Flies with colon cancer help to unravel the genetic keys to disease in humans
Posted by IRBBarcelona, on 8 October 2014
Researchers generate for the first time Drosophila melanogaster with intestinal cancer and reveal key genetic factors behind human colon cancer.
The scientists identify a human gene that favours the proliferation of tumour cells in early stages of colon cancer.
Furthermore, the flies are useful for faster and more economic drug screening.
Researchers at the Institute for Research in Biomedicine (IRB Barcelona) have managed to generate a fruit fly (Drosophila melanogaster) model that reproduces human colon cancer. With two publications appearing in PLoS One and EMBO Reports, the IRB team also unveil the function of a key gene in the development of the disease.
“The breakthrough is that we have generated cancer in an adult organism and from stem cells, thus reproducing what happens in most types of human cancer. This model has allowed us to identify subtle interactions in the development of cancer that are practically impossible to detect in mice with the current technology available,” explains the biologist Andreu Casali, Associate Researcher at IRB Barcelona and leader of the Drosophila project.
Although the flies do not have a colon, they have an intestine that includes a colon and rectum and that works in the same way as the human colon. The scientists generated mutant flies for two genes that are altered in most human colon tumours, namely APC and Ras. Thanks to the ease with which genetic studies can be performed in Drosophila, the researchers were able to examine the effect of 250 genes that are altered in these types of tumour and found that, of these, 30% affected growth while the others had no significant effects.
“The advantage of the model is that it allows us to explore genetic alterations more quickly, to distinguish between those that are important and those that are not, and to see what role they play,” explains Òscar Martorell, first author of the paper that appears in EMBO Reports published today. “Performing these genetic experiments in mice is time-consuming and costly and the Drosophila model allows us to rapidly analyse new pathways that could be relevant for colon cancer,” adds the co-author of the study, Francisco Barriga, a postdoctoral fellow working on colon cancer in vertebrate models. Undertaken over five years, the study is the result of collaboration between the Development and Morphogenesis in Drosophila Lab and the Colorectal Cancer Lab, both at IRB Barcelona.
Of all the genes that have a relevant function, the group focused on one called Mirror in Drosophila and lrx in humans. The experiments with flies led to the finding that this gene favours tumour growth in early stages of human cancer. “The problem with human cancer is that we know very little about what happens in the early stages. Our models allows us to better study its development.” Also, Casali goes on to speculate that the human gene lrx may become a good drug target “for example, to prevent benign adenomas from developing further.” However, first the validity of the gene as a therapeutic target has to be tested in mice.
A good in vivo guinea pig for drugs
The researchers also expound that flies can be used to study candidate drug molecules to combat cancer. Drosophila would serve as an intermediate step between the in vitro phase and testing in vertebrates. On the one hand, this model has the in vitro advantages because many molecules can be tested at a minimum dose, and on the other, it shares the advantage of animals models because, as it is a living organism, toxic molecules or those with poor absorption could be omitted very quickly.
“If there are 2000 promising molecules among a million tested in vitro, instead of testing them in mice, Drosophila could offer a good alternative to identify the two or three that are most appropriate. Both time and costs would be reduced,” explains Casali.
With this aim, Casali has start collaborating with the group headed by Ernest Giralt (IRB Barcelona)—an authority on pharmacological chemistry and peptide design—to use flies to test new families of molecules against cancer.
Reference articles:
Iro/Irx transcription factors negatively regulate Dpp/TGF-b pathway activity during intestinal tumorigenesis
Òscar Martorell, Francisco M. Barriga, Anna Merlos-Suárez, Camille Stephan-Otto Attolini, Jordi Casanova, Eduard Batlle, Elena Sancho and Andreu Casali
EMBO Reports (2014 Oct 8). 10.15252/embr.201438622
Conserved mechanisms of tumorigenesis in the Drosophila adult midgut
Òscar Martorell, Anna Merlos-Suárez, Kyra Campbell, Francisco M. Barriga, Christo P. Christov,
Irene Miguel-Aliaga, Eduard Batlle, Jordi Casanova, Andreu Casali
PLoS One (2014 Feb) doi: 10.1371/journal.pone.0088413


 (1 votes)
(1 votes)


 The repair of cartilage and bone following damage remains a clinical challenge. Current cell-based therapies rely mostly on adult mesenchymal stromal cells, but the expansion of these into correctly differentiated and functionally competent chondrocytes, which give rise to cartilage and then bone, remains problematic. Here, Naoki Nakayama and colleagues develop a small molecule-based approach that mimics the embryonic somitic chondrogenesis programme and can be used to differentiate mouse embryonic stem cells (ESCs) into chondrocytes in vitro (p.
The repair of cartilage and bone following damage remains a clinical challenge. Current cell-based therapies rely mostly on adult mesenchymal stromal cells, but the expansion of these into correctly differentiated and functionally competent chondrocytes, which give rise to cartilage and then bone, remains problematic. Here, Naoki Nakayama and colleagues develop a small molecule-based approach that mimics the embryonic somitic chondrogenesis programme and can be used to differentiate mouse embryonic stem cells (ESCs) into chondrocytes in vitro (p.  The gene orthodenticle homologue 2 (Otx2) encodes a paired-type homeodomain transcription factor that is known to play a role in head morphogenesis. In the mouse, Otx2 is expressed in the anterior neurectoderm, where it is required for the differentiation of anterior neural tissues. Otx2 is also expressed in the anterior mesendoderm (AME) but its role here is unknown. On p.
The gene orthodenticle homologue 2 (Otx2) encodes a paired-type homeodomain transcription factor that is known to play a role in head morphogenesis. In the mouse, Otx2 is expressed in the anterior neurectoderm, where it is required for the differentiation of anterior neural tissues. Otx2 is also expressed in the anterior mesendoderm (AME) but its role here is unknown. On p.  Numerous transcription factors (TFs), including PU.1 and Scl, are known to play important roles during haematopoiesis, but how these act within wider TF networks is unclear. Now, Berthold Göttgens and colleagues use transcription activator-like effectors (TALEs) to manipulate the expression of PU.1 and Scl and determine how these TFs function during developmental haematopoiesis (p.
Numerous transcription factors (TFs), including PU.1 and Scl, are known to play important roles during haematopoiesis, but how these act within wider TF networks is unclear. Now, Berthold Göttgens and colleagues use transcription activator-like effectors (TALEs) to manipulate the expression of PU.1 and Scl and determine how these TFs function during developmental haematopoiesis (p.  Pattern formation during development often depends on the differential regulation of gene expression in response to a morphogen gradient, but how such gradients govern gene expression is unclear. A simplified view suggests that the morphogen activates a transcriptional activator, and that differential gene expression is dependent on the affinity or number of binding sites for this activator within target genes. However, this model does not account for bifunctional transcriptional effectors – those that function as activators and repressors – and has also been questioned by recent experimental results. Here, James Briscoe and colleagues describe a unifying mathematical model of morphogen-dependent gene expression that can explain recent counterintuitive findings (p.
Pattern formation during development often depends on the differential regulation of gene expression in response to a morphogen gradient, but how such gradients govern gene expression is unclear. A simplified view suggests that the morphogen activates a transcriptional activator, and that differential gene expression is dependent on the affinity or number of binding sites for this activator within target genes. However, this model does not account for bifunctional transcriptional effectors – those that function as activators and repressors – and has also been questioned by recent experimental results. Here, James Briscoe and colleagues describe a unifying mathematical model of morphogen-dependent gene expression that can explain recent counterintuitive findings (p.  The T-box family of transcription factors exhibits widespread involvement throughout development in all metazoans. Here, Virginia Papaioannou provides an overview of the key features of T-box transcription factors and highlights their roles and mechanisms of action during various stages of development and in stem/progenitor cell populations. See the Primer on p.
The T-box family of transcription factors exhibits widespread involvement throughout development in all metazoans. Here, Virginia Papaioannou provides an overview of the key features of T-box transcription factors and highlights their roles and mechanisms of action during various stages of development and in stem/progenitor cell populations. See the Primer on p. 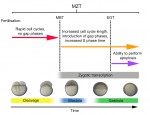 The initial phases of embryonic development occur in the absence of de novo transcription and are instead controlled by maternally inherited mRNAs and proteins. Following this period of transcriptional silence, zygotic transcription begins, the maternal influence on development starts to decrease, and dramatic changes to the cell cycle take place. Here, Steven Harvey and colleagues discuss recent work that is shedding light on the maternal to zygotic transition. See the Review on p.
The initial phases of embryonic development occur in the absence of de novo transcription and are instead controlled by maternally inherited mRNAs and proteins. Following this period of transcriptional silence, zygotic transcription begins, the maternal influence on development starts to decrease, and dramatic changes to the cell cycle take place. Here, Steven Harvey and colleagues discuss recent work that is shedding light on the maternal to zygotic transition. See the Review on p.  (No Ratings Yet)
(No Ratings Yet)