Frogs shed new light on the midblastula transition
Posted by Gary McDowell, on 3 September 2013
After fertilization, embryos undergo rapid, synchronous cell divisions until the point of the midblastula transition (MBT) where the cell cycle lengthens. This transition is also known as the maternal-zygotic transition (MZT), as the embryo switches from relying on maternally-deposited mRNAs to undergoing its own transcription.
The trigger for this transition was previously unknown but thought to be related to the increasing ratio of nuclear material to cytoplasm. The embryo does not grow in these early stages, but keeps dividing its cytoplasm between more and more cells. So as the amount of cytoplasm decreases relative to the nuclear component of the cells, it was proposed that at a critical ratio the embryo undergoes a transition where the cell cycles lengthen and become asynchronous, and embryos begin transcribing mRNA.
Now a recent paper in Science has proposed a mechanism for this observation. Four DNA replication factors – Cut5, RecQ4, Treslin and Drf1 – are implicated in the cell cycle elongation, which occurs in concert with a decrease in the density and synchrony of DNA replication initiation events, directly related to a decrease in the abundance of mRNA and protein of these replication factors.
This was demonstrated in vitro using Xenopus egg extracts containing sperm nuclei, by addition of in vitro translated protein of the replication factors. There was an increase in DNA synthesis compared to extract without the addition of protein. This was also carried out with addition of more nuclei to extract to artificially increase the nucleus: cytoplasm ratio, which reduced the rate of synthesis; addition of proteins to this extract increased the rate of synthesis again.
mRNA microinjection, to direct protein overexpression in vivo in Xenopus embryos, of the replication factors demonstrated no slowdown or loss of synchrony in the cell cycle at the MBT and an increase in the number of cells and DNA content compared to controls. Animal caps – the top portion of blastulae that later forms ectoderm – were dissected out of injected embryos and dissociated, then used for pulse-chase labelling of replication forks, showing that overexpression of the replication factors resulted in an increased rate of DNA replication. This was achieved not by affecting elongation, but rather increasing the number of origins of replication.
The authors also found that in embryos overexpressing the four replication factors, there was an earlier activation of the cell cycle checkpoint kinase Chk1, normally activated during the MBT, that had been linked previously to the depletion of the nucleotides required for DNA synthesis.
Whilst overexpressing the four replication factors resulted in increased cell number and no cell cycle lengthening, these embryos were also severely restricted in their ability to undergo gastrulation and form closed blastopores, with high rates of embryo death by the stage of neurulation. This phenotype was rescued partially by co-injection of morpholinos, small synthetic oligonucleotides that prevent mRNA translation, targeting Cdc6, a protein involved in prereplicative complex formation. The idea behind targeting this protein was to reduce the extent of origin licensing, illustrating that developmental problems caused by overexpression of the replication factors were indeed caused by increased rates of replication initiation. The rescued embryos also showed normal lengthening of the cell cycle at the MBT as well as normal activation of Chk1.
Overall, the paper has illustrated the role of replication initiation rates in regulating normal embryonic development, tying in nicely to the observation that an increasing nuclear: cytoplasmic ratio correlates with events at the MBT. Furthermore the authors speculate that this may be an important mechanism in regulating the length of S-phase across development, and across eukaryotes.
References:
Collart, C. et al. Titration of four replication factors is essential for the Xenopus laevis midblastula transition. Science 341, 893-896 (2013).


 (5 votes)
(5 votes) (No Ratings Yet)
(No Ratings Yet)








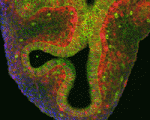 Lung development in mice involves specification of the primary lung field followed by the formation of lung buds, which subsequently undergo outgrowth and branching morphogenesis to form the stereotypic bronchial tree. Localised expression of Fgf10 in the distal mesenchyme adjacent to the sites of lung bud formation has long been thought to drive branching morphogenesis in the lung but now, on
Lung development in mice involves specification of the primary lung field followed by the formation of lung buds, which subsequently undergo outgrowth and branching morphogenesis to form the stereotypic bronchial tree. Localised expression of Fgf10 in the distal mesenchyme adjacent to the sites of lung bud formation has long been thought to drive branching morphogenesis in the lung but now, on 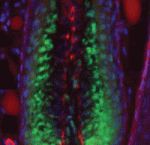 Hair follicles cyclically degenerate and regenerate through adult life: after an initial growth phase, hair follicles enter a destructive phase and then go through a quiescent stage before re-entering the next growth phase. This cycling involves hair follicle stem cells (HFSCs) but how these cells transition between the phases of the hair follicle cycle is unclear. Here, Hoang Nguyen and colleagues report that the forkhead transcription factor Foxp1 is crucial for maintaining HFSC quiescence (
Hair follicles cyclically degenerate and regenerate through adult life: after an initial growth phase, hair follicles enter a destructive phase and then go through a quiescent stage before re-entering the next growth phase. This cycling involves hair follicle stem cells (HFSCs) but how these cells transition between the phases of the hair follicle cycle is unclear. Here, Hoang Nguyen and colleagues report that the forkhead transcription factor Foxp1 is crucial for maintaining HFSC quiescence (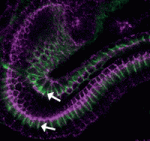 The molecular mechanisms that link intracellular signalling pathways to changes in tissue morphology are unclear. Using the Drosophila embryonic hindgut as a model, Martin Zeidler and co-workers demonstrate that the transmembrane protein Fasciclin III (FasIII) regulates intracellular adhesion and links signal transduction to morphogenesis (
The molecular mechanisms that link intracellular signalling pathways to changes in tissue morphology are unclear. Using the Drosophila embryonic hindgut as a model, Martin Zeidler and co-workers demonstrate that the transmembrane protein Fasciclin III (FasIII) regulates intracellular adhesion and links signal transduction to morphogenesis (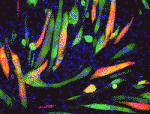 Muscle development is driven by a set of myogenic factors, but how these are regulated during normal development and during regeneration is unclear. Here (
Muscle development is driven by a set of myogenic factors, but how these are regulated during normal development and during regeneration is unclear. Here (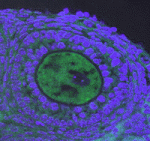 The piRNA pathway silences retrotransposons and hence maintains genome integrity in the germline. Several components of the piRNA pathway localise to a structure called the nuage, which has been detected in many animal germlines, including mouse testes and Drosophila oocytes. Now, Ai Khim Lim, Barbara Knowles and colleagues show that a nuage-like structure can be found in mouse oocytes (
The piRNA pathway silences retrotransposons and hence maintains genome integrity in the germline. Several components of the piRNA pathway localise to a structure called the nuage, which has been detected in many animal germlines, including mouse testes and Drosophila oocytes. Now, Ai Khim Lim, Barbara Knowles and colleagues show that a nuage-like structure can be found in mouse oocytes (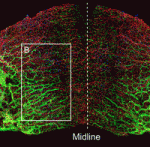 Lymphangiogenesis, the formation of lymphatic vessels, involves multiple growth factors and receptors, including vascular endothelial growth factor C (VEGFC) and its receptor VEGFR3. Here, on
Lymphangiogenesis, the formation of lymphatic vessels, involves multiple growth factors and receptors, including vascular endothelial growth factor C (VEGFC) and its receptor VEGFR3. Here, on 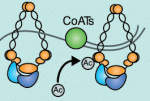 Recent studies have shown that cohesin, which was named for its ability to mediate sister chromatid cohesion, can influence gene expression during development. Here, Ana Losada and colleagues provide an overview of how cohesin functions in development and disease. See the Development at a Glance poster article on p.
Recent studies have shown that cohesin, which was named for its ability to mediate sister chromatid cohesion, can influence gene expression during development. Here, Ana Losada and colleagues provide an overview of how cohesin functions in development and disease. See the Development at a Glance poster article on p. 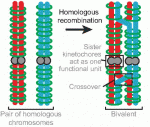 Mammalian oocytes are particularly error prone in segregating their chromosomes during their two meiotic divisions, resulting in the creation of an embryo that has inherited the wrong number of chromosomes: it is aneuploid. Here, Keith Jones and Simon Lane review recent data on factors that determine successful segregation in female meiosis and explain how this might be related to an age-related decline in female segregation accuracy. See the Primer article on p.
Mammalian oocytes are particularly error prone in segregating their chromosomes during their two meiotic divisions, resulting in the creation of an embryo that has inherited the wrong number of chromosomes: it is aneuploid. Here, Keith Jones and Simon Lane review recent data on factors that determine successful segregation in female meiosis and explain how this might be related to an age-related decline in female segregation accuracy. See the Primer article on p.