Forgotten classics- Regulating the size of the mouse embryo
Posted by the Node, on 6 April 2016
Snow, M. H. L., Tam, P. P. L. (1979) Is compensatory growth a complicating factor in mouse teratology? Nature 279, 555-557
Lewis, N. E., Rossant, J. (1982) Mechanism of size regulation in mouse embryo aggregates J. Embryol. exp. Morph 72, 169-181
Recommended by James Briscoe (Francis Crick Institute)
As our previous forgotten classic demonstrated, much can be learnt from careful observation of embryonic development. Once there is a good basis of what ‘normal’ is though, the next step is to perturb the system. These days we are used to precise genetic manipulations: removing a single gene (or even a single protein domain), affecting only a specific cell type or at a specific stage in development. However, there are certain questions that benefit from more global alterations, such as making the embryo very small or very big. The two papers recommended by James Briscoe use such approaches to address the important question of how the mouse embryo regulates its size.
The first study, published in Nature in 1979 by Snow and Tam, did not set out to examine size regulation. Mitomycin C was a known teratogen, i.e. a chemical known to cause embryonic malformations, but its effects on early mouse development were unknown. The authors found that the effects of this compound were indeed very quick: by E7 the compound had caused extensive cell death and the embryos were very small. Yet, while this treatment reduced embryonic viability and the resulting pups showed various defects, they were not noticeably smaller. As the authors stated ‘the mouse embryo can withstand a major disturbance in its early development and recover to such an extent that structural abnormalities rarely emerge’. In short, ‘the mouse embryo can be reduced to around 10% of its normal size at a time when it is about to begin organogenesis, but is nearly normal again before that phase of development is complete.’ What an extraordinary ability to regulate size, and a good example that in science you may find an interesting answer for a question that you weren’t even asking!
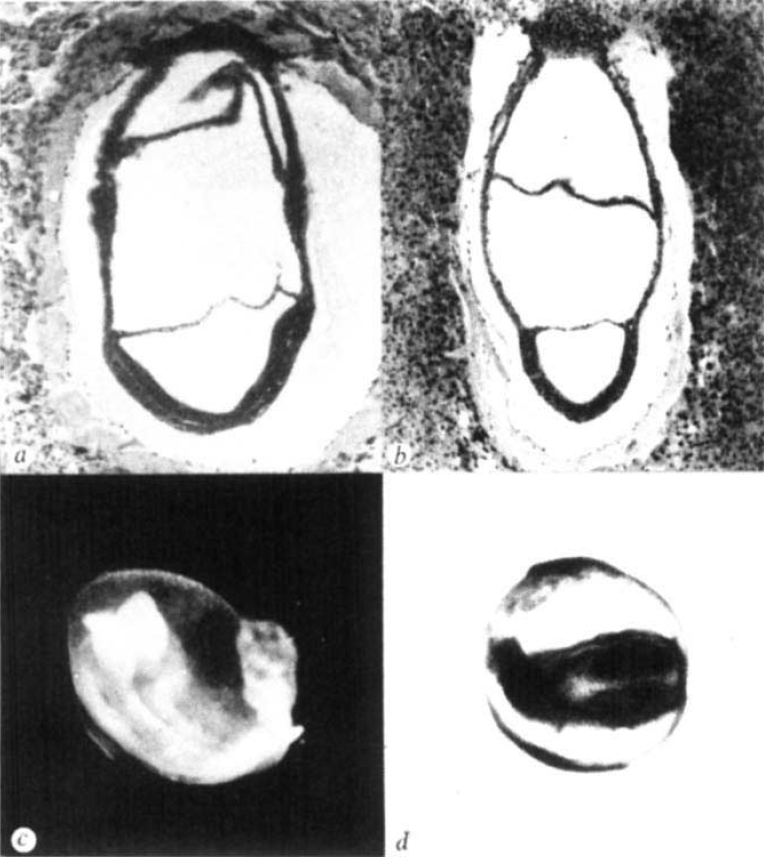
Reprinted by permission from Macmillan Publishers Ltd: Nature (279, 555-557), copyright (1979)
While Snow and Tam inadvertently discovered what happens when a mouse embryo becomes too small, Lewis and Rossant very purposefully tried the opposite in their paper in JEEM (now Development). By 1982, several groups had already shown that if you increase the size of a mouse embryo, e.g. by aggregating several embryos together, the end product is still a viable offspring of normal size. But at which stage of development does this regulation occur, and by what mechanism? To answer this question, the authors compared normal- and double-sized mouse embryos. They showed that while the embryos start off with different sizes, by E6.5-E7 (around the same time as observed by Snow and Tam) they have the same dimensions. They also suggested a mechanism by which this regulation of size may be happening. At E5, normal embryos undergo a period of high mitotic activity while double-sized embryos slow down their cell cycles, giving the normal embryos a chance to ‘catch up’.
As James Briscoe tells us ‘both papers demonstrate the surprising ability of early mouse embryo to regulate its size. Tam and Snow show that ~80% of the cells in an E7 embryo can be killed yet the embryo still recovers (albeit with reduced viability). Conversely Lewis and Rossant show that double sized embryos are twice the size up until ~E6 when they regulate their size back towards normal. Both papers suggest that size regulation happens at around the time of gastrulation, indeed they hint that gastrulation might depend on passing a size checkpoint.’ Despite their important findings, these papers have been cited relatively little, maybe because, as James says ‘the mechanism of size regulation continues to be a mystery today, although it is receiving increasing attention. It is a fascinating example of the self-organising ability of embryogenesis.’
From the authors:
The Lewis and Rossant paper is one of my personal favourites and I still quote it, even if it is often ignored by the field! In fact it is a still unsolved question of how the embryo manages to size regulate so well in such a relatively short period of time. I have been trying to interest new post-docs in the lab with returning to the problem and investigating whether Hippo or other growth control pathways might be involved and/or whether there is a metabolic link to mTor pathways. But so far noone has taken up the challenge!
Janet Rossant, The Hospital for Sick Children (Canada)
The study on the effect of inhibition of DNA synthesis by mitomycin C on embryogenesis was founded on an observation made by Michael Snow: that to account for the increase in cell number during mouse gastrulation, the whole epiblast cell population has to undergo a doubling in about every 6-7 hours and, in particular, a small sub-population of cells would have to proliferate at a much faster rate with a cell cycle time of about 3 hours. It was not known previously that cell cycling could be that fast in mouse embryos. We hypothesized that this highly proliferative population may be hit more severely by blocking DNA synthesis, and the subsequent deficiency of any cell types in the embryo might reveal the developmental fate of the descendants of this population. As it turned out, the unexpected outcome of the experiment pointed instead to a mechanism for size regulation by compensatory growth. We followed up this study years later where under-sized mouse gastrula were created, not by cell killing, but by removing one to two blastomeres from the 4-cell stage pre-implantation embryo (Power and Tam, 1993). Results of this study further highlighted the attribute of size (or cell number) up-regulation in the control of post-implantation development. The under-sized embryo does not initiate gastrulation until it has attained a threshold cell number and this is accomplished by extending the phase of rapid cell proliferation beyond the normal duration of 24 hours into gastrulation. Recently, there has been heightened interest in the mechanism of size regulation and the control of the timing of morphogenetic events during development and the connection of this process to the mechanosensory activity and the intercellular communication among cells in a community.
Reference: Power, M. A., Tam P. P. L. (1993) Onset of gastrulation, morphogenesis and somitogenesis in mouse embryos displaying compensatory growth. Anatomy and Embryology 187,493-504
Patrick Tam, University of Sydney (Australia)
Further thoughts from the field
These are two really fantastic papers that are indeed often overlooked, and well worth revisiting. Both studies use classic mouse embryology (and teratology – now that’s a word we don’t see too often these days) methods to describe the remarkable regulative capacity of the mammalian embryo. In the intervening (almost 4!?) decades, we’ve come full circle. Armed with sophisticated genetics tools, an increasing ability to accurately measure (and perturb) the behavior of individual cells within a population, and spurned by studies yielding information on (possibly conserved) mechanisms regulating tissue size and scaling in other systems, for example Drosophila, there’s been an upsurge of interest in this phenomenon in the mouse. Interestingly, two recent studies from Miguel Torres’ lab at the CNIC Institute in Madrid, and Tristan Rodriguez’s lab at Imperial College in London, revisited the question in a contemporary setting and pin-pointed the same window of time (approximating to the onset of gastrulation) as noted in the earlier Tam and Rossant studies, when everything seems to have happened, and any outlier embryos have either managed to catch-up or slow-down (Claveria et al., Nature 2013; Sancho et al., Dev Cell 2013). Moving forward, it’ll be interesting to see how our understanding of this phenomenon deepens, especially bearing in mind that achieving the exquisitely invariant size of mammalian embryos (and organs) likely involves the sensing, integration, and reaction to, several inputs.
Kat Hadjantonakis, Sloan Kettering Institute (USA)
Nature has kindly provided free access to the Snow and Tam paper until June 2016; the Lewis and Rossant paper is freely available.
—————————————–
by Cat Vicente
 This post is part of a series on forgotten classics of developmental biology. You can read the introduction to the series here and read other posts in this series here.
This post is part of a series on forgotten classics of developmental biology. You can read the introduction to the series here and read other posts in this series here.


 (5 votes)
(5 votes) This year you will be awarded the first Cheryll Tickle Medal by the BSDB. What does it mean to you to receive this prize?
This year you will be awarded the first Cheryll Tickle Medal by the BSDB. What does it mean to you to receive this prize? How did you first become interested in developmental biology and why did you decide to study limb development?
How did you first become interested in developmental biology and why did you decide to study limb development?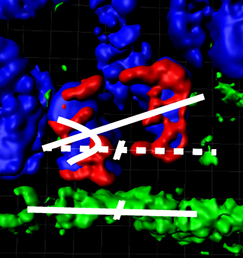 Mutations affecting tubulin genes have been implicated in a range of human neurological disorders but very little is known about the cellular mechanisms that underlie these disorders. Now, on p.
Mutations affecting tubulin genes have been implicated in a range of human neurological disorders but very little is known about the cellular mechanisms that underlie these disorders. Now, on p. 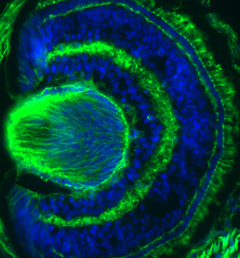
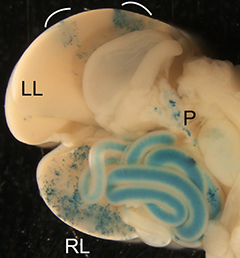 The liver is a vital organ, and understanding how it develops can provide key insights into liver disorders and regeneration. Here, Mary Weiss, Margaret Buckingham and colleagues use a retrospective clonal approach to provide an in-depth analysis of cell behaviour during liver development in mice (p.
The liver is a vital organ, and understanding how it develops can provide key insights into liver disorders and regeneration. Here, Mary Weiss, Margaret Buckingham and colleagues use a retrospective clonal approach to provide an in-depth analysis of cell behaviour during liver development in mice (p. 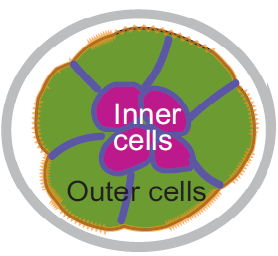 Chazaud and Yamanaka discuss recent advances in live imaging, computational modelling and single cell analyses that provide insights into how the first three cell lineages of the mouse embryo are generated.
Chazaud and Yamanaka discuss recent advances in live imaging, computational modelling and single cell analyses that provide insights into how the first three cell lineages of the mouse embryo are generated.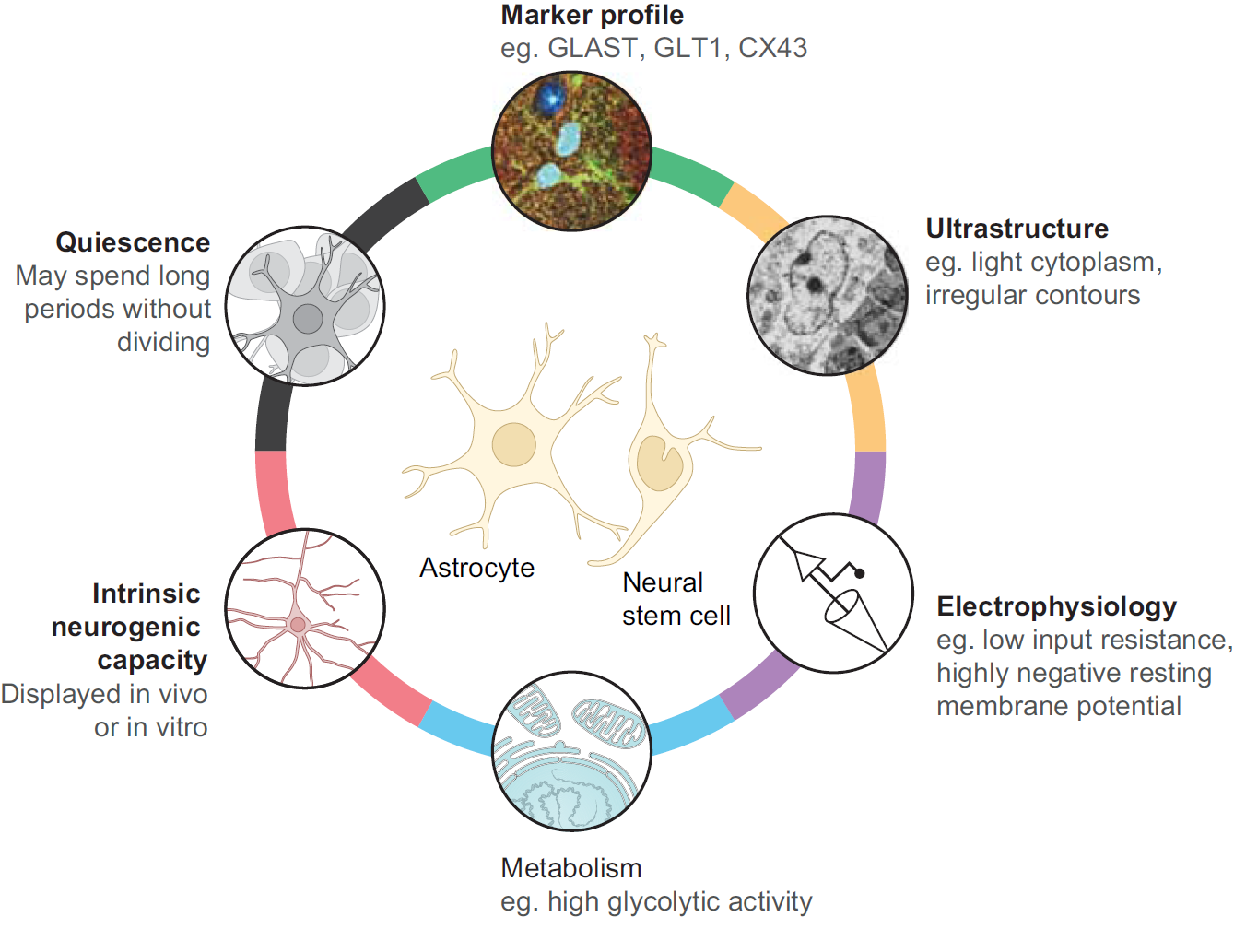 Unlocking the neurogenic potential of astrocytes in different brain regions
Unlocking the neurogenic potential of astrocytes in different brain regions
 (No Ratings Yet)
(No Ratings Yet) Following Development’s call for papers for their upcoming
Following Development’s call for papers for their upcoming  – Liangyu discussed how he adapted the auxin-inducible degradation system to C. elegans, providing a new tool for
– Liangyu discussed how he adapted the auxin-inducible degradation system to C. elegans, providing a new tool for  Also on the Node
Also on the Node