Welcome to Yokohama… ISSCR 2012
Posted by Jamesomalley, on 13 June 2012
“In a world of total darkness, a glimmer of light emboldens human spirit”. A view on the work of stem cell scientists among the closing remarks from a moving talk by the NBC journalist Charles Sabine, recently diagnosed with Huntington’s disease. It brought into sharp focus one of the many reasons why this field of research holds so much potential for the future of medicine and therapeutics, and also reflected the overall message from previous talks by heavy-hitters in stem cell science; we are on the cusp of discoveries which could make regenerative medicine a reality.
The conference began mid-afternoon, once we had claimed our passes and purple tote bags filled to the brim with leaflets from companies extoling the virtues of their newest products-I was especially drawn to the claim that I could “Have a weekend off! Change media less often!”…ah but if only my supervisor agreed!
In the huge National Conference Centre at the Pacific Yokohama hotel, Fred Gage, the outgoing president of the ISSCR welcomed the 3,150 registered attendees of the conference and outlined the future aims of the organisation: to educate, investigate and influence public policy, and to increase focus on translational science. This meeting, the first to take place in Asia, was an opportunity to allow basic and clinical researchers to collaborate and facilitate the development of safe, science-based applications of discoveries in the field of stem cell research. This call for cooperation was echoed by the president of the Japan Science and Technology Agency, Michiharu Nakamura, who encouraged “global collaboration”…considering these are the guys who support Shinya Yamanaka and have committed ¥35 billion to research in the 18 years, I think it’s a prudent message!
The keynote talks began with Rudolf Jaenisch, recipient of this year’s ISSCR Public Service Award presented by Rob and Cheryl McEwen, Canadian philanthropists and supporters of stem cell research and innovation. He described his lab’s most recent work in the area of reprogramming and their quest to understand if this process is deterministic or stocastic. He showed how their detailed analysis of cells could lead to greater control of the generation of induced pluripotent stem cells (iPSCs) and finished his talk describing how these cellsmay be used to model human disease and provided an insight into an issue in all scientific experiments: how to generate a proper control!
More fundamental in vitro research was presented by Austin Smith, a key figure in establishing our understanding of the mechanism of pluripotantcy. His talk focused on the mechanism of 2i media and in detail how it functions to maintain stem cells in a “ground state” of pluripotantcy. He also discussed it’s use in other contexts and most interestingly, widened the potential perspective on the definition of what might be considered a stem cell. Mechanisms preventing the aquisition of this pluripotent, self-renewing state were described by the legend that is Prof. John Gurdon-he of two-headed Xenopus fame! Describing his investigation of the “resistance to reprogramming”, it was fantastic to discover that older techniques such as nuclear transfer (which he had been crucial in developing) could provide insight into the questions posed by the modern field of Reprogramming. The use of iPSCs for patients was again addressed by Kazutoshi Takahashi, one half of the famous reprogramming duo. He outlined a system by which stem cells and iPSCs could be screened for their suitability for use in therapeutic applications based on highly detailed transcriptional and epigenetic investigation of multiple cell lines.
The importance of epigenetics was again touched upon by Jane Visvader, who discussed molecular regulation of mammary stem cells at the beginning of the second Planary Session. Elaine Fuchs described their investigation of the role of sweat glands and recent discovery of a stem cell population contained within-more than just a cooling system it seems! An eyeball staring back at you in a dish might sound like something from a horror film, but for Dr. Yoshiki Sasai, the spontaneous generation of optic cups from mouse and human ES cells provided an opportunity to investigate the mechanism of development and formation of this important part of the body. His films showing the folding and curving of cell-layers into 3D structures were breathtaking, and highlighted how small observations can lead to bigger stories.
Finally, the Ernest Mcculloch Memorial Lecture was given by Prof. Irving Weissman, instrumental in identifying and enumerating hematopoietic stem cells in mouse models and humans. His talk encompassed the huge number of breakthroughs in the area of haematopoietic stem cell research from it’s early beginnings will little understood lineages to it’s modern-day potential use for cancers such as lymphoma.
While the topic of the day’s talks varied widely, the singular mood was one of forward-facing hope for the future of stem cells and their use in regenerative medicine. Looking up into the neon-tinged night sky, I felt the potential for our research was as high as the towering skyscrapers of Yokohama Bay.
James
#ISSCR2012 #Japan #StemCells #Conference #iPS


 (7 votes)
(7 votes)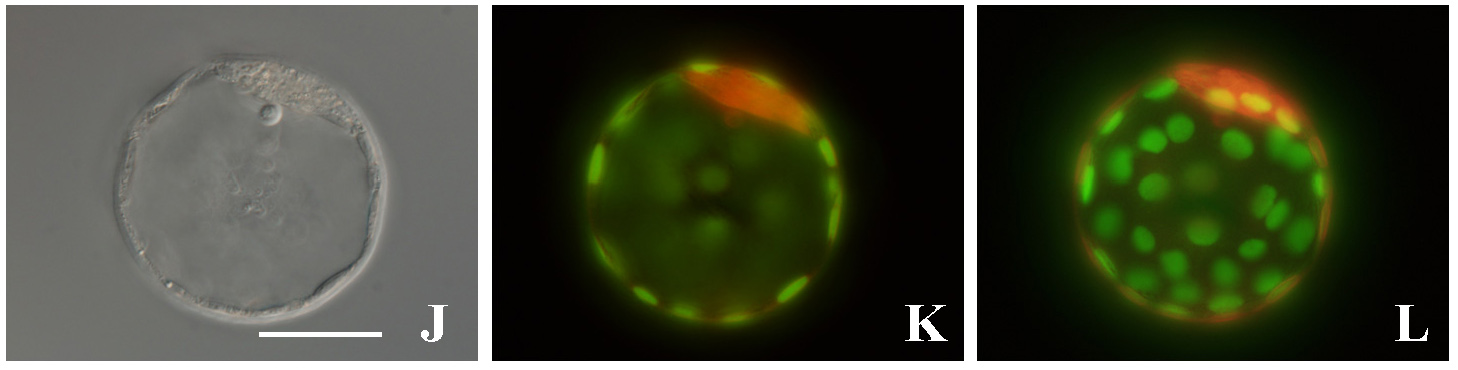
 (No Ratings Yet)
(No Ratings Yet)
 During early development, embryonic cells can form derivatives of all three embryonic layers. This pluripotency, which is regulated by a gene regulatory network that includes the transcription factors Oct4 and Nanog, is lost in mouse embryos between about E7.5 and E8.5. Here (
During early development, embryonic cells can form derivatives of all three embryonic layers. This pluripotency, which is regulated by a gene regulatory network that includes the transcription factors Oct4 and Nanog, is lost in mouse embryos between about E7.5 and E8.5. Here (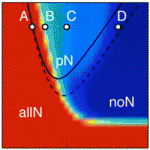 During neurogenesis, lateral inhibition controls the final number of neurons. Neuronal precursors that express high levels of Delta prevent the neuronal differentiation of neighbouring cells by inducing Notch-dependent inhibitory signals in these neighbours. However, neurogenic wavefronts spread through non-neurogenic areas during development, so why isn’t lateral inhibition disrupted where these wavefronts contact non-neurogenic tissue? José María Frade, Saúl Ares and colleagues investigate this puzzle on
During neurogenesis, lateral inhibition controls the final number of neurons. Neuronal precursors that express high levels of Delta prevent the neuronal differentiation of neighbouring cells by inducing Notch-dependent inhibitory signals in these neighbours. However, neurogenic wavefronts spread through non-neurogenic areas during development, so why isn’t lateral inhibition disrupted where these wavefronts contact non-neurogenic tissue? José María Frade, Saúl Ares and colleagues investigate this puzzle on 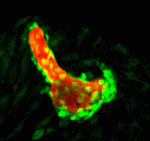 The vertebrate skeleton provides structural support and protection for vital organs but how its component bones acquire their unique shapes is unknown. Here (
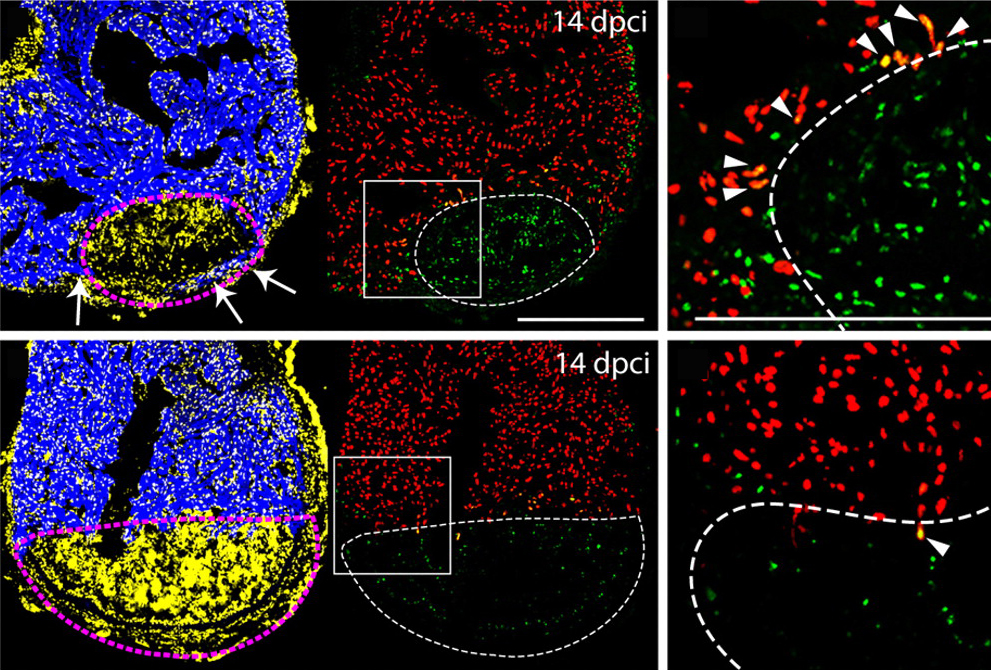
The vertebrate skeleton provides structural support and protection for vital organs but how its component bones acquire their unique shapes is unknown. Here (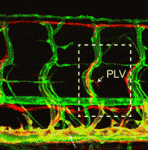 The lymphatic system regulates tissue fluid homeostasis, aids immunity and helps absorb dietary fat. Because aberrant lymphatic growth is associated with cancer metastasis and chronic inflammation, a better understanding of lymphangiogenesis could identify therapeutic targets for these and other lymphatic abnormalities. The major trunk lymphatic vessel in the zebrafish embryo is a well-established model of lymphangiogenesis but the rest of the zebrafish’s lymphatic system is poorly described. On
The lymphatic system regulates tissue fluid homeostasis, aids immunity and helps absorb dietary fat. Because aberrant lymphatic growth is associated with cancer metastasis and chronic inflammation, a better understanding of lymphangiogenesis could identify therapeutic targets for these and other lymphatic abnormalities. The major trunk lymphatic vessel in the zebrafish embryo is a well-established model of lymphangiogenesis but the rest of the zebrafish’s lymphatic system is poorly described. On 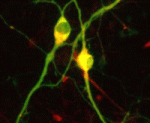 In vitro differentiation of stem cells has the potential to generate specific cell types for clinical use but, to date, this approach has mainly created cells with unsatisfactory phenotypes. Now, Sang-Hun Lee and colleagues generate mature dopamine (DA) neurons from rat neural progenitor cells (NPCs; see
In vitro differentiation of stem cells has the potential to generate specific cell types for clinical use but, to date, this approach has mainly created cells with unsatisfactory phenotypes. Now, Sang-Hun Lee and colleagues generate mature dopamine (DA) neurons from rat neural progenitor cells (NPCs; see 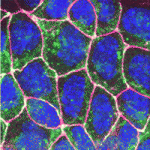 Establishment of the left-right (LR) body axis is a crucial step in embryogenesis. In mouse embryos, a leftward flow of fluid in the node establishes an initial LR signal, which is transferred to the lateral plate mesoderm (LPM) where it triggers the gene expression program responsible for LR asymmetry. But how is the LR signal transferred to the LPM? On
Establishment of the left-right (LR) body axis is a crucial step in embryogenesis. In mouse embryos, a leftward flow of fluid in the node establishes an initial LR signal, which is transferred to the lateral plate mesoderm (LPM) where it triggers the gene expression program responsible for LR asymmetry. But how is the LR signal transferred to the LPM? On 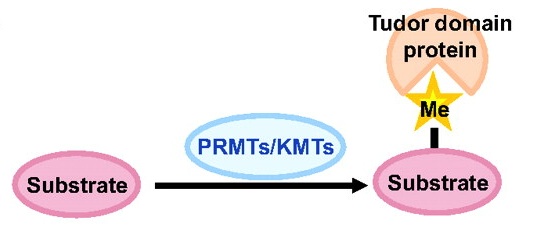 Toshie Kai and colleagues discuss the emerging roles of Tudor domain proteins during development, most notably in the Piwi-interacting RNA pathway, but also in other aspects of RNA metabolism, the DNA damage response and chromatin modification. See the Primer article on p.
Toshie Kai and colleagues discuss the emerging roles of Tudor domain proteins during development, most notably in the Piwi-interacting RNA pathway, but also in other aspects of RNA metabolism, the DNA damage response and chromatin modification. See the Primer article on p. 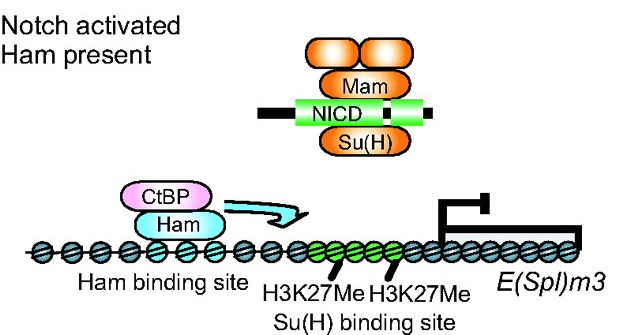 Prdm factors either act as direct histone methyltransferases or recruit a suite of histone-modifying enzymes to target promoters. In their review, Hohenauer and Moore discuss the roles played by these proteins in stem cells and throughout development. See the Review article on p.
Prdm factors either act as direct histone methyltransferases or recruit a suite of histone-modifying enzymes to target promoters. In their review, Hohenauer and Moore discuss the roles played by these proteins in stem cells and throughout development. See the Review article on p. 



 (10 votes)
(10 votes)