Regeneration thwarts ageing in newts
Posted by Tsonis, on 17 January 2016
Konstantinos Sousounis and Panagiotis A. Tsonis
The human eye is built to deliver the sense of vision. The eye lens is one of the organs playing role in focusing the light to the retina. Lens injury or disease leads to blurriness or even blindness in human patients. This is not the case for newts. These small amphibian urodeles can have their entire lens removed, and yet they can regenerate a brand new one! In fact, regeneration in newts is not limited to the lens, but extends to many organs from brain, heart, spinal cord to limbs and tails. Ever since the discovery that newts can regenerate numerous body parts, many scientists have focused on deciphering the mechanism underlying regeneration (Sanchez Alvarado and Tsonis, 2006; Tsonis and Fox, 2009). The mechanism as a whole is still unclear, despite the numerous findings of the various molecular interactions underlying regeneration. In addition to the mechanism of regeneration, two important questions remained unanswered:
(1) Knowing that body parts can be regenerated, how many times can an organ be regenerated?
(2) Does age affect/limit the regeneration capacity of an organism?
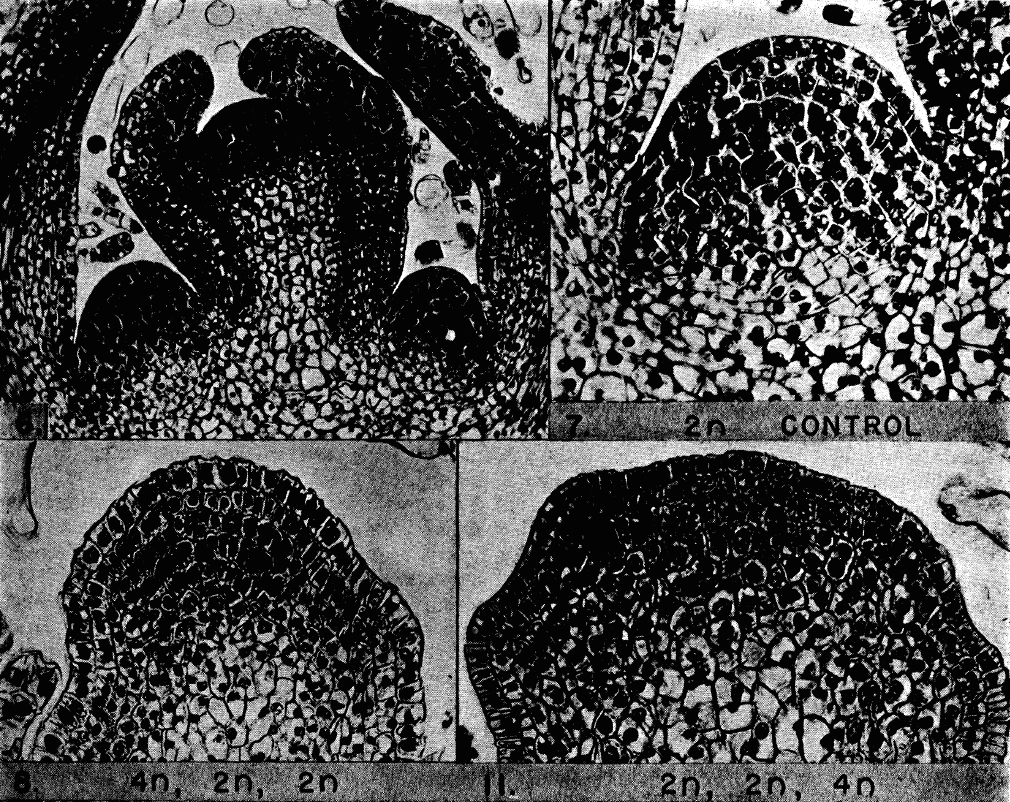
These two questions cannot be answered experimentally by the commonly used “injury induction-time point-collection-analysis” paradigm. These experiments need to include repetitive insults to the same animals over a prolonged period of time and will, thus, require a commitment from the researcher to take extremely good care of the animals while maintaining these experimental colonies for a long time. Now, this is not an easy task. To put things into perspective, keep in mind that newts are kept in water, and water is an environment that without prompt cleaning can get pretty dirty quite fast. Diseases can also spread, so that one infected newt can easily wipe out the rest of a group in a tank. So, rule number one: Extreme super animal care! Something you get used to after the first couple of weeks in an amphibian lab. Many other crucial issues arise. For how long do you need to be super vigilant for newts being used in an ageing study? When does a newt age? How many years can a newt live [keeping in mind that a mouse is considered aged after its first birthday]? Well, according to Goin et al, 1978, newts can live as long as 25 years. That is almost a three-decade-long newt ageing study, which would also mean that an actual study designed to investigate the effect(s) of ageing on regeneration in newts would span way beyond the time typically needed for students to obtain their Ph.D. or post-doctoral researchers to complete their fellowship. That is not all; surviving surgery is challenging and requires a lot of post-surgical care on the animals that will be operated twice every year since 6 months is sufficient period of time for an organ to be fully regenerated and for a newt to recover well before the next surgery. Taken together, the requirements for the newt ageing study make it seem unimaginable that a person might actually carry such a study and investigate how repetitive regeneration is affected throughout the lifespan of a newt (not to mention receiving funding). Unimaginable but not impossible for Goro Eguchi who in 1994 at the National Institute for Basic Biology at Okazaki, Japan took on this task and put a start to this project. The model chosen was that of lens regeneration in the Japanese newt. One of the main advantages of this model is the fact that the lens is in an enclosed environment (the eye), and when removed through a small slit in the cornea the injury is local and the wound closure heals very fast. The lens is regenerated from the iris via a process called transdifferentiation, whereby the uniformly pigmented cells of the dorsal iris undergo a complete morphological, transcriptomic and molecular transformation to regenerate the lost lens (Henry and Tsonis, 2010). The process is considered complete after 6 months and the newly regenerated lens is identical to the original. In our study, the first lens removal was performed on 14-year-old newts. These newts were 30-year-old at the end of the study, and they had their lenses removed 18 times over a period of 16 years. Then, at the end of the study, our team (based at the University of Dayton and the Sanford Burnham Prebys Medical Discovery Institute) had these repeatedly regenerated lenses analyzed for the first time. The conclusion was clear: repeated regeneration or ageing did not affect regenerative capacity in newts (Eguchi et al, 2011). Fiber structure and expression of crystallin and lens-specific genes were similar to never-regenerated lenses from young newts. Even though that study was groundbreaking, many questions were left unanswered. What we wanted to investigate next was whether the regenerated lenses were aged and if the process of regeneration had affected the underlying genetic profile. To get the most out of the tested samples, we decided to use RNA-sequencing. We collected the repeatedly regenerated lenses (now 19 times regenerated; 18 years after the initiation of the project), the dorsal iris (the source of regeneration) and the tails (to serve as controls since this tissue was never removed or regenerated in these animals). We also collected the same tissues from young newts that never regenerated the lens. Altogether the samples totaled 30, with n = 5 animals per each group. In the absence of a genome, de novo assembly of the newt transcriptome was required. With 30 samples at hand, multiplied by 150 million reads for each sample, we were left with a complex task that even super computers found challenging to execute this de novo assembly. With suggestions from Tritiny while using their program, the task was completed and the final gene expression data was emailed to us. Holding our breath, we checked the data and BINGO: zero genes differentially expressed in Experimental Lenses versus Control Lenses! After an overwhelming feeling of joy we went onto data analysis. The tail samples were found to have the most differentially regulated genes, followed by the iris and then the lenses with none. At first glance, it was obvious that there was a gradient of age-dependent regulation based on the degree of regeneration (or degree of cell replacement in the case of iris if you prefer) that each tissue had experienced. To clarify how we came about knowing that the tail and iris samples were indeed aged, we used previous studies on other vertebrates and even flies that had identified, for example, that down-regulation of mitochondrial genes is a common signature of ageing. We identified this pattern and other gene patterns too in our samples. That told us that there was a robust transcriptional program that is utilized with every single lens removal, which guarantees the completion of the process with the outcome of a perfectly identical lens (Sousounis et al, 2015). Another interesting finding was that the iris, the tissue that regenerates the lens, had shown signs of ageing but the regenerate, the lens, did not! That showed us that the transdifferentiation process can be really compared to an “extreme makeover – cell edition.” Considering that human lenses are vastly affected by ageing and that their composition is really not that different from newt lenses, our findings hold great translational promise for future studies. The newt is not only capable of reprogramming adult cells to dedifferentiate and regenerate, but also has inherent mechanisms to deactivate pathways that lead to ageing. These findings could pave the way to avenues of experimentation towards the prevention of the process of ageing.
References
Eguchi G, Eguchi Y, Nakamura K, Yadav MC, Millán JL, Tsonis PA. Regenerative capacity in newts is not altered by repeated regeneration and ageing. Nat Commun. 2011; 2:384. doi: 10.1038/ncomms1389.
Goin, C. J., Goin, O. B. & Zug, G. R. Introduction to Herpetology 3rd edn (W.H. Freeman, 1978).
Henry JJ, Tsonis PA. Molecular and cellular aspects of amphibian lens regeneration. Prog Retin Eye Res. 2010; 29(6):543-55. doi: 10.1016/j.preteyeres.2010.07.002.
Sánchez Alvarado A, Tsonis PA. Bridging the regeneration gap: genetic insights from diverse animal models. Nat Rev Genet. 2006; 7(11):873-84.
Sousounis K, Qi F, Yadav MC, Millán JL, Toyama F, Chiba C, Eguchi Y, Eguchi G, Tsonis PA. A robust transcriptional program in newts undergoing multiple events of lens regeneration throughout their lifespan. eLife. 2015; 4. pii: e09594. doi: 10.7554/eLife.09594
Tsonis PA, Fox TP. Regeneration according to Spallanzani. Dev Dyn. 2009; 238(9):2357-63. doi: 10.1002/dvdy.22057


 (17 votes)
(17 votes) (No Ratings Yet)
(No Ratings Yet)

 (1 votes)
(1 votes) Marcos and Julia were a brilliant young couple who instantly made the laboratory smart and fun. Marcos took up cancer research, the second person to ever work on this disease in the laboratory. He loved everything about conducting science and he was ambitious. He worked crazy hours. With a laconic Argentinian accent, he kept up a running science conversation that lasted until now. Julia worked on separate topics and they lived balanced lives. Marcos loved board sports from snowboarding to kite- and wind-surfing and was superb at them. Marcos and Julia made friends easily. The Facebook postings and the emails we’ve received and the conversations we’ve had over the past few days are a reminder just how much they were loved. We all rooted for their success because they were really good people.
Marcos and Julia were a brilliant young couple who instantly made the laboratory smart and fun. Marcos took up cancer research, the second person to ever work on this disease in the laboratory. He loved everything about conducting science and he was ambitious. He worked crazy hours. With a laconic Argentinian accent, he kept up a running science conversation that lasted until now. Julia worked on separate topics and they lived balanced lives. Marcos loved board sports from snowboarding to kite- and wind-surfing and was superb at them. Marcos and Julia made friends easily. The Facebook postings and the emails we’ve received and the conversations we’ve had over the past few days are a reminder just how much they were loved. We all rooted for their success because they were really good people.