Santa Cruz Developmental Biology Meeting
Posted by Katherine Brown, on 15 August 2012
I’ve just returned from this year’s Santa Cruz Developmental Biology meeting. Some of you may have seen the tweets I was sending out from there (see Storify for the collected set), but a combination of limited ability to multitask and limited laptop battery life meant I didn’t cover all the talks. So to add to what I missed, and for those who prefer more than 140 characters of coverage, here’s a summary of some of the highlights.
SCDB is a bi-annual broad developmental biology meeting, held at the beautiful wooded campus of UC Santa Cruz. With only around 180 participants, it’s a fairly small and very friendly event, with plenty of opportunity for informal discussions. This year, it was organised by Bob Goldstein, Amander Clark and John Tamkun, and topics discussed at the meeting ranged from the evolution of segmentation in arthropods (Mike Akam, University of Cambridge) to ligand/receptor interactions in axon guidance (Elke Stein, Yale University), with pretty much every model organism, tissue and process in between.
Despite covering the whole breadth of the field, there were some definite themes running through the meeting – aside those defined by the program. Multiple talks dealt with the germline: how you make it, put it in the right place and maintain it. Both Diana Laird (UCSF) and Jeremy Nance (NYU Skirball Institute) focussed on the earliest stages of gonad formation in the mouse and the worm respectively. Diana’s work looks at the interactions between migrating primordial germ cells and the various niches they encounter during migration through the embryo, while Jeremy presented data on the mechanisms by which C. elegans germ cells are internalised during gastrulation. Moving to later stages of the nematode, Jane Hubbard (NYU Skirball) demonstrated that nutrient status is a key determinant in regulating germ cell proliferation. The importance of environmental signals was echoed by Timothy Kelliher (Walbot lab, Stanford), who showed that hypoxia triggers germ cell formation in maize (where there is no pre-defined germline as in animals). Perhaps most spectacularly, Bruce Draper (UC Davis) presented his latest work on how mature germ cells influence sex determination in zebrafish: look out for his upcoming paper on sex-changing fish!
As is becoming standard in developmental biology meetings these days, talks were filled with beautiful movies of everything from early stage Drosophila embryos (Dan Kiehart, Duke University and Jen Zallen, Sloan Kettering) to regenerating axolotl (Saori Haigo, Center for Regenerative Therapies Dresden and UCSF) and mouse neural tube closure (Lee Niswander, University of Colorado Denver). But all were (or at least claimed to be!) put in the shade by Eric Betzig’s keynote lecture on super-resolution in vivo imaging: for unprecedented intracellular resolution in developing tissues, the future apparently lies with the Bessel beam.
In a third recurring theme, several speakers discussed the regulation of cell division and its impact on cell fate. Asako Sugimoto (Tohoku University) showed beautiful work on spindle assembly in C. elegans, directly comparing oocyte meiotic division, where the spindle is small and acentrosomic, with the following first zygotic mitosis, in which both centrosomes and chromatin direct microtubule assembly and spindle formation. Laurie Smith (UCSD) uses stomatal development in maize as a model to study asymmetric division, and shared her latest insights into the pathways regulating this process in plants. Finally, Roel Nusse (Stanford University) presented a tour-de-force study on the regulation of embryonic stem cell division by Wnt signalling – another paper to keep an eye out for in the future.
 Away from the lecture theatre, the poster sessions were very lively and interactive – congratulations to poster prize winners Shawn Chavez (human blastocyst development), Harshani Peiris (planarian stem cells) and Jacqueline Tabler (ciliopathy models in mouse), although I’m sure there could have been many more winners among the great posters I saw. The Friday evening wine tasting and social session was also enlivened by the surprise entertainment: I’m not sure how many companies get called up and asked if they would sponsor a Mariachi band, but Nikon stepped up to the plate and delivered.
Away from the lecture theatre, the poster sessions were very lively and interactive – congratulations to poster prize winners Shawn Chavez (human blastocyst development), Harshani Peiris (planarian stem cells) and Jacqueline Tabler (ciliopathy models in mouse), although I’m sure there could have been many more winners among the great posters I saw. The Friday evening wine tasting and social session was also enlivened by the surprise entertainment: I’m not sure how many companies get called up and asked if they would sponsor a Mariachi band, but Nikon stepped up to the plate and delivered.
All of this means that the next set of organisers for this meeting – Jeremy Nance, Diana Laird and Amy Ralston – have a lot to live up to: not just in topping the Mexican minstrels, but mainly in putting together a fantastic and diverse set of speakers and fostering a welcoming and collaborative atmosphere. Look out for SCDB2014: I’m sure it’ll be a good’un!


 (9 votes)
(9 votes)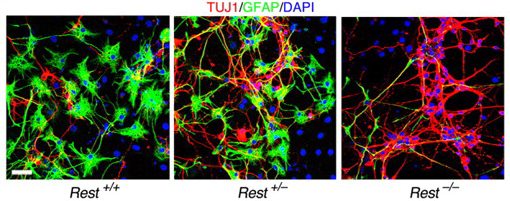
 Gene expression is translationally regulated during many developmental processes. Translation is mainly controlled at the initiation step, which involves recognition of the mRNA 5′ cap structure by the eukaryotic initiation factor 4E (eIF4E). Eukaryotic genomes often encode several eIF4E paralogues but their biological relevance is largely unknown. Here (
Gene expression is translationally regulated during many developmental processes. Translation is mainly controlled at the initiation step, which involves recognition of the mRNA 5′ cap structure by the eukaryotic initiation factor 4E (eIF4E). Eukaryotic genomes often encode several eIF4E paralogues but their biological relevance is largely unknown. Here (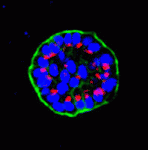 Mammary epithelial cells undergo structural and functional differentiation at late pregnancy and parturition to initiate milk secretion. TGF-β and prolactin signalling act antagonistically to regulate this process but what coordinates these pathways? On
Mammary epithelial cells undergo structural and functional differentiation at late pregnancy and parturition to initiate milk secretion. TGF-β and prolactin signalling act antagonistically to regulate this process but what coordinates these pathways? On 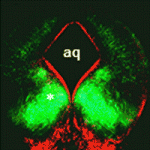 During the development of the central nervous system, neurons and/or neuronal precursors travel along diverse routes from the ventricular zones of the developing brain and integrate into specific brain circuits. Neuronal migration has been extensively studied in the forebrain but little is known about this key developmental event in the embryonic midbrain (mesencephalon). On
During the development of the central nervous system, neurons and/or neuronal precursors travel along diverse routes from the ventricular zones of the developing brain and integrate into specific brain circuits. Neuronal migration has been extensively studied in the forebrain but little is known about this key developmental event in the embryonic midbrain (mesencephalon). On 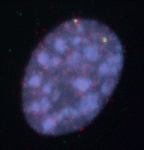 Anterior-posterior patterning of both the primary embryonic axis and the secondary body axis (limbs and digits) in mammals requires regulated Hox expression. Polycomb-mediated changes in chromatin structure control Hox expression during the first patterning event but are they also involved in the second? Here (
Anterior-posterior patterning of both the primary embryonic axis and the secondary body axis (limbs and digits) in mammals requires regulated Hox expression. Polycomb-mediated changes in chromatin structure control Hox expression during the first patterning event but are they also involved in the second? Here (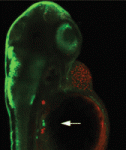 Mutations in the human Shwachman-Bodian-Diamond syndrome (SBDS) gene, which functions during maturation of the large 60S ribosomal subunit, cause a disorder characterised by exocrine pancreatic insufficiency, chronic neutropenia and skeletal defects. Steven Leach and colleagues have now refined a zebrafish model of this ‘ribosomopathy’ (see
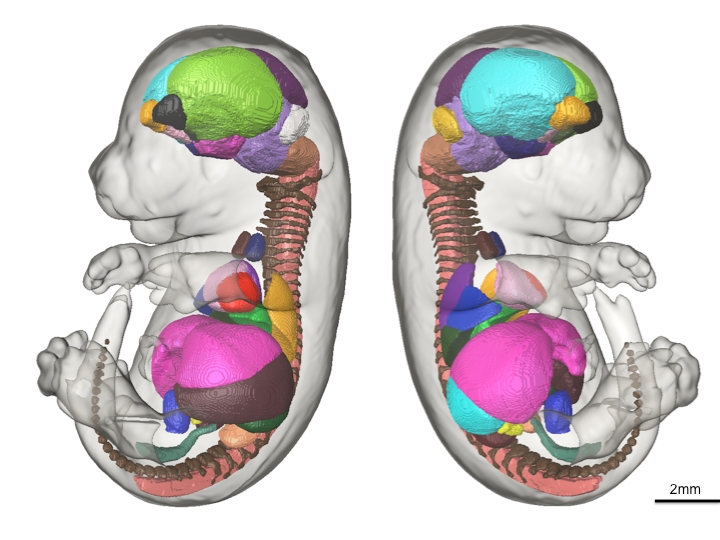
Mutations in the human Shwachman-Bodian-Diamond syndrome (SBDS) gene, which functions during maturation of the large 60S ribosomal subunit, cause a disorder characterised by exocrine pancreatic insufficiency, chronic neutropenia and skeletal defects. Steven Leach and colleagues have now refined a zebrafish model of this ‘ribosomopathy’ (see 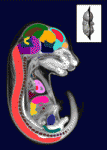 The sequence and location of every gene in the human genome is now known but our understanding of the relationships between human genotypes and phenotypes is in its infancy. To better understand the role of every gene in the development of an individual, the International Mouse Phenotyping Consortium aims to phenotype targeted gene knockout mice throughout the genome (∼23,000 genes). Because many of these mice will be embryonic lethal, methods for phenotyping mouse embryos are needed. Michael Wong and colleagues are developing such an approach and, on
The sequence and location of every gene in the human genome is now known but our understanding of the relationships between human genotypes and phenotypes is in its infancy. To better understand the role of every gene in the development of an individual, the International Mouse Phenotyping Consortium aims to phenotype targeted gene knockout mice throughout the genome (∼23,000 genes). Because many of these mice will be embryonic lethal, methods for phenotyping mouse embryos are needed. Michael Wong and colleagues are developing such an approach and, on  The tenth annual RIKEN Center for Developmental Biology symposium ‘Quantitative Developmental Biology’ held in March 2012 covered a range of topics. As reviewed by Davidson and Baum, the studies presented at the meeting shared a common theme in which a combination of physical theory, quantitative analysis and experiment was used to understand a specific cellular process in development. See the Meeting Review on p.
The tenth annual RIKEN Center for Developmental Biology symposium ‘Quantitative Developmental Biology’ held in March 2012 covered a range of topics. As reviewed by Davidson and Baum, the studies presented at the meeting shared a common theme in which a combination of physical theory, quantitative analysis and experiment was used to understand a specific cellular process in development. See the Meeting Review on p. 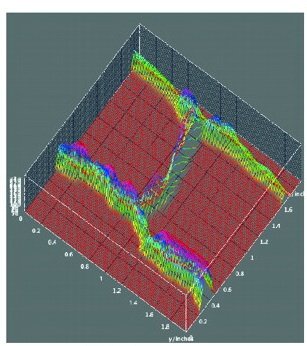 Meyerowitz and colleagues present a glossary of image analysis terms to aid biologists and discuss the importance of robust image analysis in developmental studies.
Meyerowitz and colleagues present a glossary of image analysis terms to aid biologists and discuss the importance of robust image analysis in developmental studies.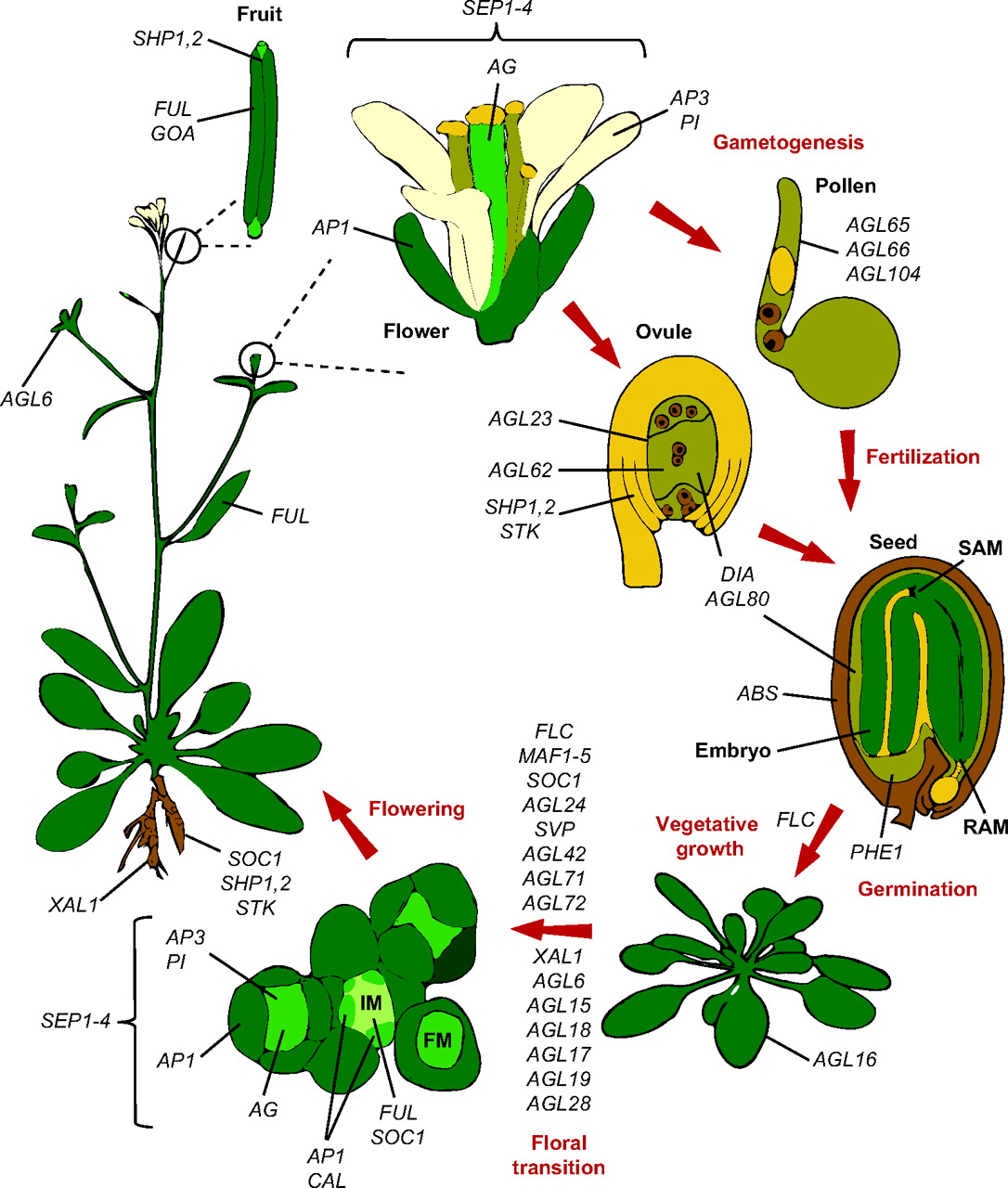 Members of the MADS-box transcription factor family play essential roles in almost every developmental process in plants. Kaufmann and colleagues review recent findings on MADS-box gene functions in Arabidopsis and discuss the evolutionary history and functional diversification of this gene family in plants.
Members of the MADS-box transcription factor family play essential roles in almost every developmental process in plants. Kaufmann and colleagues review recent findings on MADS-box gene functions in Arabidopsis and discuss the evolutionary history and functional diversification of this gene family in plants.
 (No Ratings Yet)
(No Ratings Yet)
