I make art that brings together music, animation, and performance to explore evolutionary and developmental biology themes. I aim to illuminate the magic of the biological world for a broad audience of scientists and non-scientists, young and old. In order to communicate science to the general public and fuel a sense of awe at the natural world, I intertwine biological stories with classical narratives and human emotion, and I create multimedia experiences that allow an audience of diverse backgrounds to have different entry points—musical, visual, theatrical, or scientific—to the experience. Here’s a trailer from my performance Theory of Flight, where a lecturing scientist reveals she’s been growing her own wings using avian genes, and animations and music bring the transgenic processes to life.
I was introduced to many of the evolutionary and developmental biology (evo-devo) themes featured in my work while studying biology at Yale, and while working in Dr. Antónia Monteiro’s lab. When the Node asked me to write about how I go about scientific outreach, I thought a dialogue with Dr. Monteiro would be the perfect way to delve into the process of bringing biological ideas to a broad audience.
Many thanks to Dr. Antónia Monteiro, Associate Professor, Yale-NUS-College and Department of Biological Sciences, National University of Singapore.
Dr. Antónia Monteiro (AM): Anna, you have a bachelor’s degree in biology from Yale University and have published your senior thesis project in the field of evolution of development. But you went on to pursue a masters degree in electronic arts. Did you ever think about a career in science? And if yes, what was it about it that did not quite satisfy you?
Anna Lindemann (AL): There were certainly times when I imagined pursuing a more traditional scientific career, but my life seems to be an experiment with an ever-shifting protocol. I exist in a hazy territory between art and science. This means that sometimes I feel lost—I am a “science” person around “art” people, and an “art” person around “science” people. But at the best times, this hazy territory is an exciting place to be, full of possibilities and new connections, whether that’s connecting disparate disciplines to generate new forms of creative output, or whether that’s connecting other people to new ideas that lie outside of their area of specialization.
AM: I have seen several of your performances. You are very creative with your use of music, performance, audio-visuals, etc. What is going on in your head when you are planning a new piece? Do you think first about the science you want to communicate? The music you want to use? How do you go about pulling it all together?
AL: My approach to integrating artistic and scientific disciplines is twofold. I am interested in illuminating biological processes, especially evo-devo processes, for a broad audience, and I am also interested in developing art using biological processes as a model for creation.
So, sometimes I am thinking about the most effective visual or musical elements to convey a particular biological story in an engaging, dramatic, or humorous way. And sometimes I am trying to create interesting, new, complex, absurd, and beautiful sounds and images, and I turn to biological processes in order to do that, processes that have evolved over millions of years to create interesting, complex, absurd, and beautiful life forms. To this end, many of my projects include music that has been developed algorithmically based on developmental biology. I am currently exploring a new way of developing music generated from the dynamic gene expression output of gene network simulations.
Ultimately, how the musical, theatrical, visual, and biological parts come together involves a lot of trial and error, and is certainly far from an efficient or direct assembly process.
AM: The evo-devo science behind your pieces is state-of-the-art and complex, yet, you manage clearly to convey the big picture to a broad college-educated audience. How do you go about picking the evo-devo themes for your performances? And shaping them for a broad audience?
AL: I first learned about many of the evo-devo themes featured in my work, including the modularity and co-option of genetic networks, through your evo-devo course, other courses I took at Yale, and the experience of working in your lab. The thesis research I did in your lab investigating the role of the Hedgehog signaling pathway in butterfly eyespot development was a significant influence on the investigations that the main character pursues in my performance Theory of Flight. Once I start developing a project, I also read a lot of scientific journal articles. People have commented that Theory of Flight is the first performance they’ve seen with a program that has citations.
There are a few things that stand out in terms of shaping the evo-devo themes for a broad audience. First, I want audiences to leave with a sense of wonder and appreciation for the evo-devo processes that give rise to the complex biological world. I want an audience to have that same contagious sense of awe that I remember feeling when first learning about evo-devo from you, the sense that all of the growing and evolving that organisms do is mysterious and incredible. Keeping a sense of awe in the foreground, and mechanistic details in the background involves lots of revisions of a project for me. Even though I create fictional narratives, I want the science behind those narratives to be detailed and realistic. I constantly have to reign in the tendency to bombard and overwhelm an audience with details that would ultimately dissuade rather than encourage them from engaging the evo-devo themes.
Second, I think about connecting the evo-devo themes to universal human experiences. In Theory of Flight, the exploration of the developmental mechanisms of wing growth and the evolutionary origins of flight are framed within the context of a character’s quest to develop her own flight. An audience can relate on some level to the character’s ambitious striving, failures, and isolation, even if perceiving the world through a biological lens is something new to them.
Third, I embrace a multimedia mindset. I believe that bringing together performance, music, and animation not only allows for evo-devo themes to be explored in a variety of ways, but also connects to a diverse audience by providing multiple entry points to the performance. For some audience members, the biology may be familiar, but the theatrical and musical context might be unfamiliar. For other audience members, the animation or music or performance might be familiar territory and the biology foreign territory.
AM: Writing, performing, and then trying to “market” your product must be challenging. Are there organizations or venues that can help people like you, at the intersection of performance art and science, get more visibility? Have you thought about ideal venues for how artists/scientists like yourself should be interacting with the rest of us?
AL: I am excited by the growing number of venues, series, events, and institutions that are interested in supporting work at the intersection of art and science. I’ve had a wonderful time working with some of them.
Theory of Flight was premiered at one of the black box theaters equipped with state-of-the-art multimedia performance technology at Experimental Media and Performing Arts Center (EMPAC) in Troy, NY. EMPAC opened to the public in 2008 as a place “where the arts, sciences, and technology interact with and influence each other by using the same facilities, technologies, and by breathing the same air.”
I’ve performed at the Entertaining Science series in New York City which pairs lectures by prominent scientists with performances by diverse artists. Entertaining Science started in 2002 as a monthly series organized by chemist and poet Roald Hoffman of Cornell University and neuroscientist and composer Dave Soldier of Columbia University.
I’ve presented at a marathon “Survival of the Beautiful Wonder Cabinet” organized by David Rothenberg that brought together artists and scientists that think about aesthetics and evolution in their work. And I’ve presented through the Franke Program in Science and the Humanities that started this past year at Yale.
I also work with high school students to think creatively across the arts and sciences through a program called the ArtScience Prize, which is part of a larger international organization of educational programs and exhibition spaces that focus on the creativity that can emerge from interdisciplinary investigation.
There are many other places around the world, far too many to name here, that celebrate art-science intersections in various ways, and many that I hope will continue to form. I am always interested in new presentation contexts that may or may not have an explicit mission to integrate art and science, and that might have resources and budgets big and small. For example, I would love to some day perform at a biology conference.
I think there are still many untapped possibilities for how art and science can come together to spark new forms of understanding and creativity, and to be meaningful for artists, scientists, and the generally curious. I’m interested in how venues like the Node can foster this kind of interaction online, and I’m interested to hear from readers of the Node about their experiences in art-science intersections (either as participants or observers), and their visions for future cross-disciplinary activity.
 This post is part of a series on science outreach. You can read the introduction to the series here and read other posts in this series here.
This post is part of a series on science outreach. You can read the introduction to the series here and read other posts in this series here.
 (6 votes)
(6 votes)
 Loading...
Loading...
 We’re excited to announce that our short film, Stem cells – the future: an introduction to iPS cells, is now available in French, German, Italian, Polish and Spanish as well as English, with a supporting quiz for the classroom. You can order a DVD or view the film online and download the quiz, all in your language. Read more
We’re excited to announce that our short film, Stem cells – the future: an introduction to iPS cells, is now available in French, German, Italian, Polish and Spanish as well as English, with a supporting quiz for the classroom. You can order a DVD or view the film online and download the quiz, all in your language. Read more Last Wednesday, a large, friendly audience of scientists, writers and science communicators gathered in Edinburgh to celebrate EuroStemCell’s recent writing competition by launching a booklet of winning entries, and listening to stunning readings from the booklet. A fantastic panel of writers discussed the interplay between words and science, and one audience member was so impressed with the panel’s thoughts that she wanted to ‘eat their brains’! There was lots of chatter on Twitter too, all captured in a Storify of the tweets. Read more
Last Wednesday, a large, friendly audience of scientists, writers and science communicators gathered in Edinburgh to celebrate EuroStemCell’s recent writing competition by launching a booklet of winning entries, and listening to stunning readings from the booklet. A fantastic panel of writers discussed the interplay between words and science, and one audience member was so impressed with the panel’s thoughts that she wanted to ‘eat their brains’! There was lots of chatter on Twitter too, all captured in a Storify of the tweets. Read more Many congratulations to our partner, Dr Elena Cattaneo, who is the winner of the 2013 Stem Cell Person of the Year Award run by Dr Paul Knoepfler via his well-known blog. We know Elena not only as a leading scientist, but also as a very active, fantastically energetic and supportive collaborator in public engagement with stem cell research. She’s also recently become a senator. All in all, a great choice for Stem Cell Person of the Year!
Many congratulations to our partner, Dr Elena Cattaneo, who is the winner of the 2013 Stem Cell Person of the Year Award run by Dr Paul Knoepfler via his well-known blog. We know Elena not only as a leading scientist, but also as a very active, fantastically energetic and supportive collaborator in public engagement with stem cell research. She’s also recently become a senator. All in all, a great choice for Stem Cell Person of the Year!

 (1 votes)
(1 votes)
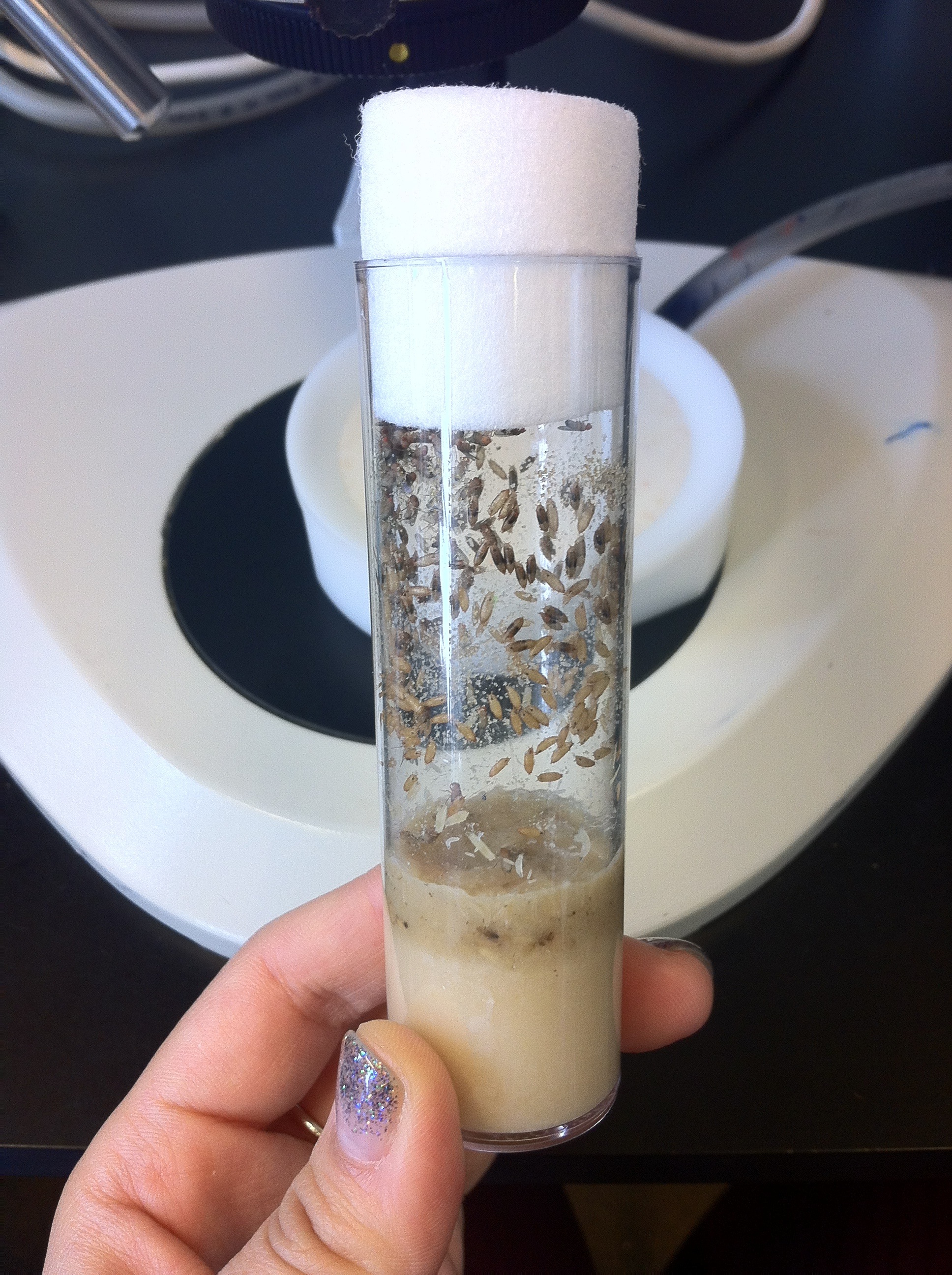
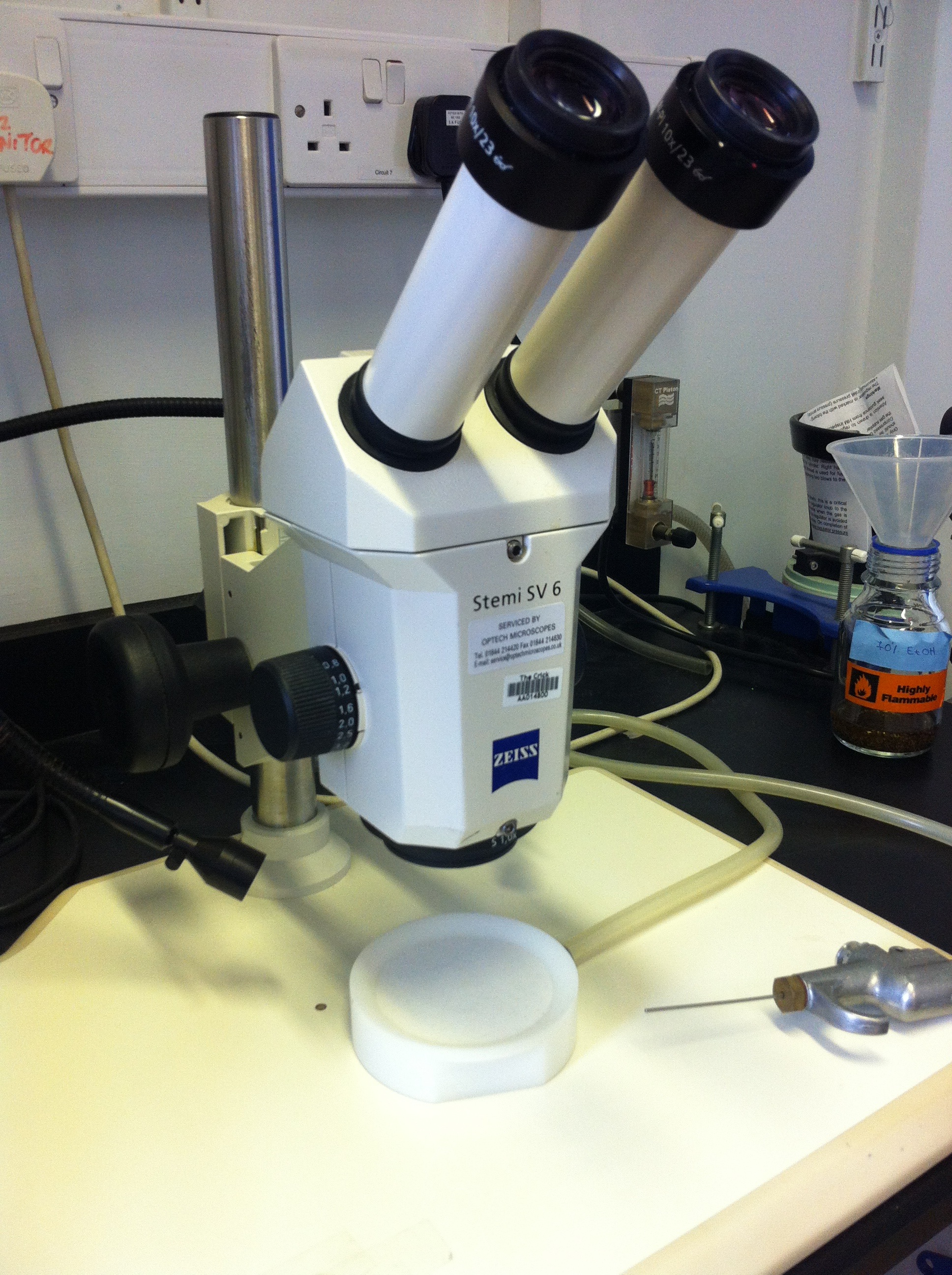
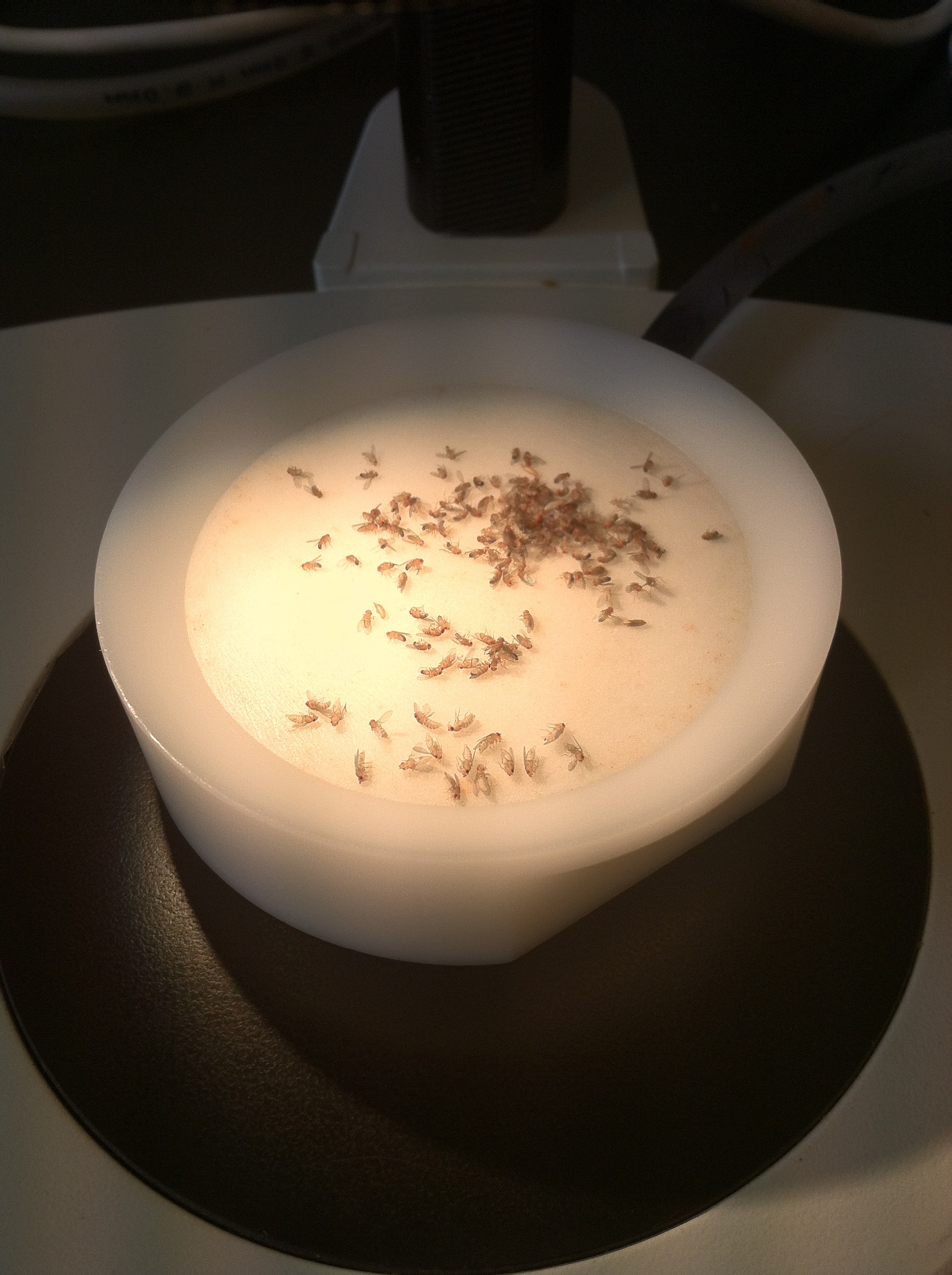



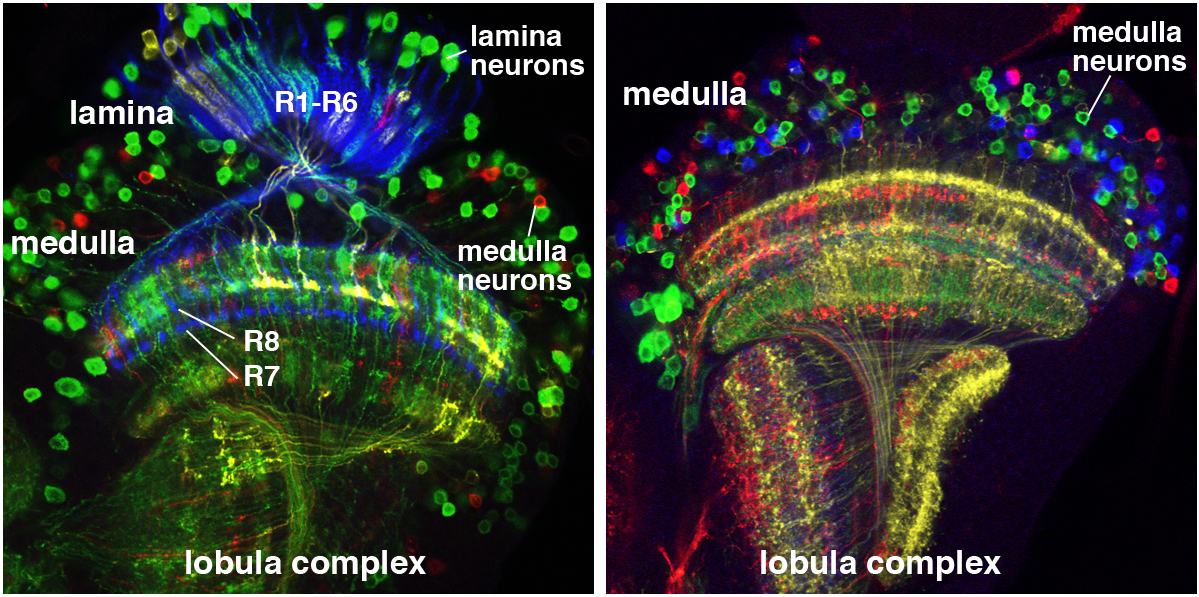
 This post is part of a series on a day in the life of developmental biology labs working on different model organisms. You can read the introduction to the series
This post is part of a series on a day in the life of developmental biology labs working on different model organisms. You can read the introduction to the series  (17 votes)
(17 votes)





 (No Ratings Yet)
(No Ratings Yet)