Forgotten classics: Making a monotreme
Posted by the Node, on 17 October 2017
T. Thomson Flynn and J.P. Hill. 1939. The Development of the Monotrema – Part IV. Growth of the Ovarian Ovum, Maturation, Fertilisation and Early Cleavage. Transactions of the Zoological Society of London, 24, 445-623.
T. Thomson Flynn and J.P. Hill. 1947. The Development of the Monotrema – Part VI. The Later Stages of Cleavage and the Formation of the Primary Germ-layers. Transactions of the Zoological Society of London, 26, 1-151.
Recommended by Guojun Sheng, IRCMS, Kumamoto University, Japan.
Mammals alive today are split into three groups: the placentals (like us), the marsupials (like Skippy), and, distantly related to the rest, the monotremes. Monotremes come in two kinds: echidnas, “shambling, animated pincushions” whose young are known as ‘puggles’, and platypuses, famous for their chimeric collection of weird anatomical features (their species name, Ornithorhyncus anatinus, means ‘duck-like bird-snout’).
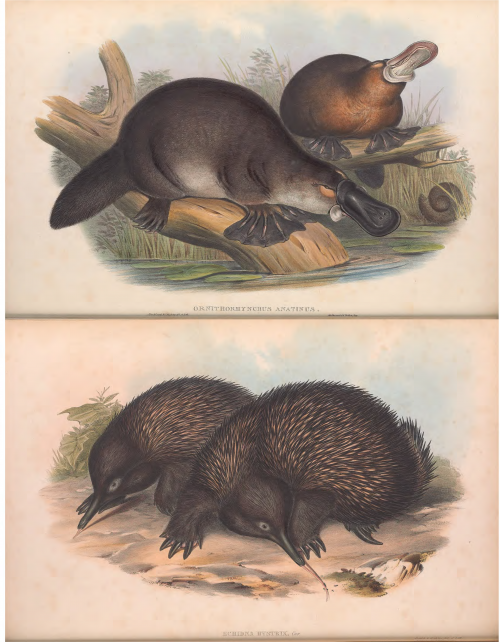

In addition to their odd appearance, monotremes stand out compared to other mammals when it comes to reproduction: rather than giving birth to live young, they lay eggs, a trait shared with birds and reptiles, which together form the sister group to mammals (mammals, birds and reptiles together form the amniotes). Given their phylogenetic position and mix of biological traits, monotremes might tell us a lot about mammalian evolution from a reptile-like ancestor, keeping in mind of course that they have not stayed still since they split from the rest of the mammals perhaps a hundred and sixty million years ago. From the perspective of developmental biology, the question is how an ancestral, reptile-like mode of embryogenesis was transformed into a mammal-like one: do monotremes develop more like other mammals, or more like birds and reptiles? (Another proviso here: ‘mammal-like’ rather obscures the diversity seen in early mammalian embryogenesis, as explored by Guojun Sheng, Marilyn Renfree, Berenika Plusa and others, but monotremes might in fact prove a useful example to understand this diversity.)
Answering these questions is not easy. Some animals are more amenable to embryological investigation than others, and monotremes are exceptionally intransigent in this regard: they can be hard to find, do not breed well in colonies (though this might be getting easier), are not really prodigious breeders in any case, and have embryos that are difficult to collect and preserve. ‘Non-model’ doesn’t quite cover it.
Nevertheless, monotreme embryology has a long history. The Scottish zoologist W.H. Caldwell was one of first Western scientists to unequivocally describe monotreme egg-laying (read more in Brian Hall’s paradoxical history of the platypus), in a famously terse telegram sent while on an expedition in Queensland in 1884 :
“Monotremes oviparous, ovum meroblastic”
‘Meroblastic’ describes the incomplete cleavage of the early embryo, which reptiles and birds do too.
Caldwell’s expeditions were torrid affairs –
“Crossing the Maclntyre River in a flood, the buggy was upset, and its contents washed away. The two following months were lost through the effects of a fever…”
But not just for him: the Aboriginal Australians employed by Caldwell to catch the animals were treated horrendously, according to the introduction to his paper describing the expedition (it’s quite a shocking read, but perhaps not an altogether surprising one).
As Caldwell describes, collecting monotreme samples in the wild was a difficult and often brutal business. Since a lot of embryonic development occurs in utero, to get earlier stages you have to kill the mothers. Platypuses posed particular problems, as described in a later account:
“…it is extremely difficult, even for residents in Australia, to procure material necessary for an investigation into the development of the egg of Ornithorhynchus….The animal itself, though pretty widely distributed, and probably still far from becoming extinct, is to be found, in any one locality, only in comparatively small numbers.
The eggs, when laid, are deposited in a burrow which it is far from easy to locate, and whose opening up involves a considerable amount of labour, since, apart from its great length, the river-bank in which it is situated is commonly enough permeated by tree-roots. And when at length the actual dwelling-chamber or nest is successfully opened up, no reward at all may be forthcoming, or the material which is obtained may be unsuitable for the immediate purpose in view.
Even when it is the intra-uterine stages of the egg which are required…the difficulties are nearly as great. The animal is extremely shy and difficult [to] approach. They are occasionally, but rarely, captured as an incident in net-fishing in the larger rivers: otherwise they are practically only obtainable with the gun. During the breeding season, however, the pregnant female appears to keep much more closely to the burrow, so that one may then commonly enough shoot five or six males to one female.”
This was published in 1908 by J.T Wilson and one J.P. Hill, who with T.T. Flynn wrote the two hefty monographs that are the subject of this Forgotten Classics piece – Volume IV and Volume VI of the Transactions of the Zoological Society of London’s Development of the Monotrema series.
Hill was born in Edinburgh but moved early in his career to Sydney (where he met Flynn who was studying there), before returning to the UK to University College London and rising to chair of embryology and histology. Flynn was an Australian who moved to Tasmania from Sydney to become a professor of biology and an eminent naturalist. In Tasmania, as well as collecting monotreme samples, he fathered Errol Flynn (yes that Errol Flynn, the actor), but after his funding was cut he moved to Queens University in Belfast. There he served as a fire martial and casualty clearing officer in the Second World War, for which he was awarded an MBE.

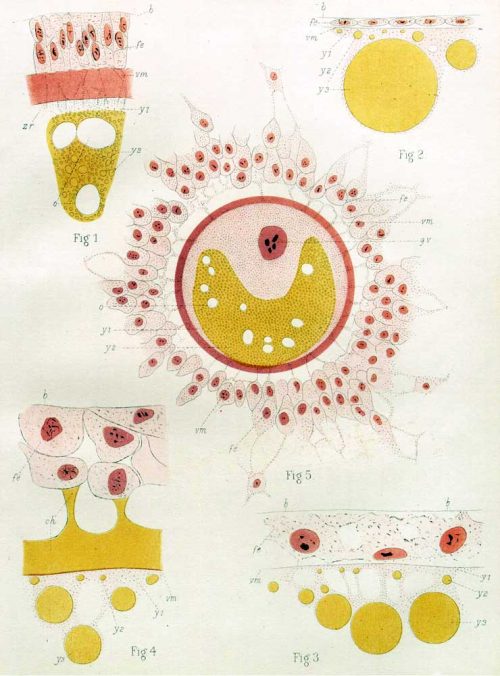
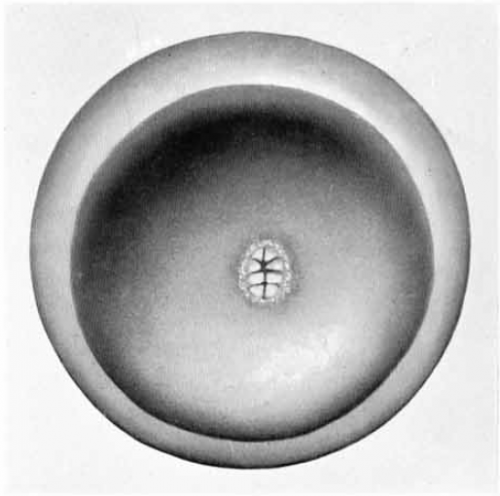
The two monographs describe observations on a collection of 150 or so monotreme eggs and embryos initiated in 1896 by Hill in Queensland, and later added to by Flynn in Tasmania. The samples were preserved in the field and then sectioned, stained and analysed in the lab, often decades later (one was embedded in wax thirty years before being sectioned with a myotome; there may have been half a century between a sample’s collection and the publication of its description). Volume IV describes the making of the monotreme oocyte, its fertilisation and the early cleavage stages of the embryo. Volume VI goes from cleavage until germ layer formation.
The monographs are methodical and descriptions of the collection – each sample is described in the most minute of details, consistent with Hill’s renown for accuracy both in sample preparation and description (“everything he does is done with the most meticulous accuracy…the exact wording of each sentence had to be discussed to make certain that it was precise, unambiguous, and made all possible reservations”, according to the introduction to a dedicated volume of the Journal of Anatomy). It can make for some daunting reading, particularly if you are unfamiliar with some of descriptive terms – one section on the oocyte features eosinophil and basal granules, pseudo-reticular strands, vitello-fatty zones, irregular sinuous folds, linin threads. But in fact much of it is written beautifully and the plates are often quite stunning.

So how do you make a monotreme? Flynn and Hill observed that many features of monotreme oocyte formation and embryogenesis look quite like their bird and reptile cousins. For instance the eggs are yolky, the germinal disc sits on top of the yolk, and early cleavages are meroblastic; early cleavage creates a cellular blastoderm that is separate from the uncleaved yolk; there is an extra-embryonic structure called a germ-ring not seen in mammalian embryos; the multi-layered blastoderm thins out into a single layered structure, and this process appears to involve migration of cells to within the embryo, which looks like reptile delamination. There are also some features that are more mammal-like, and some which seemed to Flynn and Hill to be unique to monotremes, but the key observations seem to be that these mammals share a lot of features of early embryogenesis with birds and reptiles.
As discussed by Guojun Sheng below, Flynn and Hill’s findings suggest we should do more to understand the early development of birds and reptiles, stages which have historically, and for technical reasons, been underserved. We could then move to a more complete understanding of the variations of amniote development, and hence its evolution.
The work also makes us consider the use of different models in developmental biology. There are many animals for which descriptive embryology and anatomy of the kind practised by Flynn and Hill was the only way in, and for which today perhaps only genome sequences – the platypus genome was sequenced in 2007 – can add to our understanding. When such animals are the only extant outcomes of critical evolutionary junctures, papers like Flynn and Hill’s are crucial. And, of course, we are benefiting from work that was of its time: such a collection with its collateral damage of thousands of dead monotremes simply could not have been assembled today. No one has performed a similar analysis since, so the papers really do stand alone in embryology.
But the story isn’t over, at least for echidnas: the establishment of colonies promises to provide additional opportunities to understand monotreme development, and there are even PhD positions available! For now at least, I’d like to propose a monotreme wing in William Sullivan’s Institute for the Study of Non-Model Organisms.
Thoughts from the field
Guojun Sheng, IRCMS, Kumamoto University, Japan
These two related, heavy-weight (literally) papers by the same authors (TT Flynn and JP Hill) [for biographical accounts, see http://adb.anu.edu.au/biography/hill-james-peter-6669 and http://adb.anu.edu.au/biography/flynn-theodore-thomson-6202 ] describe pre-gastrulation development of monotreme embryos. Despite the length (each over 150 pages), these two papers are easy and enjoyable to read. The clarity and quality in their writing style and data presentation can only be appreciated by reading the original papers, rather than from summaries in a handful of review papers which cited them.
What makes them essential readings for developmental biologists is their relevance to modern-day stem cell biology, the foundation of which is based on our understanding of mammalian early development. The monotremes are a group of prototherians (early-branched out mammals) which retain many developmental features of their reptilian ancestor. Our knowledge on cellular and molecular regulation of early lineage segregation (trophoblast, epiblast and hypoblast) in eutherians (placental mammals including mice and humans) would be incomplete, to say the least, without an understanding of these events from the perspective of comparative mammalian/amniote embryology. Although experimentation with monotreme embryos is impractical, similarities between monotreme and reptilian (including avian) early development make a strong case for redoubling our efforts in the investigation of pre-gastrulation development using avian/reptilian models.
Readers may be interested in two papers from our lab:
1) Research paper on chicken pre-ovipositional development (published in Development)
http://dev.biologists.org/content/142/7/1279.long
2) Review paper on comparative amniote pre-gastrulation development (published in Developmental Biology)
www.sciencedirect.com/science/article/pii/S0012160614005193
Both papers have been made free to view for three months courtesy of Wiley Publishers.
Aidan Maartens
This post is part of a series on forgotten classics of developmental biology. You can read the introduction to the series here and read other posts in this series here. We also would love to hear suggestions for future Forgotten Classics – let us know in the comments box.









 (8 votes)
(8 votes) (No Ratings Yet)
(No Ratings Yet) (1 votes)
(1 votes) On p.
On p.  On p.
On p. 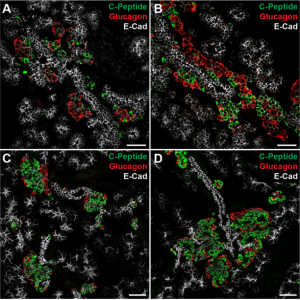 On p.
On p.  On p.
On p. 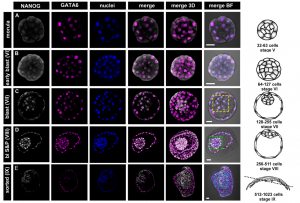 Unlike the mouse embryo, they detect co-expression of NANOG and GATA6 in late blastocysts, suggesting that mutual co-repression does not function in the initiation of lineage specification in rabbit. Furthermore, they show that inhibition of FGF signalling is not sufficient to expand the population of EPI cells, and although the population of PE cells is reduced, GATA6 expression is unaffected. These results indicate that although the key transcription factors are conserved in early mammalian embryogenesis, the way they function, and the way they interact with signalling pathways, may differ between species.
Unlike the mouse embryo, they detect co-expression of NANOG and GATA6 in late blastocysts, suggesting that mutual co-repression does not function in the initiation of lineage specification in rabbit. Furthermore, they show that inhibition of FGF signalling is not sufficient to expand the population of EPI cells, and although the population of PE cells is reduced, GATA6 expression is unaffected. These results indicate that although the key transcription factors are conserved in early mammalian embryogenesis, the way they function, and the way they interact with signalling pathways, may differ between species.