Friday Harbor Comparative Embryology Course – Applications open until Feb. 26
Posted by Mickey von Dassow, on 10 February 2015
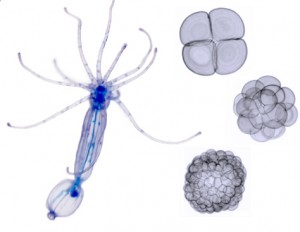
 Taking the Comparative Invertebrate Embryology course at the Friday Harbor Labs was one of the pivotal experiences of my graduate life, and it was possibly the most valuable, and enjoyable, course I’ve ever taken. I was a student in the course several years ago, when it was taught by two great scientists, Dr. Charles Lambert and Dr. Mark Martindale (it is taught by different instructors every year). I loved seeing the amazing diversity of marine invertebrate embryos and larvae, and watching their development in real time, after collecting the animals from their natural habitats. One of the things I especially appreciated about the course is that it gave me an opportunity to watch and observe organisms closely, on their own terms, rather than immediately trying to convert them into data points to answer a specific question. This let me see aspects of the embryos and larvae that I would not have noticed otherwise.
Taking the Comparative Invertebrate Embryology course at the Friday Harbor Labs was one of the pivotal experiences of my graduate life, and it was possibly the most valuable, and enjoyable, course I’ve ever taken. I was a student in the course several years ago, when it was taught by two great scientists, Dr. Charles Lambert and Dr. Mark Martindale (it is taught by different instructors every year). I loved seeing the amazing diversity of marine invertebrate embryos and larvae, and watching their development in real time, after collecting the animals from their natural habitats. One of the things I especially appreciated about the course is that it gave me an opportunity to watch and observe organisms closely, on their own terms, rather than immediately trying to convert them into data points to answer a specific question. This let me see aspects of the embryos and larvae that I would not have noticed otherwise.
The Friday Harbor embryology course transformed my research path. At the time I took it, I was a graduate student working on biomechanics and pattern formation in invertebrate colonies, and was more interested in physiology and marine biology than development. I took the course because I felt a need to get a complete picture of invertebrate biology, and it sounded like fun (it was). I loved the complexity of the larvae, with their muscles, ciliated bands, guts, skeletons, and behavior; but as I watched the embryos transform themselves from egg, to blastula, to gastrula, and beyond, I learned to love them for their own sake. At the same time I gained a feel for their variety and for their natural environments.
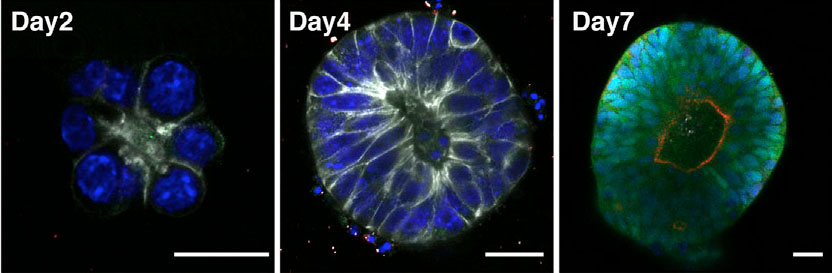
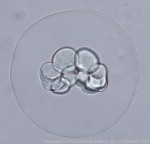 So, after finishing my Ph.D., I switched from working on adult animals to working on embryos, focusing on the biomechanics of gastrulation in frogs (with Dr. Lance Davidson), and then struck out on my own to study echinoderm blastula formation. The memory of my experiences at Friday Harbor has continued to shape my research, making me think hard about the kinds of variation organisms have to tolerate in nature, variation due to things like salinity, temperature, and maternal condition. And it made me keenly aware of the diversity of developmental processes within and among animal phyla. This gave me a broad perspective on the mechanics of morphogenesis, and how it relates to development-environment interactions and developmental evolution.
So, after finishing my Ph.D., I switched from working on adult animals to working on embryos, focusing on the biomechanics of gastrulation in frogs (with Dr. Lance Davidson), and then struck out on my own to study echinoderm blastula formation. The memory of my experiences at Friday Harbor has continued to shape my research, making me think hard about the kinds of variation organisms have to tolerate in nature, variation due to things like salinity, temperature, and maternal condition. And it made me keenly aware of the diversity of developmental processes within and among animal phyla. This gave me a broad perspective on the mechanics of morphogenesis, and how it relates to development-environment interactions and developmental evolution.
 My personal experience in the course led me to my own peculiar niche in developmental biomechanics, but other former students have gone on to study cell and molecular biology, evo-devo, larval ecology, and many other fields. One of the great things about the Friday Harbor course is that it bridges cell and molecular perspectives with ecological and evolutionary perspectives to provide an integrated view of animal development. So the course is very useful for both biologists who wish to understand diversity in developmental modes for ecological or evolutionary studies, and cell and developmental biologists who wish to broaden their knowledge of embryos beyond the standard model systems. The focus is on watching and observing living embryos and larvae from as many different kinds of marine animals as one can get one’s hands on (typically a few species from each of over a dozen phyla).
My personal experience in the course led me to my own peculiar niche in developmental biomechanics, but other former students have gone on to study cell and molecular biology, evo-devo, larval ecology, and many other fields. One of the great things about the Friday Harbor course is that it bridges cell and molecular perspectives with ecological and evolutionary perspectives to provide an integrated view of animal development. So the course is very useful for both biologists who wish to understand diversity in developmental modes for ecological or evolutionary studies, and cell and developmental biologists who wish to broaden their knowledge of embryos beyond the standard model systems. The focus is on watching and observing living embryos and larvae from as many different kinds of marine animals as one can get one’s hands on (typically a few species from each of over a dozen phyla).
I loved being a student so much that I jumped at the chance to co-teach the course this summer with Dr. Yale Passamaneck, an excellent evolutionary developmental biologist who has worked with several invertebrate phyla.
Applications are still open, and we have a few spots left that we hope to fill. Applications are needed before February 26th. Financial aid may be available.
If you are interested in the course, please see the course description at: http://depts.washington.edu/fhl/studentSummer2015.html#SumA-4
The course runs from June 15 – July 17, 2015 (5 weeks).


 (3 votes)
(3 votes)
 (No Ratings Yet)
(No Ratings Yet)
 (6 votes)
(6 votes) – Christoph and his colleagues wrote about their recent Development paper, where they
– Christoph and his colleagues wrote about their recent Development paper, where they 






 This post is part of a series on a day in the life of developmental biology labs working on different model organisms. You can read the introduction to the series
This post is part of a series on a day in the life of developmental biology labs working on different model organisms. You can read the introduction to the series