Yoshiki Sasai: stem cell Sensei
Posted by the Node, on 21 August 2014
This obituary first appeared in Development.
Stefano Piccolo looks back at the life and research of his friend and colleague Yoshiki Sasai.
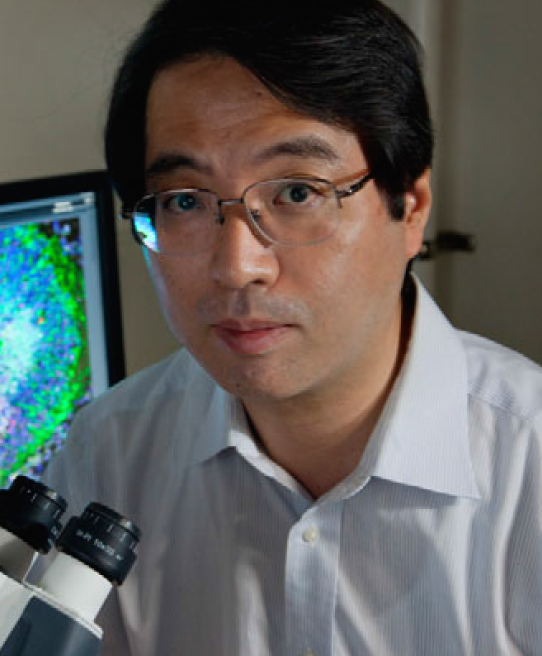 On 5 August 2014, Yoshiki Sasai died at the age of 52, near to the RIKEN Center for Developmental Biology in Kobe, Japan. This is the institute that he had helped to establish and painstakingly driven to become a world-leading research institution. The scientific community mourns the loss of one of its giants. But for those of us fortunate enough to have enjoyed Yoshiki’s friendship, or to have been transformed by his teachings and intellectual intensity, the grief of his death is devastating. He was a pioneer in the fields of developmental and stem cell biology, and here I will try to summarize and celebrate his exceptional legacy.
On 5 August 2014, Yoshiki Sasai died at the age of 52, near to the RIKEN Center for Developmental Biology in Kobe, Japan. This is the institute that he had helped to establish and painstakingly driven to become a world-leading research institution. The scientific community mourns the loss of one of its giants. But for those of us fortunate enough to have enjoyed Yoshiki’s friendship, or to have been transformed by his teachings and intellectual intensity, the grief of his death is devastating. He was a pioneer in the fields of developmental and stem cell biology, and here I will try to summarize and celebrate his exceptional legacy.
Yoshiki Sasai was born in Hyogo, Japan, and grew up playing baseball and cultivating the noble discipline of Kendo (the Japanese martial art that uses bamboo swords). Like other members of his family, he entered medical school, receiving his MD from the University of Kyoto in 1986. However, after completing an internship in internal medicine, he was frustrated by the limited biological understanding involved in routine clinical practice. He wanted to get at the root of the fundamental principles by which cells and tissues, particularly the brain, operate in health and disease. He therefore left the hospital halls to join the neurobiology laboratory of Shigetada Nakanishi as a PhD student. There, he became intrigued by how neural cells control their differentiation status. He was a gifted molecular biologist and, during his PhD, he identified mammalian HES family members and revealed their anti-neurogenic properties (Sasai et al., 1992).
For his postdoc, his instinct for cutting edge research brought him to southern California, to work on early Xenopus embryology in the laboratory of Eddy De Robertis (HHMI, UCLA). I vividly remember the day I first met Yoshiki. I was a newcomer to the De Robertis laboratory, just arrived in Los Angeles from Italy, and he was the senior postdoc of the lab. The respect he garnered from Eddy and the rest of the team was palpable. A few weeks after arriving at UCLA, Yoshiki had already cloned a new secreted factor, Chordin. This discovery held the key for what was then one of the biggest mysteries in developmental biology: the workings of the Spemann organizer. This fragment of the early embryo serves as signaling source to induce the nervous tissue and pattern the body plan, but its inner workings were yet unknown (De Robertis, 2009). Yoshiki had found that Chordin was expressed precisely in the organizer; moreover, injection of chordin mRNA was sufficient to generate a twin body, thus recapitulating the effects of transplantation of the organizer tissue. His first paper had already appeared in Cell (Sasai et al., 1994), and when I arrived in Los Angeles, Yoshiki was about to publish his second landmark discovery in Nature: the observation that neuralization of naïve cells induced by Chordin could be reversed by BMP4 (Sasai et al., 1995). Eddy made it clear that now it was my turn to venture into the biochemical mechanism for the Chordin-BMP antagonism. Of course, I had no idea where to start, so I was directed to Yoshiki for practical advice. At our first meeting, he effortlessly noted down a long list of ‘to-do’ experiments, with such rigor, logic and in-depth analysis of potential pitfalls and necessary controls to leave me at the same time captivated and intimidated. I still have those lab notes with me! By following that to-do list, it took only a few months to show that Chordin is not providing the embryo with any signal, but rather depriving it from the BMP signal, physically trapping BMPs in the extracellular space (Piccolo et al., 1996). Yoshiki was thus a central figure in establishing the notion in the late 1990s that the neural fate is a default state (Sasai and De Robertis, 1997).
Yoshiki treasured the essence of these discoveries: for embryonic cells, we often must remove external instructions, rather than adding them. Remarkably, with minimal external cues, cells apparently ‘know’ what they have to do, and can initiate entire developmental programs. By 1996, Yoshiki had returned to Japan to take up a position as associate and then full professor at Kyoto University. In 2003, he moved to Kobe, to the newly established RIKEN center. Through the years, he continued to work with the frog model system, providing seminal findings on the transcription factors involved in neural patterning and on the mechanisms responsible for sizing the embryo (Inomata et al., 2008). In parallel, he was using embryonic stem cells (ESCs), which he essentially considered the mammalian counterpart of the naïve frog embryo ectoderm cells that he had neuralized with Chordin. In his lab, lessons obtained in the frog model system were applied to ESCs, and vice versa (Sasai et al., 2008).
Yoshiki had a unique ability to see things clearly while others were left wandering in the dark. Creative intuition was then coupled with an ability to conceive straightforward experimental approaches, many requiring a patient, almost ritual, optimization in perfect Japanese style. He established a mouse ESC culture system containing minimal exogenous growth factors, a system that allowed cells to spontaneously slip into a telencephalic progenitor fate (Watanabe et al., 2005). Another major innovation was the discovery of an efficient method to culture human ESCs (Watanabe et al., 2007; Ohgushi et al., 2010). Until the mid-2000s, advances using human ESCs had been hampered by the fact that, unlike mouse ESCs, human ESCs are vulnerable to dissociation, and thus are lost through passaging. Yoshiki was not discouraged by this trivial, yet apparently insuperable, limitation: he systematically searched for chemical compounds able to sustain human ES passaging. One of these, a ROCK inhibitor, instantly did the trick and allowed human ESCs to survive through multiple passages; this compound is now routinely used in the field of ES and induced pluripotent stem cell (iPS) research.
It was the follow-up to his initial ESC work that made Yoshiki a scientific superstar. Inducing some specific types of neural cell types was too easy for Yoshiki; his challenge was to generate entire parts of the mammalian brain in the Petri dish. At that time, no techniques existed for generating organs from stem cells in culture. Earlier attempts to coax cells into organs by putting them on artificial scaffolds had been met with mixed success or had floundered. Keeping faith with his ‘less-is-more’ approach, Yoshiki not only removed growth factors from his ESC cultures; he also removed cells from the tissue culture plastic on which they are normally maintained, and grew them in suspension as floating spheres in Matrigel. He sensed that freeing stem cells from any external impediments would allow them to follow their own inner biological script (Sasai, 2013).
The results were awe-inspiring. Yoshiki and his colleagues showed that when their neuroepithelial ‘balls’ reached a given size, they started to form complex three-dimensional brain structures. In a series of seminal papers, they reported the generation in a dish of cortical tissue, of optic cups covered with a multilayered retina and of functioning pituitary glands (Eiraku et al., 2011, 2008; Suga et al., 2011). Yoshiki saw the emergence of these tissues under his microscope as a sort of living origami: he had only to adjust the initial conditions and then the tissue would continue its ordered folding and progressive assembly spontaneously, without external instructions. Size, topology and differentiation were all orchestrated by mysterious ‘self-organizing’ principles. When he repeated his famous optic-cup experiments with human ESCs, the induced eyes were, in fact, very human in terms of size and photoreceptor types (Nakano et al., 2012). Species-specific differences appeared to be intrinsically encrypted in cells, and blossomed under his in vitro organogenesis conditions. When I met him at a meeting and asked whether he would now rush to apply these astonishing findings to human retinal disorders, he answered “No way, the specialists should take credit for that. I will only be their consultant”. Indeed, he had already conceived what was for him the next frontier: getting a handle on the nature of tissue’s self-organizing scripts, which he sensed were a consequence of both a chemical and a biomechanical chain of events (Eiraku et al., 2012).
Those who met Yoshiki on the conference circuit will certainly remember that he was an outstanding lecturer. His talks were perfectly punctuated with wit and a sense of playfulness. Others may have met him in the evenings of the many symposia organized at his institute as the exuberant RIKEN bartender (he expected tips, too!). The more private Yoshiki was a positive, charismatic and generous man. And it was impossible to resist being fascinated by a man who could talk passionately about so many diverse things, whether the topic was Japanese culture, the delicacies of an Italian risotto or evolutionary plasticity.
Hopefully, Yoshiki’s ‘eyes-in-vitro’ will help treating blindness in the future. But whatever the translational legacy of his work, he was certainly a visionary scientist, who opened our eyes to the wonders of developmental and stem cell biology and its potential for mankind.
Goodbye Sensei, we miss you.
References:
De Robertis, E. M. (2009). Spemann’s organizer and the self-regulation of embryonic fields. Mech. Dev. 126, 925-941.
Eiraku, M., Watanabe, K., Matsuo-Takasaki, M., Kawada, M., Yonemura, S., Matsumura, M., Wataya, T., Nishiyama, A., Muguruma, K. and Sasai, Y. (2008). Self-organized formation of polarized cortical tissues from ESCs and its active manipulation by extrinsic signals. Cell Stem Cell 3, 519-532.
Eiraku, M., Takata, N., Ishibashi, H., Kawada, M., Sakakura, E., Okuda, S., Sekiguchi, K., Adachi, T. and Sasai, Y. (2011). Self-organizing optic-cup morphogenesis in three-dimensional culture. Nature 472, 51-56.
Eiraku, M., Adachi, T. and Sasai, Y. (2012). Relaxation-expansion model for self driven retinal morphogenesis: a hypothesis from the perspective of biosystems dynamics at the multi-cellular level. BioEssays 34, 17-25.
Inomata, H., Haraguchi, T. and Sasai, Y. (2008). Robust stability of the embryonic axial pattern requires a secreted scaffold for chordin degradation. Cell 134, 854-865.
Nakano, T., Ando, S., Takata, N., Kawada, M., Muguruma, K., Sekiguchi, K., Saito, K., Yonemura, S., Eiraku, M. and Sasai, Y. (2012). Self-formation of optic cups and storable stratified neural retina from human ESCs. Cell Stem Cell 10, 771-785.
Ohgushi, M.,Matsumura, M., Eiraku, M.,Murakami, K., Aramaki,T., Nishiyama, A., Muguruma, K., Nakano, T., Suga, H. and Ueno, M. et al. (2010). Molecular pathway and cell state responsible for dissociation-induced apoptosis in human pluripotent stem cells. Cell Stem Cell 7, 225-239.
Piccolo, S., Sasai, Y., Lu, B. and De Robertis, E. M. (1996). Dorsoventral patterning in Xenopus: inhibition of ventral signals by direct binding of chordin to BMP-4. Cell 86, 589-598.
Sasai, Y. (2013). Next-generation regenerative medicine: organogenesis from stem cells in 3D culture. Cell Stem Cell 12, 520-530.
Sasai, Y. and De Robertis, E. M. (1997). Ectodermal patterning in vertebrate embryos. Dev. Biol. 182, 5-20.
Sasai, Y., Kageyama, R., Tagawa, Y., Shigemoto, R. and Nakanishi, S. (1992). Two mammalian helix-loop-helix factors structurally related to Drosophila hairy and Enhancer of split. Genes Dev. 6, 2620-2634.
Sasai, Y., Lu, B., Steinbeisser, H., Geissert, D., Gont, L. K. and De Robertis, E. M. (1994). Xenopus chordin: a novel dorsalizing factor activated by organizer-specific homeobox genes. Cell 79, 779-790.
Sasai, Y., Lu, B., Steinbeisser, H. and De Robertis, E. M. (1995). Regulation of neural induction by the Chd and Bmp-4 antagonistic patterning signals in Xenopus. Nature 376, 333-336.
Sasai, Y., Ogushi, M., Nagase, T. and Ando, S. (2008). Bridging the gap from frog research to human therapy: a tale of neural differentiation in Xenopus animal caps and human pluripotent cells. Dev. Growth Differ. 50 Suppl. 1, S47-S55.
Suga, H., Kadoshima, T., Minaguchi, M., Ohgushi, M., Soen, M., Nakano, T., Takata, N., Wataya, T., Muguruma, K. and Miyoshi, H. et al. (2011). Self formation of functional adenohypophysis in three dimensional culture. Nature 480, 57-62.
Watanabe, K., Kamiya, D., Nishiyama, A., Katayama, T., Nozaki, S., Kawasaki, H., Watanabe, Y., Mizuseki, K. and Sasai, Y. (2005). Directed differentiation of telencephalic precursors from embryonic stem cells. Nat. Neurosci. 8, 288-296.
Watanabe, K., Ueno, M., Kamiya, D., Nishiyama, A., Matsumura, M., Wataya, T., Takahashi, J. B., Nishikawa, S., Nishikawa, S.-i. and Muguruma, K. et al. (2007). A ROCK inhibitor permits survival of dissociated human embryonic stem cells. Nat. Biotechnol. 25, 681-686


 (2 votes)
(2 votes)
 (No Ratings Yet)
(No Ratings Yet)



 (8 votes)
(8 votes)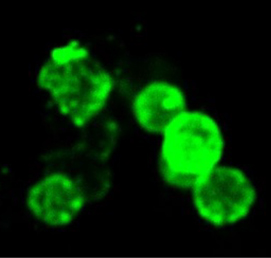 The emergence of haematopoietic stem cells (HSCs) during early mammalian development is crucial for the formation of all blood cell lineages. Previous studies indicate that Runx1 is required for the endothelial-haematopoietic transition that gives rise to definitive HSCs; however, this transition occurs in multiple steps and the precise stage at which Runx1 is required has been unclear. In this issue (p.
The emergence of haematopoietic stem cells (HSCs) during early mammalian development is crucial for the formation of all blood cell lineages. Previous studies indicate that Runx1 is required for the endothelial-haematopoietic transition that gives rise to definitive HSCs; however, this transition occurs in multiple steps and the precise stage at which Runx1 is required has been unclear. In this issue (p.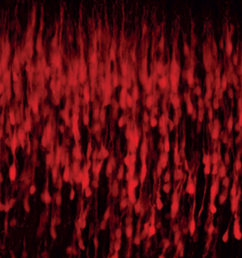 The formation of the mammalian neocortex relies on the migration of projection neurons into specific layers, which is in turn regulated by cyclin-dependent kinase 5 (Cdk5). Cdk5 has previously been shown to interact with the G protein-coupled receptor serotonin 6 receptor (5-HT6R); however, whether and how this interaction might be important for neocortical migration remains unclear. Now, on p.
The formation of the mammalian neocortex relies on the migration of projection neurons into specific layers, which is in turn regulated by cyclin-dependent kinase 5 (Cdk5). Cdk5 has previously been shown to interact with the G protein-coupled receptor serotonin 6 receptor (5-HT6R); however, whether and how this interaction might be important for neocortical migration remains unclear. Now, on p.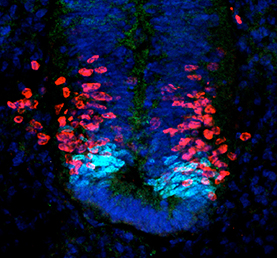 The sonic hedgehog (Shh) signalling pathway is a crucial mediator of cell proliferation, morphogenesis and fate, influencing the development and homeostasis of multiple organ systems during all stages of life. A central premise of Shh signalling is that the receptor patched 1 (Ptch1) mediates the Shh response, and that this response is greatest when Ptch1 is absent, allowing the release of pathway activator smoothened (Smo) and the eventual activation of Gli proteins. In this issue (p.
The sonic hedgehog (Shh) signalling pathway is a crucial mediator of cell proliferation, morphogenesis and fate, influencing the development and homeostasis of multiple organ systems during all stages of life. A central premise of Shh signalling is that the receptor patched 1 (Ptch1) mediates the Shh response, and that this response is greatest when Ptch1 is absent, allowing the release of pathway activator smoothened (Smo) and the eventual activation of Gli proteins. In this issue (p.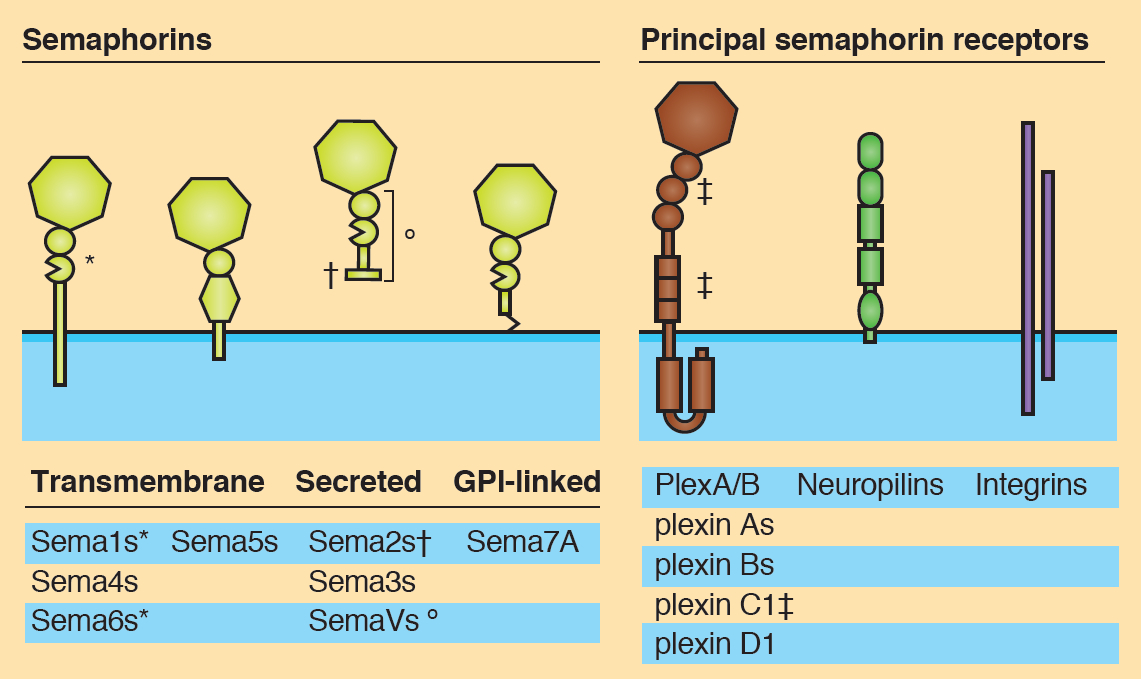 Semaphorins are secreted and membrane-associated proteins that regulate many cellular and developmental processes. In this article and accompanying poster, Jongbloets and Pasterkamp review the molecular biology of semaphorin signalling in different contexts. See the Review on p.
Semaphorins are secreted and membrane-associated proteins that regulate many cellular and developmental processes. In this article and accompanying poster, Jongbloets and Pasterkamp review the molecular biology of semaphorin signalling in different contexts. See the Review on p.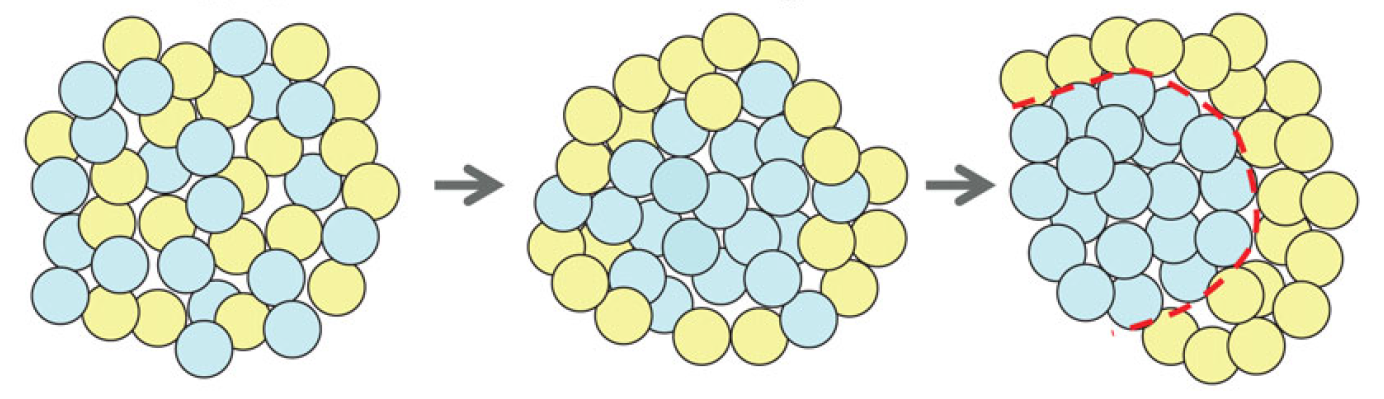 The separation of the embryo into physically distinct regions is one of the most important processes in development. François Fagotto discusses various boundary formation models and summarizes recent studies that have examined this process at the cellular level. See the Review on p.
The separation of the embryo into physically distinct regions is one of the most important processes in development. François Fagotto discusses various boundary formation models and summarizes recent studies that have examined this process at the cellular level. See the Review on p.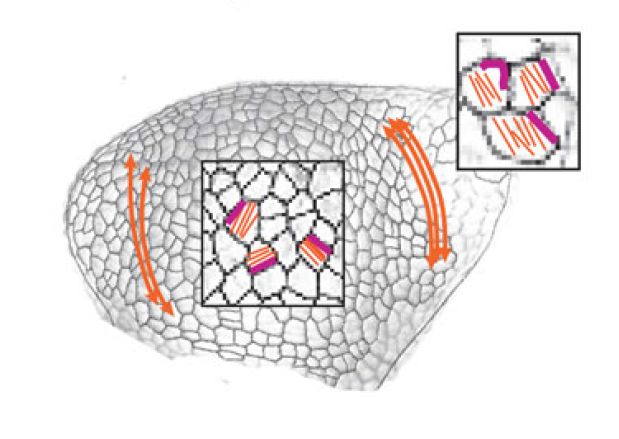 Jeffrey Axelrod and Dominique Bergmann report from The Company of Biologists workshop ‘Coordinating Cell Polarity’, which brought together researchers working on plant and animal systems to discuss the emerging themes in the field. See the Meeting Review on p.
Jeffrey Axelrod and Dominique Bergmann report from The Company of Biologists workshop ‘Coordinating Cell Polarity’, which brought together researchers working on plant and animal systems to discuss the emerging themes in the field. See the Meeting Review on p.