Alejandro Sánchez Alvarado is an investigator at the Stowers Institute and Howard Hughes Medical Institute, and has worked for many years on regeneration in a little flatworm- the planaria. The Node interviewed Alejandro at the 2013 International Society of Developmental Biology meeting, and asked him about his career, his work on regeneration, and his role as co-director of the Woods Hole embryology course.
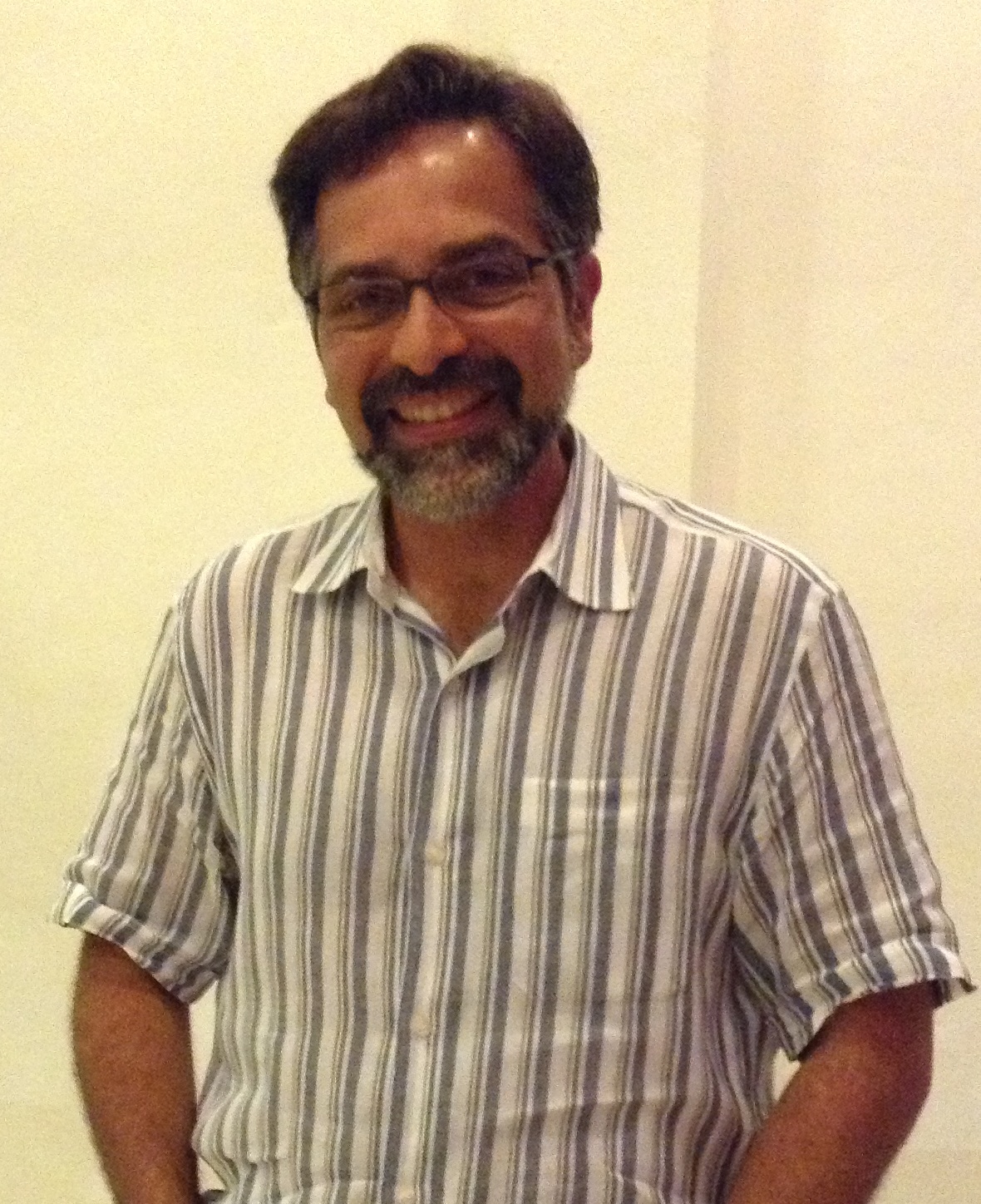 When did you first become interested in Biology?
When did you first become interested in Biology?
My interest in Biology arose as the result of a phenomenal high school teacher named Maldonado, who had a very unique method of teaching. We walked into his classroom on the first day of Biology classes and he asked us: ‘What is the minimum number of characters that you need to make a language?’. My friend Nestor said one letter: ‘a’ could be one word, ‘aa’ could be two words, etc. And Maldonado said ‘well, that is how life works. Only it uses 4 letters, ACGT’. And he then started explaining DNA and other such concepts. He was very good at challenging us. Once, after he described how DNA is a double helix, he asked us if we could think of a way it could copy itself. My friend Francisco was very close, and suggested a conservative rule. Following this Maldonado explained Meselson and Stahl’s experiment [showing that DNA replication is semi-conservative], and this is when I got really hooked- I understood that you could arrive at all these fundamental conclusions without even seeing the molecules. I wanted to know more about this awesome molecular biology. He had a fundamental effect on how I did things down the road, because I was on my way to medical school, the only way to do Biology in Latin America at the time, and he opened my eyes to the possibility of molecular biology.
You are originally from Venezuela, but you moved to the US to study molecular Biology. Why did you decide to move abroad?
In Venezuela, the only way to study biological sciences was to go to medical school. But Maldonado inspired me to study molecular biology. I went to embassies in Caracas to check university catalogues- I considered going to France, England and even Russia, but I realized that there were many more opportunities in the United States, and I looked for universities where I could get a degree in molecular biology. I spoke no English at the time, and of the 4 universities that offered a molecular biology course I ended up choosing Vanderbilt, in Nashville Tennessee, primarily because nobody spoke Spanish there and I wanted to learn English. It was a little crazy, but when you are young everything looks like an adventure and I really wanted to study molecular biology.
You didn’t do a PhD straight after your undergraduate studies, and instead you decided to take some time off and travel around South America. Why did you do this?
I went back to Venezuela assuming that I would be able to practice as a molecular biologist at a research institution, but the economic situation at the time did not allow it. So I decided to do something that I had always wanted to: travel around South America. The trip was supposed to take 3 months, but it took almost 8 months to complete! I ran out of money in Brazil, then I got robbed, loosing everything I might be able to sell to make ends meet. From that point onwards I was at the mercy of people who helped me get all the way back home, and it was a long trajectory. By the time I arrived back in Caracas again, everyone thought I was gone, because I had not been in touch for months. My dad could hardly recognize me as I had lost so much weight. It was a terrific adventure! And it did teach me something: things are never as bad as they look.
Because of this adventure I couldn’t go to graduate school at Washington University in Saint Louis as I had planned. Instead I went to Cincinnati to be a technician and try to go to graduate school. There I met some terrific people, such as Jeff Robinson, Tom Doetschman and Arnold Schwartz who took me under their wing and gave me opportunities, so everything just worked out. But that hiatus… I would do it again for sure. The trip convinced me that what I really wanted to do was science- the best thing I could do for my fellow human beings was to give back by trying to expand knowledge. That trip also reinforced my desire to do basic research. Translational research is fine, but I don’t want to map already discovered continents- I want to find new ones.
You work on regeneration. Why is regeneration your continent to be discovered?
I came in to the field of developmental biology by accident. When I was a graduate student I joined the lab of Jeff Robbins to work with embryonic stem (ES) cells. I became absolutely hooked by the notion that these cells were capable of giving rise to everything in the mouse. I wanted to understand how genomic output and plasticity were regulated, and hence the early decision steps that those cells took. However, these processes were hard to study in vivo in mice, so I decided that I was going to work on animals that grow outside of the mother.
Later I was offered a staff position by Don Brown at the Carnegie Institution. I was not prepared to run my own lab- I barely knew any developmental biology! But I thought that if I went to an embryology department and was surrounded by developmental biologists I would learn. When trying to decided what to do for my NRSA (research fellowship) there was an article from an Indian researcher (Mohanty-Hejmadi) describing an unusual species of frog that could undergo a phenomenal homeotic transformation when exposed to a particular chemical: the tails would regenerate as limbs. Malcolm Maden showed the same thing in Rana temporaria, a species I had access to. I thought this was an amazing process. So I wrote my NRSA proposal on this topic, and began to compare normal tadpole tail regeneration with this homeotic transformation to find out what genes were being up or down regulated.
The whole idea was plasticity. When I began to study regeneration I realized that most people thought that regeneration was an epiphenomenon and therefore not worth studying- if you study embryogenesis, you understand regeneration. But I was never quite convinced of that. In regeneration, you have a situation where cells already know what they are: they are blood vessels, blood, connective tissue. You amputate them, and that amputation somehow resets the state of the tissue so that new tissues are made, and are functionally integrated into the pre-existing tissue. This does not happen in embryogenesis. I was completely hooked.
You mentioned how you started working with frogs, but you eventually moved on to the planarian. Why?
After I finished my screen, I had a beautiful collection of up and down regulated genes. The only way I could address the function of these molecules was to perturb them, but there was no way to do this in frogs at the time. I had to make a really hard choice: I could develop gene perturbation methodologies for the tadpole, but would have to be in this crazy species, Rana temporaria, that can only mate once a year; or I could take a bigger step back and find a new system to study regeneration that I could try to manipulate molecularly.
I took the Woods Hole Embryology Course in 1995 because I wanted to expand my understanding of the model systems available. I went back in 1996 to do some independent research, cutting everything that came out from the sea to see what regenerated. I had previously thought that regeneration was a rare event, but I realized that almost everything that I cut could regenerate! Then, also at Woods Hole, I found a book from 1901 called ‘Regeneration’ written by Thomas Hunt Morgan, who had actually worked on planarians (11-12 publications) before he moved a lot of his efforts to Drosophila. I read the book from cover to cover. I couldn’t believe it- why wasn’t anybody studying this phenomenal problem? This was the chance encounter that lead me to think that planarians might be a good system to study regeneration.
I heard the rumour that the planarians that established your current colony were collected from a fountain in Barcelona…
This was another of these chance encounters. I went to a British Society of Developmental Biology meeting in York and I concluded my presentation by saying ‘I am done with the frogs, I want to start working on planarians’. Someone in the audience mentioned that they had a postdoc in his lab who worked on planarians, and wanted to return to the US. That is how I met Phil [Newmark]. I invited him to come to my lab, and he brought with him a seed population of the Mediterranean species that we work with. We were just getting RNAi to work when all the planarians died: because of water conditions and so on- the usual things that happen when you are trying to establish a system.
Phil was ready to give up, but I said ‘Phil, we are going to Spain. It is rainy season, and you know these planarians come from some abandoned fountain. We’ll fly on Thursday, on Friday we’ll collect these animals and we’ll be back Monday or Tuesday.’ And that is what we did. We hopped on a plane with a green cooler that I still have, full with all kinds of bottles and traps, enough to seed every fountain in this specific park in Barcelona: we knew the planarians could be obtained from one of the fountains, but we didn’t know which. When we landed in Barcelona the first thing that I did was to check if it had rained: we knew that the fountain was broken, and that the worms would only come out when it was filled with rain water. Luckily it had rained. From the airport, we asked the cab driver to take us to a butcher, where we bought some liver to use as bait. Planarians love liver, but chopping it up in our hotel room was a bloody business! The word must have gone around because people in the hotel and the taxi drivers looked at us in a funny way! We seeded all the fountains in the park, and eventually, on the last day, we found a fountain teeming with planarians! We brought them back on the ice cooler. So yes, our entire planarian colony, and the planarians that most American and international labs are using, came from that fountain.
Where do you think the field of regeneration is heading? What are the big questions that excite you?
To me regeneration today its what embryology was at the beginning of the twentieth century, when people looked at all these very different embryos and discovered their shared properties. First principles are being uncovered, some similarities and dissimilarities are arising, and there is a great deal of things that don’t fit together. What I am excited about is the distinct possibility that we will be able to begin asking questions across multiple phyla, and get to the bottom of whether or not regeneration is something that was invented multiple times and independently in evolution or whether it is a fundamental attribute of multicellular life. We are beginning to really gain the tools to ask these questions, and perturb the system in ways that will allow us to test hypotheses. I am really excited by that.
I am also excited about how studying regeneration might allow us to understand some aspects of embryogenesis, such as scale and proportion. If a salamander regenerated its hand to the size of a embryo hand, it would be a useless hand. Instead, the hand grows to the perfect size. How scale and proportion are maintained is something that we don’t understand, but it’s fundamental. There are a whole bunch of other intriguing questions, like how do these newly formed tissues functionally integrate with pre-existing tissues, and how is it decided which cells are killed and which ones survive – both in regeneration and in tissue homeostasis.
I think that regeneration can provide the context to understand biological attributes that may be responsible for malfunctions, pathologies in humans or other organisms. It really is a new continent, one of the last unexplored frontiers of developmental biology.
You have previously expressed your interest in the history of science. Is this interest in pre-molecular literature strictly historical or do you think it is beneficial for research to explore ‘old’ literature?
I think it is both. I like to understand the humans behind the science but I also like to understand the context in order to understand the genesis of ideas. For example: who had the first idea of what a stem cell is? How far in biological thinking does it go? That is a big draw for me, and usually this happened in the pre-molecular era. I like to go back and read as far as I can, trying to identify who may have formulated those ideas. And I think that is a great exercise intellectually, because it allows you to interrogate biases that we immediately accept. For example, the notion that stem cells were just naïve cells waiting to be induced. We have a bias introduced by this concept, whereas now we know stem cells are a highly specialized cell type that is fighting constantly not to differentiate.
You have been the co-director of the Woods Hole Embryology Course for the last two years. Do you have a vision for where you want the course to go? How do you see your role as the director?
I think both Richard [Beringher] and I took on this position because we really believe that despite what everybody is trying to tell us, developmental biology and embryology are not dying arts. The magic of the course is to let people who have never seen a sea urchin or an ascidian embryo before watch them unfold, from some nondescript cell into organisms, in all their glory. There is a great deal of inspiration in that. There are thousands of different species out there, doing truly remarkable things that we are not even aware of. We try to plant the seed, in the few brains we get every year, that there is merit in looking beyond what is readily accessible.
The particular vision that I have for the course is to try to bring molecular tools to organisms that have been under-studied. We bring in organisms and say “let’s make an RNA seq library from this embryo, and sequence it. Let’s get a transcriptome for this species this year, the next species next year. Let’s see what we can learn from these diverse organisms.” I think there is some merit in that because it will allow us to expand our understanding of the early developmental stages.
But I think the main vision for the course is to try to convey to the students the tremendous value of thinking about discovery. Most of the students are hearing what I hear from my colleagues- how difficult it is to get grants if our research is not translational. I know that these are the exigencies of the time, but that is not the only way things should happen. I want to be able to convey to the students that other people can mind the shop, but that they should be doing discovery research. There are many more labs studying C.elegans than there are cells in these worms! Why? Because there are remarkable amounts of Biology there to be discovered. I want to be able to encourage the students to think a little bit about this.
We hope that there is a palpable difference between how the students think about their biological problems before they come to the course and when they leave. I don’t want them to specialize too much- specialization is for insects, not for humans! We should have an expertise, but that should not dictate what we can do.
This is quite an important year for the Woods Hole Embryology course- it is its 120th anniversary. Are you doing anything to celebrate?
We are. The MBL is celebrating 125 years, and the course has been going for 120 years. There’s a mini-symposium on July 12th, where prior directors, faculty and students are giving presentations about their current science and how Woods Hole affected their way of thinking. I want the students to see the origins of the ideas of these incredibly smart, established people, who have contributed enormously to the field.
It will be an opportunity for the students to see how connected the community really is. The developmental biology community is a tight-knit community, and I think this is one of the reasons why it has been able to weather all sorts of challenges. I think we are about to witness a renaissance on the importance of this field, as our ability to interrogate more and more species, and more and more diversity becomes more accessible. It is just a matter of time. I am hoping that this embryology course will be continuing for another 120 years.
 (26 votes)
(26 votes)
 Loading...
Loading...



 (2 votes)
(2 votes)
 I think you do challenge people, partially because you present many different views, not just your own.
I think you do challenge people, partially because you present many different views, not just your own.


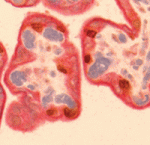 Correct trophoblast development is essential for placenta function and embryogenesis. In humans, the early stages of this process can only be modelled in vitro using embryonic stem cells (ESCs); however, owing to the failure to identify the stepwise progression that occurs during differentiation, the identity of the resulting cells is not clear. Now, on
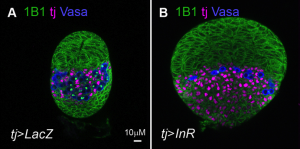
Correct trophoblast development is essential for placenta function and embryogenesis. In humans, the early stages of this process can only be modelled in vitro using embryonic stem cells (ESCs); however, owing to the failure to identify the stepwise progression that occurs during differentiation, the identity of the resulting cells is not clear. Now, on 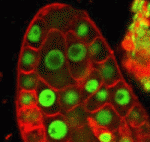 Unlike animals, plants do not set aside germ cells during embryogenesis. Instead, the precursors of these cells, called spore mother cells (SMCs), are generated via a somatic-to-reproductive transition that occurs later in life. Although epigenetic remodelling has been largely studied in the post-meiotic phase of germline development, it is unknown whether pre-meiotic events contribute to cellular reprogramming in the reproductive lineage. Now, on
Unlike animals, plants do not set aside germ cells during embryogenesis. Instead, the precursors of these cells, called spore mother cells (SMCs), are generated via a somatic-to-reproductive transition that occurs later in life. Although epigenetic remodelling has been largely studied in the post-meiotic phase of germline development, it is unknown whether pre-meiotic events contribute to cellular reprogramming in the reproductive lineage. Now, on 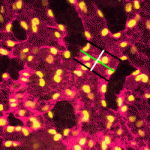 Mechanical forces such as blood flow play a key role in regulating vascular remodelling and angiogenesis. Vessel diameter must be tightly controlled to establish correct hierarchical vascular architecture, but how this is achieved is unclear. Now, on
Mechanical forces such as blood flow play a key role in regulating vascular remodelling and angiogenesis. Vessel diameter must be tightly controlled to establish correct hierarchical vascular architecture, but how this is achieved is unclear. Now, on 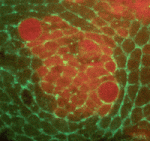 On
On 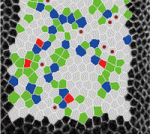 The role of mechanical stress in regulating cell shape and tissue morphogenesis is further examined by Shuji Ishihara and Kaoru Sugimura (
The role of mechanical stress in regulating cell shape and tissue morphogenesis is further examined by Shuji Ishihara and Kaoru Sugimura (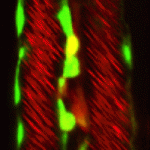 The emergence of haematopoietic stem cells (HSCs) from the aorta-gonad-mesenephros (AGM) region requires the coordinated activity of multiple transcriptional networks. The transcription factor stem cell leukemia (SCL) is necessary for this process; however, multiple scl isoforms exist and the exact stage at which each is required is unknown. In this issue (
The emergence of haematopoietic stem cells (HSCs) from the aorta-gonad-mesenephros (AGM) region requires the coordinated activity of multiple transcriptional networks. The transcription factor stem cell leukemia (SCL) is necessary for this process; however, multiple scl isoforms exist and the exact stage at which each is required is unknown. In this issue ( The formation and maintenance of adipose tissue is essential to many biological processes and when perturbed leads to significant diseases. Here, Jon Graff and colleagues highlight recent efforts to unveil adipose developmental cues, adipose stem cell biology and the regulators of adipose tissue homeostasis and dynamism. See the
The formation and maintenance of adipose tissue is essential to many biological processes and when perturbed leads to significant diseases. Here, Jon Graff and colleagues highlight recent efforts to unveil adipose developmental cues, adipose stem cell biology and the regulators of adipose tissue homeostasis and dynamism. See the  Hox genes encode a family of transcriptional regulators that elicit distinct developmental programmes along the head-to-tail axis of animals. Here, Mallo and Alonso examine the spectrum of molecular mechanisms that control Hox gene expression in model vertebrates and invertebrates. See the
Hox genes encode a family of transcriptional regulators that elicit distinct developmental programmes along the head-to-tail axis of animals. Here, Mallo and Alonso examine the spectrum of molecular mechanisms that control Hox gene expression in model vertebrates and invertebrates. See the 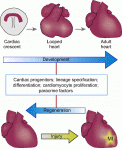 The recent EMBO/EMBL-sponsored symposium ‘Cardiac Biology: From Development to Regeneration’ gathered cardiovascular scientists from across the globe to discuss the latest advances in our understanding of the development and growth of the heart, and application of these advances to improving the limited innate regenerative capacity of the mammalian heart. Christoffels and Pu summarize some of the exciting results and themes that emerged from the meeting. See the
The recent EMBO/EMBL-sponsored symposium ‘Cardiac Biology: From Development to Regeneration’ gathered cardiovascular scientists from across the globe to discuss the latest advances in our understanding of the development and growth of the heart, and application of these advances to improving the limited innate regenerative capacity of the mammalian heart. Christoffels and Pu summarize some of the exciting results and themes that emerged from the meeting. See the  (No Ratings Yet)
(No Ratings Yet)

 (26 votes)
(26 votes)