small fraction of the total RNA) is translated while unlocalized RNA is translationally repressed. From the beginning, I have been amazingly lucky to have incredibly talented graduate students and postdocs in my lab, not only did they make all these discoveries but they also taught me a lot.
In 1996, I was recruited to the newly founded Skirball Institute at NYU School of Medicine. Here my lab focused completely on germ cell biology studying the germ line life cycle. We are particularly interested in three areas:
1. How the germ line-soma dichotomy is initially established in the early embryo. Critical for the initial distinction between soma and germ line are the properties of membraneless germ granules that co-ordinate the posttranscriptional regulation of RNAs specifically needed for germ cell formation, specification, transcriptional silencing and germ cell migration to the somatic gonad. Crucial for progress here has been the development of ever so powerful imaging modalities (from light sheet microscopy and super resolution microscopy to electron microscopy) and the development of many different ways to mark molecules and observe them in vivo.
2. How interactions between cells from different origins coordinate growth and differentiation of the gonad so that primordial germ cells mature into germ line stem cells and deposit eggs during adult life. Initially, we relied on forward genetic screens to address this big and fascinating problem of organ development but now we are increasingly take advantage of whole genome genetic analysis by RNAi and Crispr/Cas9 as well as high-throughput molecular analysis such as single cell sequencing at different developmental stages.
3. Finally, we are interested in how the unique role of germ cells as the only cells of the body with the potential to give rise to a complete new organism manifests specialized adaptions. So, we have become intrigued by the broadest sense of mutual ‘host-pathogen’ interactions as they relate to the germ line. Here, we are interested in how germ line regulatory mechanisms manage to control transposable elements activity, the interplay between genome and mitochondria during the germ line life cycle and emerging optional relationships like the ability of the intracellular bacterium Wolbachia to grow in the Drosophila host without direct harm. We reason, that such germline specific control and defence mechanisms protect the germ line, but may also provide opportunity for species to evolve via changes to the germline.
RXC l(3)mbt was identified in the laboratory of Elisabeth Gateff, who was the first to use Drosophila to identify tumor-suppressor genes (Gateff, 1978, Gateff et al., 1993). As the name says, ‘l(3) malignant brain tumor’ mutations cause brain tumors in Drosophila larvae. Many years later, Chris Yohn, a postdoc in our lab, identified several alleles of l(3)mbt in a clonal screen for maternally expressed genes that affected germ cell formation in the embryo. While Chris determined that the PGC formation defect was a secondary consequence of L(3)mbt’s role in early embryonic nuclear divisions, he also observed that l(3)mbt females were viable at the permissive temperature (brain tumors form only at high temperature) but they were completely sterile and produced no eggs. This suggested an additional role for L(3)mbt in gonadogenesis (Yohn et al., 2003). The project rested for a while, until Cayetano Gonzalez and his group at the IRB in Barcelona published an intriguing observation. The Gonzalez lab profiled l(3)mbt larval brain tumors and showed that in these tumors many germline genes were apparently derepressed (Janic et al., 2010). We were intrigued by this possible soma-to-germline transformation, so when I joined the lab as a graduate student I started two projects on l(3)mbt. First, I asked whether l(3)mbt brain tumor cells indeed behaved like bona fide germ cells and could be used to identify novel regulators of the germline fate. Second, I wanted to test whether the sterility phenotype was also due to a soma-to-germline transformation. Our paper describes the results of the later study that proved much more successful than the former.
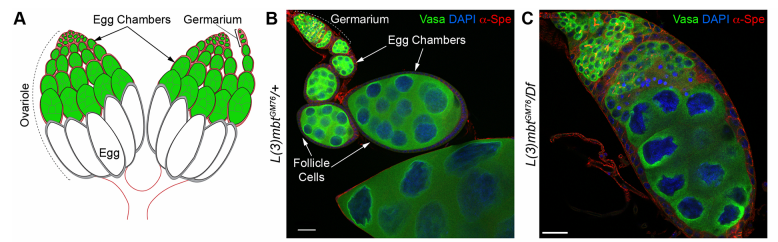
RL&RXC In addition to its role in suppressing brain tumors in the larvae, L(3)mbt functions in two tissues of the ovary to safeguard oogenesis: the somatic support cells and the germline. We found that in both tissues, L(3)mbt prevents expression of genes incompatible with normal development: in the somatic ovarian cells, L(3)mbt represses genes normally expressed in the germline while in the germ cells, it silences testis and neuronal genes. This, and the fact that l(3)mbt mutant tissues still express genes characteristic of the tissue of origin revealed a function broader than previously thought. Our study therefore suggests that L(3)mbt functions as a tissue specific transcription repressor rather than simply silencing germline genes in somatic tissues. L(3)mbt binding sites overlap with insulator elements (Richter et al., 2012) so it is also possible that L(3)mbt functions in insulator complexes.
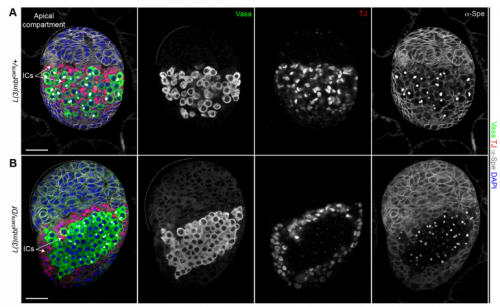
RL&RXC Indeed, L(3)mbt has been found to function with the dREaM/MMB and LINT complexes in S2 and Kc167 somatic embryonic cells (Georlette et al., 2007; Meier et al., 2012). However, our genetic studies suggest that L(3)mbt functions independently of dREaM in the ovary. Several components of the dREaM complex are required for endo-replication (polyploidization) of both somatic ovarian cells and nurse cells in the germline, but these processes seem not to require L(3)mbt function. It would be interesting to test if L(3)mbt functions in only one of these two chromatin complexes in other tissues besides the ovary and how the potential switch between complexes is regulated.
RL&RXC This is a conceptual question we really struggled with: the “orthodox” definition of transdifferentiation is complete acquisition of another cell identity. However, functionally testing fate switching in vivo is very challenging. Jarrault and colleagues beautifully showed that in C. elegans the Y epithelial cell transdifferentiates into a fully functional neuron (PDA) in wild-type larvae (Jarrault et al., 2008). It is one of the only examples of complete transdifferentiation to our knowledge. There are many more examples, especially with mutations in chromatin factors, where cell-specific gene signatures are misexpressed. At the end, it comes down to the assay that is used to define a ‘cell fate switch’. For example, can we call a cell that aberrantly expresses most of another cell type’s transcriptome trans-differentiated without a functional assay?
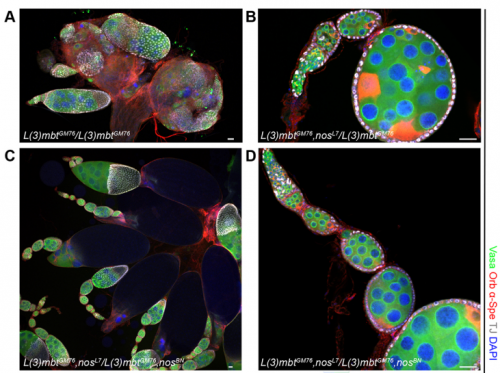
RXC Yes, I was very glad to observe that nanos mutations suppress the l(3)mbt mutant ovarian phenotypes. It was late at the confocal and I immediately emailed Ruth. The morning after, we checked that embryos laid by l(3)mbt, nos double mutant females had the typical patterning defects caused by nos mutations. When this control was done, we were really excited!!!
RXC I tried transplanting l(3)mbt tumorous brain cells into embryos devoid of germline to test if they could, at least partially, behave as germ cells. This experiment was quite challenging, and we could not detect the cells a few hours after transplantation, which was very frustrating! We now know that this experiment likely did not work because these neural-origin cells may not be completely transformed.
RXC I just started a postdoc in the Cohen-Tannoudji and Navarro-Gil labs in the Stem Cell Biology and Development Department, Pasteur Institute in Paris. I will study and characterize Transcription Factor bookmarking in the early mouse embryo.
RL Further study of L(3)mbt may provide us with clues about the transcriptional mechanisms underlying germline-soma dichotomy. We found in our study that mutating a key regulator of germline fate, the translational repressor Nanos, suppresses the somatic gonadal defects of l(3)mbt mutant ovaries almost completely. Thus, Nanos targets may be key regulators that distinguish between the germline and soma program and the l(3)mbt mutants may guide us towards their identification.
RXC In my free time, I enjoy music, modern art, playing rugby and outdoor activities such as sailing and mountain activities.
RL I like hiking with my Aussie Luke.
Rémi-Xavier Coux, Felipe Karam Teixeira, Ruth Lehmann. 2018. L(3)mbt and the LINT complex safeguard cellular identity in the Drosophila ovary. Development 2018 145: dev160721
This is #38 in our interview series. Browse the archive here
Posted by frankschnorrer, on 4 April 2018
Closing Date: 15 March 2021
Muscle building: bridging molecular order to macroscopic morphogenesis
PHD2018-11 (PDF link)
Posted by the Node, on 4 April 2018
One of the most talked about preprints this month was a corrigendum to that Nature Methods paper reporting widespread off-target mutations following CRISPR-Cas9 editing in mice. The paper was challenged quickly on bioRxiv, and four days after the corrigendum went up, it was retracted, as reported by Retraction Watch! Blink and you’ll miss it.
Otherwise this month we found plenty of plant patterning, lovely larval lampreys and ovulating roundworms, and the developing diaphragm finding its voice.
The preprints were hosted on bioRxiv, PeerJ, and arXiv. Use these links to get to the section you want:
| Stem cells, regeneration & disease modelling
Positional information encoded in the dynamic differences between neighbouring oscillators during vertebrate segmentation.
Marcelo Boareto, Tomas Tomka, Dagmar Iber
Checkpoint signaling abrogation after cell cycle reentry reveals that differentiated neurons are mitotic cells
Chaska Carlos Walton, Wei Zhang, Iris Patiño-Parrado, Estíbaliz Barrio-Alonso, Juan-José Garrido, José María Frade
Epidermal YAP Activity Drives Canonical WNT16/β-catenin Signaling to Promote Keratinocyte Proliferation in vitro and in the Murine Skin
Veronica Mendoza-Reinoso, Annemiek Beverdam
Extracellular Pax6 regulates tangential Cajal-Retzius cell migration in the developing mouse neocortex
Hadhemi Kaddour, Eva Coppola, Ariel Di Nardo, Andrea Wizenmann, Michel Volovitch, Alain Prochiantz, Alessandra Pierani
Megan B O’Hare, Alamin Mohammed, Kyle J Connolly, Katelyn C Aitchison, Niki C Anthoney, Amy L Roberts, Matthew J Taylor, Bryan A Stewart, Richard I Tuxworth, Guy Tear
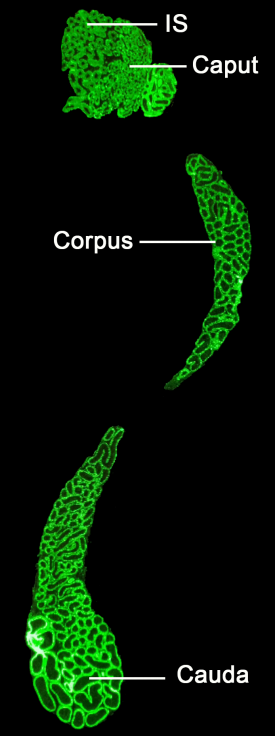
Cell-lineage specificity of primary cilia during epididymis post-natal development
Agathe Bernet, Alexandre Bastien, Denis Soulet, Olivia Jerczynski, Christian Roy, Maira Bianchi Rodrigues Alves, Cynthia Lecours, Marie-Eve Tremblay, Janice Bailey, Claude Robert, Clemence Belleannee
Hideki Katow, Hiromi Yoshida, Tomoko Katow, Masato Kiyomoto
Embryonic and postnatal neurogenesis produce functionally distinct subclasses of dopaminergic neuron
Elisa Galliano, Eleonora Franzoni, Marine Breton, Annisa N. Chand, Darren J. Byrne, Venkatesh N. Murthy, Matthew S. Grubb
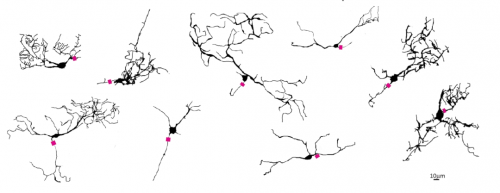
Formation of retinal direction-selective circuitry initiated by starburst amacrine cell homotypic contact
Thomas A Ray, Suva Roy, Christopher Kozlowski, Jingjing Wang, Jon Cafaro, Samuel Hulbert, Christopher V E Wright, Greg D Field, Jeremy N Kay

Conservation and Divergence of YODA MAPKKK Function in Regulation of Grass Epidermal Patterning
Emily Abrash, Maria Ximena Anleu Gil, Juliana L Matos, Dominique C Bergmann
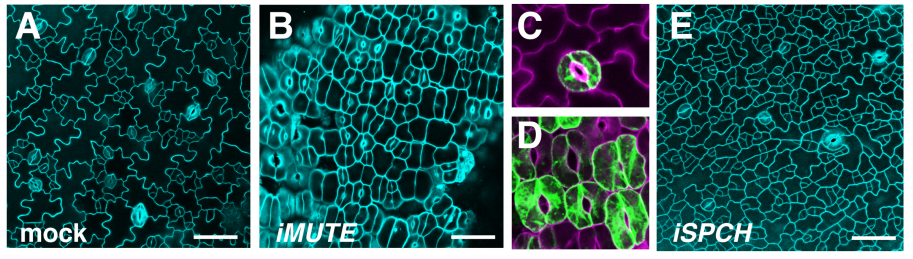
MUTE Directly Orchestrates Cell State Switch and the Single Symmetric Division to Create Stomata
Soon-Ki Han, Xingyun Qi, Kei Sugihara, Jonathan Dang, Takaho A Endo, Kristen Miller, Eun-Deok Kim, Takashi Miura, Keiko Torii
Clonal analysis reveals gradual recruitment of lateral root founder cells and a link between root initiation and cambium formation in Arabidopsis thaliana
Joseph G. Dubrovsky
Auxin is not asymmetrically distributed in initiating Arabidopsis leaves
Neha Bhatia, Marcus G Heisler
A Lotus japonicus E3 ligase interacts with the Nod factor receptor 5 and positively regulates nodulation
Daniela Tsikou, Estrella E. Ramirez, Ioanna S Psarrakou, Jaslyn E Wong, Dorthe B Jensen, Erika Isono, Simona Radutoiu, Kalliope K Papadopoulou
Pollen Patterns Form from Modulated Phases
Asja Radja, Eric M. Horsley, Maxim O. Lavrentovich, Alison M. Sweeney
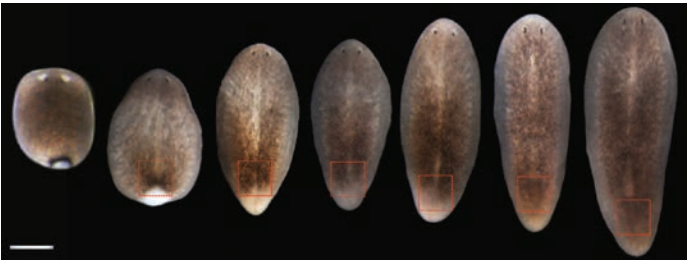
A nuclear hormone receptor and lipid metabolism axis are required for the maintenance and regeneration of reproductive organs
Shasha Zhang, Longhua Guo, Carlos Guerrero-Hernández, Eric J Ross, Kirsten Gotting, Sean A. McKinney, Wei Wang, Youbin Xiang, R. Scott Hawley, Alejandro Sánchez Alvarado
PIP5k1 β controls bone homeostasis through modulating both osteoclast and osteoblast differentiation
Xiao-ying Zhao, Guoli Hu, Chuandong Wang, Lei Jiang, Jingyu Zhao, Jiake Xu, Xiaoling Zhang
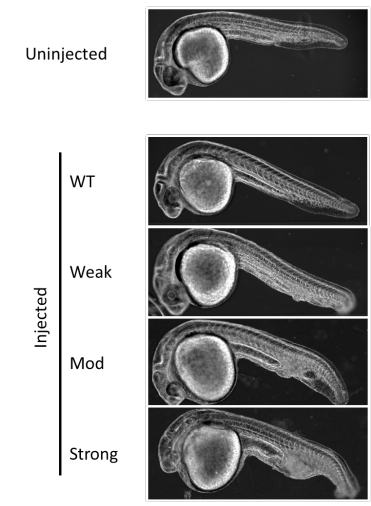
Analysis of novel domain-specific mutations in the zebrafish ndr2/cyclops gene generated using CRISPR-Cas9 RNPs
Ashley N Turner, Reagan S Andersen, Ivy E Bookout, Lauren N Brashear, James C Davis, David M Gahan, John P Gotham, Baraa A Hijaz, Ashish S Kaushik, Jordan B McGill, Victoria L Miller, Zachariah P Moseley, Cerissa L Nowell, Riddhi K Patel, Mia C Rodgers, Yazen A Shihab, Austin P Walker, Sarah R Glover, Samantha D Foster, Anil Kumar Challa
Deep proteomic analysis of chicken erythropoiesis
Marjorie Leduc, Emilie-Fleur Gautier, Anissa Guillemin, Cédric Broussard, Virginie Salnot, Catherine Lacombe, Olivier Gandrillon, François Guillonneau, Patrick Mayeux
PRY-1/AXIN signaling regulates lipid metabolism in Caenorhabditis elegans
Ayush Ranawade, Avijit Mallick, Bhagwati P Gupta
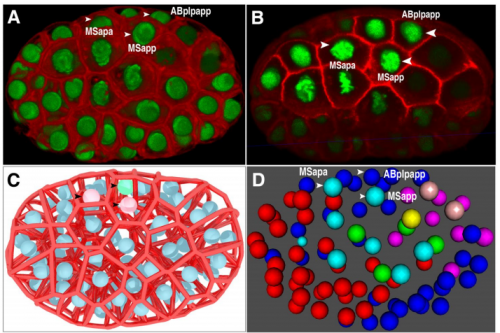
Establishment of signaling interactions with cellular resolution for every cell cycle of embryogenesis
Zhongying Zhao, Long Chen, Vincy WS Ho, Ming-Kin Wong, Xiaotai Huang, Lu-yan Chan, Kaoru HC Ng, Xiaoliang Ren, Hong Yan
Murine SEC24D Can Substitute Functionally for SEC24C in vivo
Elizabeth J. J. Adams, Rami Khoriaty, Anna Kiseleva, Audrey C.A. Cleuren, Kart Tomberg, Martijn A. van der Ent, Peter Gergics, K. Sue O Shea, Thomas L. Saunders, David Ginsburg
E93 expression and links to the juvenile hormone in hemipteran mealybugs with insights on female neoteny
Isabelle M Vea, Sayumi Tanaka, Tomohiro Tsuji, Takahiro Shiotsuki, Akiya Jouraku, Chieka Minakuchi
Anisotropic Crb accumulation, modulated by Src42A, orients epithelial tube growth in Drosophila
Ivette Olivares-Castineira, Marta Llimargas
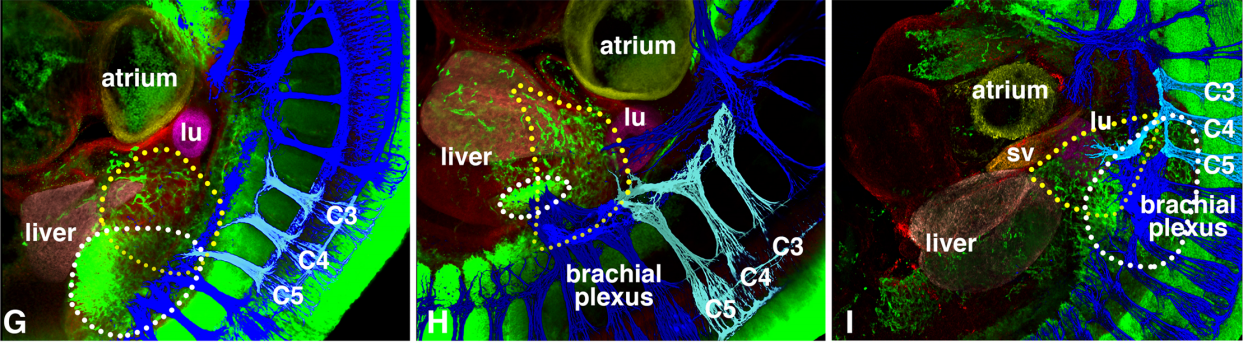
Developmental origin and morphogenesis of the diaphragm, an essential mammalian muscle
Elizabeth M Sefton, Mirialys Gallardo, Gabrielle Kardon
Spatial and temporal PCP protein dynamics coordinate cell intercalation during neural tube closure
Mitchell Butler, John B Wallingford
The nidogen-domain containing protein DEX-1 is required for epidermal remodeling in C. elegans dauers
Kristen Flatt, Caroline Beshers, Cagla Unal, Nathan Schroeder
Intermediate filaments EXC-2 and IFA-4 Maintain Luminal Structure of the Tubular Excretory Canals in Caenorhabditis elegans
HIkmat Imad Al-Hashimi, David H. Hall, Brian D Ackley, Erik A. Lundquist, Matthew Buechner
Bayesian Detection of Abnormal ADS in Mutant Caenorhabditis elegans Embryos
Wei Liang, Yuxiao Yang, Yusi Fang, Zhongying Zhao, Jie Hu
Retinal Ganglion Cell Axon Sorting at the Optic Chiasm Requires Dystroglycan
Reena Clements, Kevin M Wright
Protocadherin-mediated cell repulsion controls the central topography and efferent projections of the abducens nucleus
Kazuhide Asakawa, Koichi Kawakami
4D imaging reveals stage dependent random and directed cell motion during somite morphogenesis
James McColl, Gi Fay Mok, Anna H Lippert, Alex Ponjavic, Leila Muresan, Andrea Munsterberg
Systematic studies of all PIH proteins in zebrafish reveal their distinct roles in axonemal dynein assembly
Hiroshi Yamaguchi, Toshiyuki Oda, Masahide Kikkawa, Hiroyuki Takeda
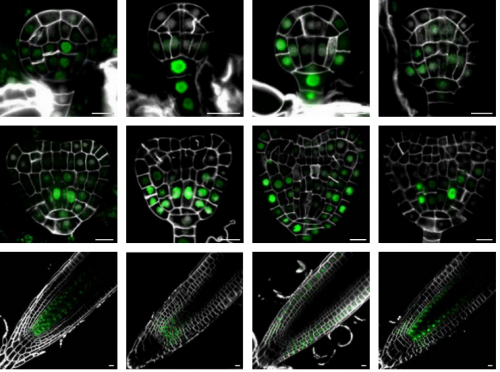
IQD proteins integrate auxin and calcium signaling to regulate microtubule dynamics during Arabidopsis development
Jos Wendrich, Bao-Jun Yang, Pieter Mijnhout, Hong-Wei Xue, Bert De Rybel, Dolf Weijers
OsIQD14 regulates rice grain shape through modulating the microtubule cytoskeleton
Bao-Jun Yang, Jos R. Wendrich, Bert De Rybel, Dolf Weijers, Hong-Wei Xue
A novel cell segmentation method for developing embryos using machine learning
Rikifumi Ota, Takahiro Ide, Tatsuo Michiue
A genome-wide assessment of the ancestral neural crest gene regulatory network
Dorit Hockman, Vanessa Chong-Morrison, Daria Gavriouchkina, Stephen Green, Chris Amemiya, Jeramiah J Smith, Marianne Bronner, Tatjana Sauka-Spengler
A post-transcriptional regulatory code for mRNA stability during the zebrafish maternal-to-zygotic transition
Charles E Vejnar, Mario Abdel Messih, Carter M Takacs, Valeria Yartseva, Panos Oikonomou, Romain Christiano, Marlon Stoeckius, Stephanie Lau, Miler T Lee, Jean-Denis Beaudoin, Hiba Darwich-Codore, Tobias C Walther, Saeed Tavazoie, Daniel Cifuentes, Antonio J Giraldez
mRNA structure dynamics identifies the embryonic RNA regulome
Jean-Denis Beaudoin, Eva Maria Novoa, Charles E Vejnar, Valeria Yartseva, Carter Takacs, Manolis Kellis, Antonio J Giraldez
Dosage sensitivity of X-linked genes in human embryonic single cells
Xiaoshu Chen, Jian-Rong Yang
Two Active X-chromosomes Modulate the Growth, Pluripotency Exit and DNA Methylation Landscape of Mouse Naive Pluripotent Stem Cells through Different Pathways
Juan Song, Adrian Janiszewski, Natalie De Geest, Lotte Vanheer, Irene Talon, Taeho Oh, Vincent Pasque
A molecular network for functional versatility of HECATE transcription factors
Christophe Gaillochet, Suraj Jamge, Froukje van der Wal, Gerco Angenent, Richard Immink, Jan U. Lohmann
P-bodies and the miRNA pathway regulate translational repression of bicoid mRNA during Drosophila melanogaster oogenesis
John M. McLaughlin, Daniel FQ Smith, Irina E. Catrina, Diana P. Bratu
Bicoid gradient formation mechanism and dynamics revealed by protein lifetime analysis
Lucía Durrieu, Daniel Kirrmaier, Tatjana Schneidt, Ilia Kats, Sarada Raghavan, Michael Knop, Timothy E Saunders, Lars Hufnagel
Novel approach to quantitative spatial gene expression uncovers genetic stochasticity in the developing Drosophila eye
Sammi Ali, Sarah Signor, Konstantin Kozlov, Sergey Nuzhdin
Embryonic gene transcription in the spiny mouse (Acomys cahirinus): an investigation into the embryonic genome activation.
Jared Mamrot, David K Gardner, Peter Temple-Smith, Hayley Dickinson
Genetically regulated human NODAL splice variants are differentially post-transcriptionally processed and functionally distinct
Scott D Findlay, Olena Bilyk, Kiefer Lypka, Andrew J Waskiewicz, Lynne-Marie Postovit
A critical role for miR-142 in alveolar epithelial lineage formation
Amit Shrestha, Carraro Gianni, Nicolas Nottet, Ana Ivonne Vasquez-Armendariz, Susanne Herold, Julio Cordero, Indra Bahadur Singh, Jochen Wilhelm, Guillermo Barreto, Cho-Ming Chao, Elie El Agha, Bernard Mari, Jin-San Zhang, Saverio Bellusci
Small non-coding RNA expression in developing mouse nephron progenitor cells
Yu Leng Phua, Andrew Clugston, Kevin Hong Chen, Dennis Kostka, Jacqueline Ho
Temporal Control of Transcription by Zelda in living Drosophila embryos
Jeremy Dufourt, Antonio Trullo, Jennifer Hunter, Carola Fernandez, Jorge Lazaro, Matthieu Dejean, Lucas Morales, Katharine N Schulz, Melissa M Harrison, Ovidiu Radulescu, Cyril Favard, Mounia Lagha
MicroRNA-202 (miR-202) controls female fecundity by regulating medaka oogenesis
Stephanie Gay, Jerome Bugeon, Amine Bouchareb, Laure Henry, Jerome Montfort, Aurelie Le Cam, Julien Bobe, Violette Thermes
Lost in translation: egg transcriptome reveals molecular signature to predict developmental success and novel maternal-effect genes
Caroline T Cheung, Thaovi Nguyen, Aurelie Le Cam, Amelie Patinote, Laurent Journot, Christelle Reynes, Julien Bobe
Transcriptome Landscape of Human Oocytes and Granulosa Cells Throughout Folliculogenesis
Yaoyao Zhang, Zhiqiang Yan, Qingyuan Qing, Vicki Nisenblat, Yang Yu, Tianren Wang, Cuiling Lu, Ming Yang, Shuo Yang, Ying Yao, Xiaohui Zhu, Xi Xia, Yujiao Dang, Yixin Ren, Peng Yuan, Rong Li, Ping Liu, Hongyan Guo, Jinsong Han, Haojie He, Yu Wu, Meng Li, Kun Zhang, Yiting Wang, Jie Qiao, Jie Yan, Liying Yan
Numerous recursive sites contribute to accuracy of splicing of long introns in flies
Athma A Pai, Joseph Paggi, Karen Adelman, Christopher B Burge
Ldb1 mediates trans enhancement in mammals
Kevin Monahan, Adan Horta, Alice Mumbay-Wafula, Liqi Li, Yangu Zhao, Paul Love, Stavros Lomvardas
Down-regulated Long Noncoding RNA HOXA11-AS affects trophoblast cell proliferation and migration by regulating RND3 and HOXA7 expression in preeclampsia
Lizhou Sun, Yetao Xu, Dan Wu, Jie Liu, Zhonghua Ma, Bingqing Hui, Jing Wang, Yanzi Chen, Sailan Wang, Yifan Lian
Chromatin accessibility dynamics across C. elegans development and ageing
Jurgen Janes, Yan Dong, Michael Schoof, Jacques Serizay, Alex Appert, Chiara Cerrato, Carson Woodbury, Ron Chen, Carolina Gemma, Ni Huang, Djem Kissiov, Przemyslaw Stempor, Annette Steward, Eva Zeiser, Sascha Sauer, Julie Ahringer
The tumor suppressor BRCA1/BARD1 complex localizes to the synaptonemal complex and regulates recombination under meiotic dysfunction in Caenorhabditis elegans
Qianyan Li, Takamune T Saito, Alison J Deshong, Marina Martinez Garcia, Saravanapriah Nadarajan, Katherine S Lawrence, Paula M Checchi, Monica P Colaiacovo, JoAnne Engebrecht
The BRCA1-BARD1 complex associates with the synaptonemal complex and pro-crossover factors and influences RAD-51 dynamics during Caenorhabditis elegans meiosis
Eva Janisiw, Maria Rosaria Dello Stritto, Verena Jantsch, Nicola Silva
Previously characterized Caenorhabditis elegans RNA interference-resistant mutants retain substantial silencing capacity
Jennifer L Watts, Jason S Watts, James Dalelio, Quentin Guenther
Synaptonemal complex architecture facilitates the chromosome-specific regulation of recombination in Drosophila
Cori Cahoon, G. Matthew Heenan, Zulin Yu, Jay R Unruh, Sean McKinney, R. Scott Hawley
Evolutionary analysis implicates RNA polymerase II pausing and chromatin structure in nematode piRNA biogenesis
Toni Beltran, Consuelo Barruso, Timothy Birkle, Lewis Stevens, Hillel T Schwartz, Paul Sternberg, Helene Fradin, Kristin Gunsalus, Fabio Piano, Enrique Martinez-Perez, Mark Blaxter, Peter Sarkies
The conserved phosphatase GSP-2/PP1 promotes germline immortality via small RNA-mediated genome silencing during meiosis
Katherine K Billmyre, Anna-lisa Doebley, Bree Heestand, Tony Belicard, Aya Sato-Carlton, Stephane Flibotte, Matt Simon, Megan Gnazzo, Ahna Skop, Donald Moerman, Peter Mark Carlton, Peter Sarkies, Shawn Ahmed
Transgenerational sterility of small RNA genome silencing mutants in response to germ granule dysfunction
Katherine Kretovich Billmyre, Bree Heestand, Maya Spichal, Shawn Ahmed
Epigenetic inheritance of telomere length in wild birds
Christina Bauch, Jelle J. Boonekamp, Peter Korsten, Ellis Mulder, Simon Verhulst
QTL mapping of natural variation reveals that the developmental regulator bruno reduces tolerance to P-element transposition in the Drosophila female germline
Erin Kelleher, Jaweria Jaweria, Uchechukwu Akoma, Lily Ortega, Wenpei Tang
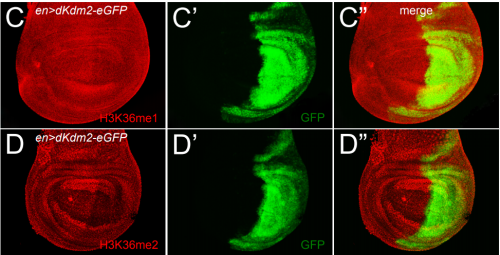
The lysine demethylase dKDM2 is non-essential for viability, but regulates circadian rhythms in Drosophila
Yani Zheng, Yongbo Xue, Xingjie Ren, Xiao-Jun Xie, Mengmeng Liu, Yu Jia, Xiao Li, Ye Niu, Jian-Quan Ni, Yong Zhang, Jun-Yuan Ji
Nonparametric Bayesian inference of transcriptional branching and recombination identifies regulators of early human germ cell development
Christopher Andrew Penfold, Anastasiya Sybirna, John Reid, Yun Huang, Lorenz Wernisch, Zoubin Ghahramani, Murray Grant, M. Azim Surani
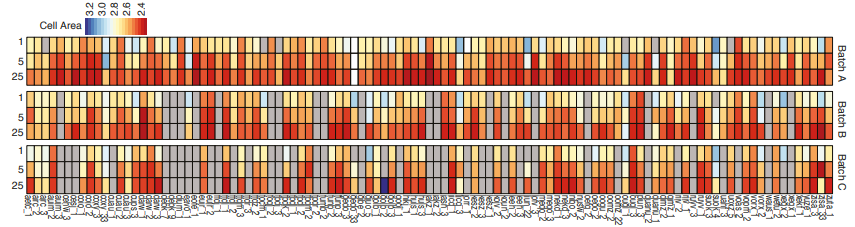
Identifying the genetic basis of variation in cell behaviour in human iPS cell lines from healthy donors
Alessandra Vigilante, Anna Laddach, Nathalie Moens, Ruta Meleckyte, Andreas Leha, Arsham Ghahramani, Oliver J. Culley, Annie Kathuria, Chloe Hurling, Alice Vickers, Mukul Tewary, Peter Zandstra, HipSci Consortium, Richard Durbin, Franca Fraternali, Oliver Stegle, Ewan Birney, Nicholas M Luscombe, Davide Danovi, Fiona M Watt
Profiling of pluripotency factors in individual stem cells and early embryos
Sarah J Hainer, Ana Boskovic, Oliver J Rando, Thomas G Fazzio
Partial reprogramming induces a steady decline in epigenetic age before loss of somatic identity
Nelly Olova, Daniel J Simpson, Riccardo Marioni, Tamir Chandra
Distinct and stage-specific contributions of TET1 and TET2 to stepwise cytosine oxidation in the transition from naive to primed pluripotency
Christopher B Mulholland, Franziska R Traube, Edris Parsa, Eva-Maria Eckl, Maximillian Schoenung, Miha Modic, Michael D Bartoschek, Paul Stolz, Joel Ryan, Thomas Carell, Heinrich Leonhardt, Sebastian Bultmann
Transcriptional repression by FACT is linked to regulation of chromatin accessibility at the promoter of ES cells
Peter Tessarz, Constantine Mylonas
Acidic microenvironment shaped by lactate accumulation promotes pluripotency through multiple mechanisms
Wen-Ting Guo, Shao-Hua Wang, Xiao-Shan Zhang, Ming Shi, Jing Hao, Xi-Wen Wang, Kai-Li Gu, Fei-Fei Duan, Ying Yan, Xi Yang, Chao Zhang, Le-Qi Liao, Yangming Wang
Chromosome compartmentalization replacement during stem cell differentiation
Yuri Eidelman, Svetlana Slanina, Valentina Pyatenko, Sergey Andreev
Coupling initial hydrogel microenvironments with mesenchymal stem cell remodeling to define a dynamic hematopoietic stem cell niche
Aidan E Gilchrist, Sunho Lee, Yuhang Hu, Brendan Harley
Longitudinal single cell fate in hematopoiesis in vivo using cellular barcoding and DiSNE movie visualization
Jerry Gao, Dawn Lin, Edmund Crampin, Shalin Naik
Urothelial organoids originate from Cd49f-High stem cells and display Notch-dependent differentiation capacity
Francisco X Real, Catarina P Santos, Eleonora Lapi, Laura Alvaro-Espinosa, Asuncion Fernandez-Barral, Antonio Barbachano, Diego Megias, Alberto Munoz
Onset of differentiation is posttranscriptionally controlled in adult neural stem cells
Avni Baser, Yonglong Dang, Maxim Skabkin, Guelce Guelcueler Balta, Georgios Kalamakis, Susanne Kleber, Manuel Goepferich, Roman Schefzik, Alejandro Santos Lopez, Enric Llorens Bobadilla, Carsten Schultz, Bernd Fischer, Ana Martin-Villalba
Single cell transcriptome profiling of mouse and hESC-derived pancreatic progenitors
Nicole AJ Krentz, Michelle Lee, Eric E Xu, Shugo Sasaki, Francis C Lynn
A Comparative Assessment of Human and Chimpanzee iPSC-derived Cardiomyocytes with Primary Heart Tissues
Bryan J Pavlovic, Lauren E Blake, Julien Roux, Claudia Chavarria, Yoav Gilad
Temporal epigenomic profiling identifies AHR and GLIS1 as super-enhancer controlled regulators of mesenchymal multipotency
Deborah Gerard, Florian Schmidt, Aurelien Ginolhac, Martine Schmitz, Rashi Halder, Peter Ebert, Marcel H. Schulz, Thomas Sauter, Lasse Sinkkonen
Correlated random walks of human embryonic stem cell in-vitro
L E Wadkin, S Orozco-Fuentes, I Neganova, G Swan, A Laude, M Lako, A Shukurov, N G Parker
An Epiblast Stem Cell derived multipotent progenitor population for axial extension
Shlomit Edri, Penelope Hayward, Peter Baillie-Johnson, Benjamin Steventon, Alfonso Martinez Arias
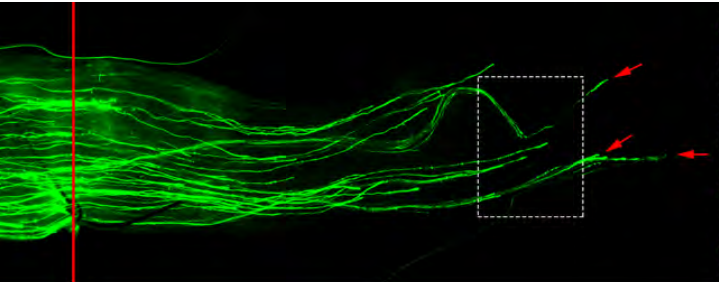
Novel role of Lin28 signaling in regulation of mammalian PNS and CNS axon regeneration
Xue-Wei Wang, Chang-Mei Liu, Philip A Hall, Jing-Jing Jiang, Christopher D Katchis, Sehwa Kang, Bryan C Dong, Shuxin Li, Feng-Quan Zhou
Positional information specifies the site of organ regeneration and not tissue maintenance in planarians
Eric M. Hill, Christian P Petersen
GABA promotes survival and axonal regeneration in identifiable descending neurons after spinal cord injury in larval lampreys
Daniel Romaus-Sanjurjo, Rocio Ledo-Garcia, Blanca Fernandez-Lopez, Kendra Hanslik, Jennifer R Morgan, Anton Barreiro-Iglesias, Maria Celina Rodicio
Experimentally Induced Metamorphosis in Axolotl (Ambystoma mexicanum) Under Constant Diet Restructures Microbiota Accompanied by Reduced Limb Regenerative Capacity
Turan Demircan, Guvanch Ovezmyradov, Berna YILDIRIM, Iknur Keskin, Ayse Elif ilhan, Ece Cana Fescioglu, Gurkan Ozturk, Suleyman Yildirim
Capn5 expression in the healthy and regenerating zebrafish retina
Cagney E. Coomer, Ann C Morris
A 3D model of human skeletal muscle innervated with stem cell-derived motor neurons enables epsilon-subunit targeted myasthenic syndrome studies
Mohsen Afshar Bakooshli, Ethan S Lippmann, Ben Mulcahy, Kayee Tung, Elena Pegoraro, Henry Ahn, Howard Ginsberg, Mei Zhen, Randolph S Ashton, Penney M Gilbert
Aneuploidy Promotes Intestinal Dysplasia in Drosophila
Luis P Resende, Augusta Monteiro, Rita Bras, Tatiana Lopes, Claudio E Sunkel
All-optical electrophysiology for high-throughput functional characterization of human iPSC-derived motor neuron model of ALS
Evangelos Kiskinis, Joel M Kralj, Peng Zou, Eli N Weinstein, Hongkang Zhang, Konstantinos Tsioras, Ole Wiskow, J. Alberto Ortega, Kevin Eggan, Adam E Cohen
Transcriptional and electrophysiological aberrations in an induced pluripotent stem cell-derived model of spinocerebellar ataxia type 7
Lauren M Watson, Danielle C Smith, Joseph V Raimondo, Richard J Burman, Robea Ballo, Janine Scholefield, Lyn Tyers, Sally A Cowley, Matthew JA Wood, Susan H Kidson, Leslie Jacquie Greenberg
Adipose fin development and its relation to the evolutionary origins of median fins
Thomas Stewart, Robert Ho, Melina A Hale
Gene regulatory network architecture in different developmental contexts influences the genetic basis of morphological evolution
Sebastian Kittelmann, Alexandra D. Buffry, Franziska A. Franke, Isabel Almudi, Marianne Yoth, Gonzalo Sabaris, Juan Pablo Couso, Maria D. S. Nunes, Nicolas Frankel, Jose Luis Gomez-Skarmeta, Jose Pueyo-Marques, Saad Arif, Alistair P. McGregor
The X chromosome of the German cockroach, Blattella germanica, is homologous to a fly X chromosome despite 400 million years divergence
Richard P Meisel, Judith R Wexler
The Hedgehog signaling domain was acquired from a prokaryote.
Henk Roelink
Evolution of the D. melanogaster chromatin landscape and its associated proteins
Elise Parey, Anton Crombach
Thirty clues to the exceptional diversification of flowering plants
Susana Magallon, Luna Sanchez-Reyes, Sandra Gomez Acevedo
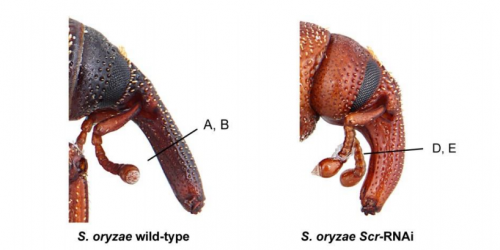

Developmental genetics in a complex adaptive structure, the weevil rostrum
Steven Davis
Germ layer specific regulation of cell polarity and adhesion gives insight into the evolution of mesoderm.
Miguel Salinas-Saavedra, Amber Q. Rock, Mark Q. Martindale
Placozoa and Cnidaria are sister taxa
Christopher E. Laumer, Harald Gruber-Vodicka, Michael G. Hadfield, Vicki B. Pearse, Ana Riesgo, John C. Marioni, Gonzalo Giribet
Functional Characterization of Enhancer Evolution in the Primate Lineage
Jason Chesler Klein, Aidan Keith, Vikram Agarwal, Timothy Durham, Jay Shendure
Tempo, mode, and fitness effects of mutation in Caenorhabditis elegans over 400 generations of minimal selection
Ayush Shekhar Saxena, Matthew P Salomon, Chikako Matsuba, Shu-Dan Yeh, Charles F Baer
Transposable elements and gene expression during the evolution of amniotes
Lu Zeng, Stephen Pederson, Dan Kortschak, David Adelson
Diverse RNA interference strategies in early-branching metazoans
Andrew D Calcino, Selene L Fernandez-Valverde, Ryan J Taft, Bernard M Degnan
What is an archaeon and are the Archaea really unique?
Ajith Harish
CHARACTERIZATION OF THE bHLH FAMILY OF TRANSCRIPTIONAL REGULATORS IN THE ACOEL S. roscoffensis AND THEIR PUTATIVE ROLE IN NEUROGENESIS
Elena Perea-Atienza, Simon G. Sprecher, Pedro Martinez
Multiple large-scale gene and genome duplications during the evolution of hexapods
Zheng Li, George Tiley, Sally Galuska, Chris Reardon, Thomas Kidder, Rebecca Rundell, Michael S. Barker
Finding Nemo’s Genes: A chromosome-scale reference assembly of the genome of the orange clownfish Amphiprion percula
Robert Lehmann, Damien J Lightfoot, Celia Schunter, Craig T Michell, Hajime Ohyanagi, Katsuhiko Mineta, Sylvain Foret, Michael Berumen, David J Miller, Manuel Aranda, Takashi Gojobori, Philip L Munday, Timothy Ravasi
Improved genome assembly and annotation for the rock pigeon (Columba livia)
Carson Holt, Michael Campbell, David Keays, Nathaniel Edelman, Aurelie Kapusta, Emily Maclary, Eric Domyan, Alexander Suh, Wesley Warren, Mark Yandell, Thomas Gilbert, Michael D. Shapiro
A phylogenomic framework and timescale for comparative studies of tunicates
Frederic Delsuc, Herve Philippe, Georgia Tsagkogeorga, Paul Simion, Marie-Ka Tilak, Xavier Turon, Susanna Lopez-Legentil, Jacques Piette, Patrick Lemaire, Emmanuel J. P. Douzery
Natural selection on gene-specific codon usage bias is common across eukaryotes
Zhen Peng, Hani Zaher, Yehuda Ben-Shahar
Spatial Control of Draper Receptor Signaling Initiates Apoptotic Cell Engulfment
Adam P. Williamson, Ronald D. Vale
Assembly of a Parts List of the Human Mitotic Cell Cycle Machinery
Bruno Giotti, Sz-Hau Chen, Mark W Barnett, Tim Regan, Tony Ly, Stefan Wiemann, David A Hume, Tom C Freeman
CDC20B is required for deuterosome-mediated centriole production in multiciliated cells
Diego R. Revinski, Laure-Emmanuelle Zaragosi, Camille Boutin, Sandra Ruiz-Garcia, Marie Deprez, Virginie Thome, Olivier Rosnet, Anne-Sophie Gay, Olivier Mercey, Agnes Paquet, Nicolas Pons, Gilles Ponzio, Brice Marcet, Laurent Kodjabachian, Pascal Barbry
Scaffolding of RhoA contractile signaling by anillin: a regulatory analogue of kinetic proofreading
Srikanth Budnar, Kabir B. Husain, Guillermo A. Gomez, Maedeh Naghibosidat, Suzie Verma, Nicholas A. Hamilton, Richard G. Morris, Alpha S. Yap
A mechanosensitive RhoA pathway that protects epithelia against acute tensile stress.
Bipul R. Acharya, Alexander Nestor-Bergmann, Xuan Liang, Srikanth Budnar, Oliver E. Jensen, Zev Bryant, Alpha S. Yap
Dynamics of sister chromatid resolution during cell cycle progression
Rugile Stanyte, Johannes Nuebler, Claudia Blaukopf, Rudolf Hoefler, Roman Stocsits, Jan-Michael Peters, Daniel W Gerlich
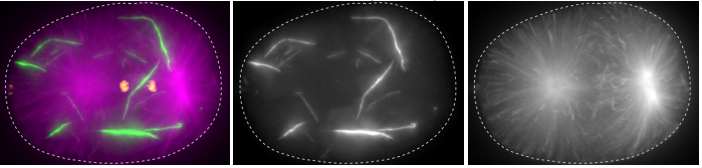
Self-assembly of the RZZ complex into filaments drives kinetochore expansion in the absence of microtubule attachment
Reto Gassmann, Cláudia Pereira, Rita M Reis, José B Gama, Dhanya K Cheerambathur, Ana X Carvalho
Dynamic Kinetochore Size Regulation Promotes Microtubule Capture And Chromosome Biorientation In Mitosis
Carlos Sacristan, Misbha Ahmad, Jenny Keller, Job Fermie, Vincent Groenewold, Eelco Tromer, Alexander Fish, Roberto Melero, Jose Maria Carazo, Judith Klumperman, Andrea Musacchio, Anastassis Perrakis, Geert JPL Kops
Optogenetic reconstitution reveals that Dynein-Dynactin-NuMA clusters generate cortical spindle-pulling forces as a multi-arm ensemble
Masako Okumura, Toyoaki Natsume, Masato T Kanemaki, Tomomi Kiyomitsu
A novel microtubule nucleation pathway for meiotic spindle assembly in oocytes
Pierre ROME, Hiroyuki OHKURA
A compartmentalized signaling network mediates crossover control in meiosis
Liangyu Zhang, Simone Koehler, Regina Rillo-Bohn, Abby F. Dernburg
Microtubule end tethering of a processive Kinesin-8 motor Kif18b is required for spindle positioning
Toni McHugh, Agata Gluszek-Kustusz, Julie Welburn
Mitotic progression, arrest, exit or death is determined by centromere integrity and independent of de novo transcription
Marco Cruz, Maria Alba Fernandez, Wilfred van Ijcken, Niels Galjart, Arockia Jeyaprakash Arulanandam, Helder Maiato, Cristina Ferras
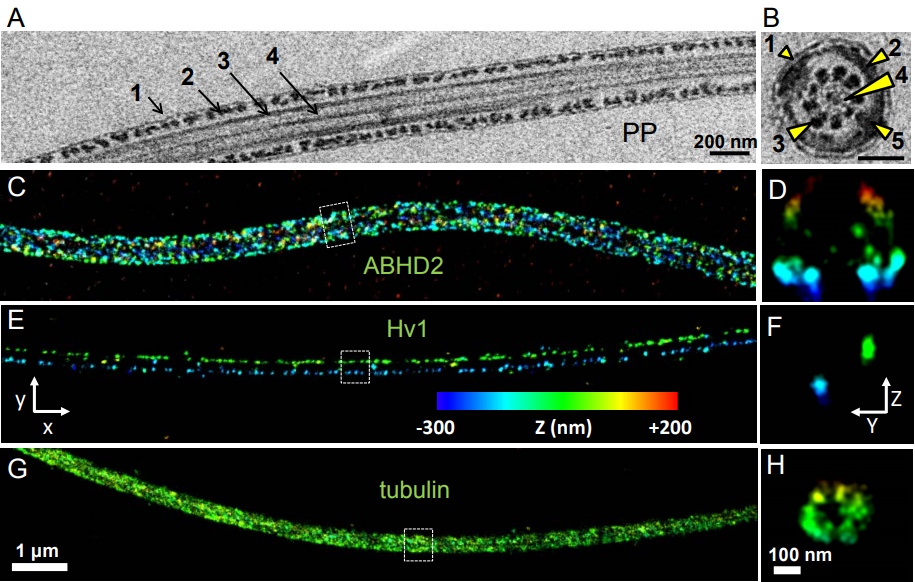
Asymmetrically Positioned Flagellar Control Units Regulate Human Sperm Rotation
Melissa Miller, Samuel Kenny, Nadja Mannowetz, Steven Mansell, Michal Wojcik, Sarah Mendoza, Robert Zucker, Ke Xu, Polina V Lishko
Regulation of DVL2 ubiquitylation by USP9X biases participation in canonical or non-canonical WNT signalling
Casey P Nielsen, Kristin K Jernigan, Jason A. MacGurn
Spatiotemporal manipulation of ciliary glutamylation reveals its roles in intraciliary trafficking and Hedgehog signaling
Shi-Rong Hong, Cuei-Ling Wang, Yao-Shen Huang, Yu-Chen Chang, Ya-Chu Chang, Ganesh V. Pusapati, Chun-Yu Lin, Ning Hsu, Hsiao-Chi Cheng, Yueh-Chen Chiang, Wei-En Huang, Nathan C. Shaner, Rajat Rohatgi, Takanari Inoue, Yu-Chun Lin
Cytoplasmic sequestration of the RhoA effector mDiaphanous1 by Prohibitin2 promotes muscle differentiation
Jyotsna Dhawan, Amena Saleh, Gunasekaran Subramaniam, Swasti Raychaudhuri
Polyamines support myogenesis by facilitating myoblast migration
Shirley Brenner, Yulia Feiler, Chaim Kahana
Stabilising and destabilising kinesin complexes queue at plus tips to ensure microtubule catastrophe at cell ends
John C Meadows, Liam J Messin, Anton Kamnev, Theresa C Lancaster, Mohan K Balasubramanian, Robert A Cross, Jonathan B.A. Millar
CLASP promotes microtubule array reorientation by acting as a specific severing rescue factor
Jelmer J Lindeboom, Masayoshi Nakamura, Marco Saltini, Anneke Hibbel, Ankit Walia, Tijs Keetlaar, Anne Mie C Emons, John C Sedbrook, Viktor Kirik, Bela M Mulder, David W Ehrhardt
Three-dimensional cell geometry controls excitable membrane signaling in Dictyotelium cells
Marcel Hörning, Tatsuo Shibata
An experimental and computational framework to build a dynamic protein atlas of human cell division
Yin Cai, M. Julius Hossain, Jean-Karim Heriche, Antonio Z. Politi, Nike Walther, Birgit Koch, Malte Wachsmuth, Bianca Nijmeijer, Moritz Kueblbeck, Marina Martinic Kavur, Rene Ladurner, Jan-Michael Peters, Jan Ellenberg
Bradly J Alicea, Richard Gordon
DevoWorm: data-theoretical synthesis of C. elegans development.
Bradly John Alicea, Richard Gordon, Thomas E. Portegys

Beyond activator-inhibitor networks: the generalised Turing mechanism
Stephen Smith, Neil Dalchau
Selection for synchronized cell division in simple multicellular organisms
Jason Olejarz, Kamran Kaveh, Carl Veller, Martin A. Nowak
Branch-recombinant Gaussian processes for analysis of perturbations in biological time series
Christopher A Penfold, Anastasiya Sybirna, John Reid, Yun Huang, Lorenz Wernisch, Zoubin Ghahramani, Murray Grant, M Azim Surani
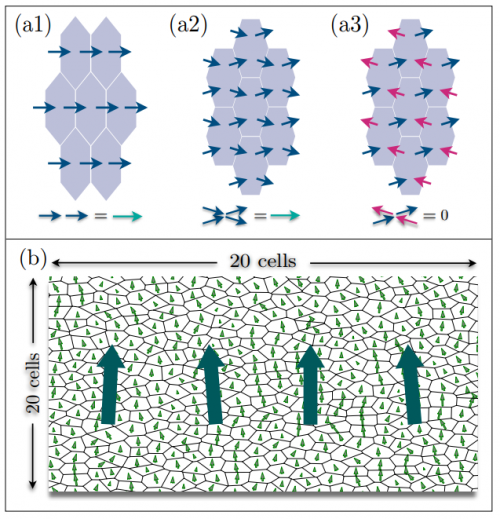
The Role of Cytoplasmic Interactions in the Collective Polarization of Tissues and its Interplay with Cellular Geometry
Shahriar Shadkhoo, Madhav Mani
Interplay between epidermal stem cell dynamics and dermal deformations
Yasuaki Kobayashi, Yusuke Yasugahira, Hiroyuki Kitahata, Mika Watanabe, Ken Natsuga, Masaharu Nagayama
Confluent and non-confluent phases in a model of cell tissue
Eial Teomy, David A. Kessler, Herbert Levine
The Role of Cytoplasmic Interactions in the Collective Polarization of Tissues and its Interplay with Cellular Geometry
Shahriar Shadkhoo, Madhav Mani
Optimal feedback mechanisms for regulating cell numbers
Saurabh Modi, Abhyudai Singh
Transition from actin-driven to water-driven cell migration depends on external hydraulic resistance
Yizeng Li, Sean X. Sun
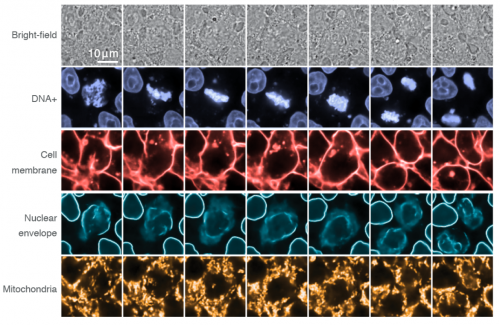

Label-free prediction of three-dimensional fluorescence images from transmitted light microscopy
Chawin Ounkomol, Sharmishtaa Seshamani, Mary M Maleckar, Forrest Collman, Gregory Johnson
Third-generation in situ hybridization chain reaction: multiplexed, quantitative, sensitive, versatile, robust
Harry M.T. Choi, Maayan Schwarzkopf, Mark E. Fornace, Aneesh Acharya, Georgios Artavanis, Johannes Stegmaier, Alexandre Cunha, Niles A. Pierce
Automated sequential chromogenic IHC double staining with two HRP substrates.
Kenneth H Petersen, Jesper Lohse, Lasse Ramsgaard
Forget Pixels: Adaptive Particle Representation of Fluorescence Microscopy Images
Bevan L. Cheeseman, Ulrik Günther, Mateusz Susik, Krzysztof Gonciarz, Ivo F. Sbalzarini
Label-free assessment of pre-implantation embryo quality by the Fluorescence Lifetime Imaging Microscopy (FLIM)-phasor approach
Ning Ma, Nabora Reyes de Mochel, Paula Pham, Tae Yoo, Ken Cho, Michelle Digman
Visualizing multiple inter-organelle contact sites using the organelle-targeted split-GFP system
Yuriko Kakimoto, Shinya Tashiro, Rieko Kojima, Yuuki Morozumi, Toshiya Endo, Yasushi Tamura
Signal-to-Noise ratio made easy: A tool to assess your confocal performance
Alexia Ferrand, Kai D Schleicher, Nikolaus Ehrenfeuchter, Wolf Heusermann, Oliver Biehlmaier
Smart Region-Growing: a novel algorithm for the segmentation of 3D clarified confocal image stacks
Alejandro Callara, Chiara Magliaro, Arti Ahluwalia, Nicola Vanello
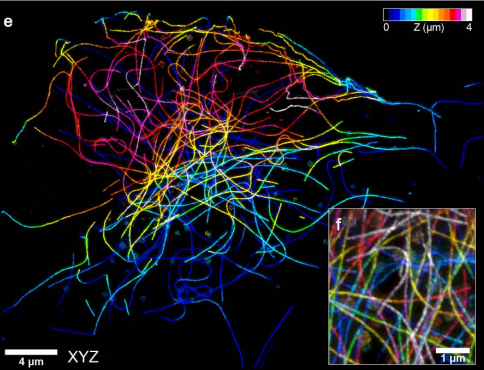
Expansion stimulated emission depletion microscopy (ExSTED)
Mengfei Gao, Riccardo Maraspini, Oliver Beutel, Amin Zehtabian, Britta Eickholt, Alf Honigmann, Helge Ewers
Complementary studies of lipid membrane dynamics using iSCAT and STED Fluorescence Correlation Spectroscopy
Francesco Reina, Silvia Galiani, Dilip Shrestha, Erdinc Sezgin, Gabrielle de Wit, Daniel Cole, B. Christoffer Lagerholm, Philipp Kukura, Christian Eggeling
Label-free high-speed wide-field imaging of single microtubules using interference reflection microscopy
Mohammed Mahamdeh, Steve Simmert, Anna Luchniak, Erik Schaeffer, Jonathon Howard
A new approach to design artificial 3D micro-niches with combined chemical, topographical and rheological cues
Celine Stoecklin, Zhang Yue, Wilhelm W. Chen, Richard de Mets, Eileen Fong, Vincent Studer, Virgile Viasnoff
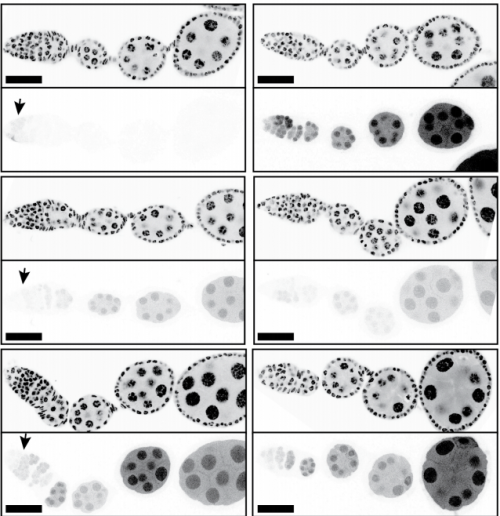
Efficient expression of genes in the Drosophila germline using a UAS-promoter free of interference by Hsp70 piRNAs
Steven Z DeLuca, Allan C Spradling
Germline silencing of UASt depends on the piRNA pathway
Yi-Chun Huang, Henry Moreno, Sarayu Row, Dongyu Jia, Wu-Min Deng
Toward Precision Molecular Surgery: Robust, Selective Induction of Microhomology-mediated End Joining in vivo
Hirotaka Ata, Thomas L Ekstrom, Gabriel Martinez-Galvez, Carla M Mann, Alexey V Dvornikov, Kyle J Schaefbauer, Alvin C Ma, Drena Dobbs, Karl J Clark, Stephen C Ekker
Corrigendum and follow-up: Whole genome sequencing of multiple CRISPR-edited mouse lines suggests no excess mutations.
Kellie A. Schaefer, Benjamin W. Darbro, Diana F. Colgan, Stephen H. Tsang, Alexander G. Bassuk, Vinit B. Mahajan
A large-scale whole-genome sequencing analysis reveals highly specific genome editing by both Cas9 and Cpf1 nucleases in rice
Xu Tang, Guanqing Liu, Jianping Zhou, Qiurong Ren, Qi You, Li Tian, Xuhui Xin, Zhaohui Zhong, Binglin Liu, Xuelian Zheng, Dengwei Zhang, Aimee Malzahn, Zhiyun Gong, Yiping Qi, Tao Zhang, Yong Zhang
CRISPR-Tag: an Efficient DNA Tagging System in Living Cells
Baohui Chen, Wei Zou, Bo Huang
A homing CRISPR mouse resource for barcoding and lineage tracing
Reza Kalhor, Kian Kalhor, Kathleen Leeper, Amanda Graveline, Prashant Mali, George M Church
High-throughput antibody engineering in mammalian cells by CRISPR/Cas9-mediated homology-directed mutagenesis
Derek M Mason, Cedric R Weber, Cristina Parola, Simon M Meng, Victor Greiff, William J Kelton, Sai T Reddy
An optimized electroporation approach for efficient CRISPR/Cas9 genome editing in murine zygotes
Simon E. Tröder, Lena K. Ebert, Linus Butt, Sonja Assenmacher, Bernhard Schermer, Branko Zevnik
Ultra-parallel ChIP-seq by barcoding of intact nuclei
Laura Arrigoni, Hoor Al-Hasani, Fidel Ramirez, Ilaria Panzeri, Devon P Ryan, Diana Santacruz, Nadia Kress, Andrew J Pospisilik, Ulrike Boenisch, Thomas Manke
Generation and validation of homozygous fluorescent knock-in cells using genome editing
Birgit Koch, Bianca Nijmeijer, Moritz Kueblbeck, Yin Cai, Nike Walther, Jan Ellenberg
A comprehensive toolkit to enable MinION long-read sequencing in any laboratory
Miriam Schalamun, David Kainer, Eleanor Beavan, Ramawatar Nagar, David Eccles, John Rathjen, Robert Lanfear, Benjamin Schwessinger
Topographer Reveals Dynamic Mechanisms of Cell Fate Decisions from Single-Cell Transcriptomic Data
Jiajun Zhang, Tianshou Zhou, Qing Nie
A biological-computational human cell lineage discovery platform based on duplex molecular inversion probes
Liming Tao, Ofir Raz, Zipora Marx, Tamir Biezuner, Shiran Amir, Lilach Milo, Rivka Adar, Amos Onn, Noa Chapal-Ilani, Veronika Berman, Ron Levy, Barak Oron, Ehud Shapiro
Towards Improving Embryo Selection: Simultaneous Next Generation Sequencing Of DNA And RNA From A Single Trophectoderm Biopsy
Noga Fuchs Weizman, Brandon Wyse, Ran Antes, Zenon Ibarrientos, Mugundhine Sangaralingam, Gelareh Motamedi, Valeriy Kuznyetsov, Svetlana Madjunkova, Clifford L Librach
NanoMod: a computational tool to detect DNA modifications using Nanopore long-read sequencing data
Qian Liu, Daniela C. Georgieva, Dieter Egli, Kai Wang
Auxin-Mediated Sterility Induction System for Longevity and Mating Studies in Caenorhabditis elegans
Katja R Kasimatis, Megan J. Moerdyk-Schauecker, Patrick C. Phillips
DECODE-ing sparsity patterns in single-cell RNA-seq
Shahin Mohammadi, Jose Davila-Velderrain, Manolis Kellis, Ananth Grama
Take ACTION to characterize the functional identity of single cells
Shahin Mohammadi, Vikram Ravindra, David Gleich, Ananth Grama
Pisces: An Accurate and Versatile Variant Caller for Somatic and Germline Next-Generation Sequencing Data
Tamsen Dunn, Gwenn Berry, Dorothea Emig-Agius, Yu Jiang, Serena Lei, Anita Iyer, Nitin Udar, Han-Yu Chuang, Jeff Hegarty, Michael Dickover, Brandy Klotzle, Justin Robbins, Marina Bibikova, Marc Peeters, Michael Stromberg
Twenty steps towards an adequate inferential interpretation of p-values
Norbert Hirschauer, Sven Grüner, Oliver Mußhoff, Claudia Becker
Adjutant: an R-based tool to support topic discovery for systematic and literature reviews
Anamaria Crisan, Tamara Munzner, Jennifer L. Gardy
Quantifying the impact of public omics data
Yasset Perez-Riverol, Andrey Zorin, Gaurhari Dass, Mihai Glonț, Juan Antonio Vizcaino, Andrew Jarnuczak, Robert Petryszak, Peipei Ping, Henning Hermjakob
Changes in Acceptance of Evolution and Associated Factors during a Year of Introductory Biology: The Shifting Impacts of Biology Knowledge, Politics, Religion, Demographics, and Understandings of the Nature of Science
Ryan D.P. Dunk, Jason R. Wiles
Persistent Underrepresentation of Women’s Science in High Profile Journals
Yiqin Alicia Shen, Jason M Webster, Yuichi Shoda, Ione Fine
Female grant applicants are equally successful when peer reviewers assess the science, but not when they assess the scientist
Holly O Witteman, Michael Hendricks, Sharon Straus, Cara Tannenbaum
A Data Citation Roadmap for Scientific Publishers
Helena Cousijn, Amye Kenall, Emma Ganley, Melissa Harrison, David Kernohan, Thomas Lemberger, Fiona Murphy, Patrick Polischuk, Simone Taylor, Maryann Martone, Timothy Clark
We need to talk about reliability: Making better use of test retest studies for study design and interpretation
Granville James Matheson
Dhanush Kumar, Rakhi Radhamani, Nijin Nizar, Krishnashree Achuthan, Bipin Nair, Shyam Diwakar
Researcher engagement in policy deemed societally beneficial yet unrewarded
Gerald G Singh, Vinicius Farjalla, Bing Chen, Andrew Pelling, Elvan Ceyhan, Martin Dominik, Eva Alisic, Jeremy Kerr, Noelle Selin, Ghada Bassioni, Elena Bennett, Andrew Kemp, Kai MA Chan
Chris HJ Hartgerink, Marino van Zelst
An Observational Study of Ballooning in Large Spiders: Nanoscale Multi-Fibres Enable Large Spiders’ Soaring Flight
Moonsung Cho, Peter Neubauer, Christoph Fahrenson, Ingo Rechenberg
Quantifying the unquantifiable: why Hymenoptera — not Coleoptera — is the most speciose animal order
Andrew A Forbes, Robin K Bagley, Marc A Beer, Alaine C Hippee, Heather A Widmayer
Ancient DNA reveals the chronology of walrus ivory trade from Norse Greenland
Bastiaan Star, James H Barrett, Agata T Gondek, Sanne Boessenkool